Nexelia Academy · Official Revision Notes
Complete A-Level revision notes · 19 chapters
This chapter explores cells as the fundamental units of life, differentiating between simpler prokaryotic and complex eukaryotic cells. It details the structure and function of various organelles, explains the principles of microscopy including magnification and resolution, and compares the unique features of bacteria and viruses.
cell — The basic unit of all living organisms; it is surrounded by a cell surface membrane and contains genetic material (DNA) and cytoplasm containing organelles.
Cells are the fundamental building blocks of life, separating internal biochemical reactions from the external environment. Their partially permeable membrane is crucial for controlling material exchange, maintaining the distinct internal conditions necessary for life. Think of a cell like a tiny, self-contained factory. It has walls (cell membrane), machinery (organelles), a control room (nucleus/DNA), and a fluid environment (cytoplasm) where work happens, all separated from the outside world.
organelle — A functionally and structurally distinct part of a cell, e.g. a ribosome or mitochondrion.
Organelles are like 'little organs' within the cell, each performing a specialised task. Many are membrane-bound, allowing their activities to be compartmentalised and separated from the surrounding cytoplasm, which increases cellular efficiency. If a cell is a factory, organelles are the specialised machines or departments within it, like the power generator (mitochondrion), the assembly line (ribosome), or the packaging and shipping department (Golgi apparatus).
eukaryote — An organism whose cells contain a nucleus and other membrane-bound organelles.
Eukaryotic cells are typically larger and more complex than prokaryotic cells, characterised by the presence of a true nucleus ('eu' means true, 'karyon' means nucleus) and a system of internal membranes that form various organelles. This compartmentalisation allows for greater specialisation and efficiency in cellular processes. Eukaryotes are like modern, multi-room houses with specialised areas (kitchen, bedroom, bathroom), whereas prokaryotes are more like single-room studios.
prokaryote — An organism whose cells do not contain a nucleus or any other membrane-bound organelles.
Prokaryotic cells are simpler and generally smaller than eukaryotic cells, lacking a membrane-bound nucleus and other internal membrane-bound organelles. Their genetic material (DNA) is typically circular and free in the cytoplasm, and they are thought to be the earliest forms of life. Prokaryotes are like a basic, open-plan workshop where all tools and materials are in one main space, unlike the compartmentalised factory of a eukaryote.
Magnification
Ensure 'observed size of the image' and 'actual size' are in the same units before calculation. This formula can be rearranged to find any of the three variables if the other two are known.
Cells are the basic units of life, and their study relies heavily on microscopy. Light microscopes use light and lenses to magnify specimens, allowing observation of common structures. Electron microscopes, however, use electron beams, providing significantly higher resolution and magnification, which reveals the intricate ultrastructure of cells. Understanding magnification and resolution is crucial for interpreting microscopic images.
magnification — The number of times larger an image of an object is than the real size of the object; magnification = image size ÷ actual (real) size of the object.
Magnification is the extent to which an image is enlarged compared to the actual specimen. While high magnification makes an object appear larger, it does not necessarily reveal more detail unless accompanied by high resolution. Magnification is like zooming in on a photo; it makes the image bigger, but if the original photo was blurry, zooming in won't make it clearer.
resolution — The ability to distinguish between two objects very close together; the higher the resolution of an image, the greater the detail that can be seen.
Resolution determines the clarity and detail of an image, indicating the minimum distance at which two separate points can still be seen as distinct. It is limited by the wavelength of the radiation used for viewing, with shorter wavelengths (like electrons) providing higher resolution. Resolution is like the sharpness of a TV screen; a higher resolution screen shows more distinct details and less blur, even if the image is magnified.
Students often confuse magnification with resolution. Remember that magnification is about making an image larger, while resolution is about seeing more detail and distinguishing between two close points.
Practice calculating magnification and actual size using the formula: Magnification = Image size / Actual size. Ensure consistent units (e.g., both in μm or both in mm) before calculation to avoid errors.
eyepiece graticule — Small scale that is placed in a microscope eyepiece.
An eyepiece graticule is a transparent ruler inserted into the microscope eyepiece, allowing the observer to measure the size of specimens in arbitrary 'eyepiece units'. It must be calibrated using a stage micrometer to convert these units into actual measurements. The eyepiece graticule is like a transparent ruler placed directly over your eye when looking through a magnifying glass, allowing you to measure what you see.
stage micrometer — Very small, accurately drawn scale of known dimensions, engraved on a microscope slide.
A stage micrometer is a precisely measured scale placed on the microscope stage, used to calibrate the eyepiece graticule. By superimposing the two scales, the value of each eyepiece unit can be determined for a specific objective lens magnification. The stage micrometer is like a standard ruler used to check and set the accuracy of a custom-made ruler (the eyepiece graticule).
Remember that the eyepiece graticule needs calibration with a stage micrometer to convert arbitrary units into actual lengths, and this calibration must be done for each objective lens.
micrograph — A picture taken with the aid of a microscope; a photomicrograph (or light micrograph) is taken using a light microscope; an electron micrograph is taken using an electron microscope.
Micrographs are photographic records of microscopic specimens, providing visual documentation of cellular structures. The type of micrograph (light or electron) indicates the microscope used and thus the level of detail and resolution captured. A micrograph is like a photograph taken through a telescope, capturing an image of something far away and making it visible for study.
Eukaryotic cells, both animal and plant, share several fundamental organelles that perform vital functions. These include the cell surface membrane, nucleus, cytoplasm, mitochondria, and Golgi apparatus. Each organelle is structurally adapted to its specific role, contributing to the overall efficiency and survival of the cell.
cell surface membrane — A very thin membrane (about 7 nm diameter) surrounding all cells; it is partially permeable and controls the exchange of materials between the cell and its environment.
This essential membrane forms the outer boundary of every cell, regulating what enters and leaves. Its partial permeability is vital for maintaining the cell's internal environment, allowing necessary substances in while keeping harmful ones out and retaining essential cellular components. Think of the cell surface membrane as the security gate and border control of a city (the cell), carefully checking and controlling all traffic (materials) entering and exiting.
Students often confuse the cell surface membrane with the cell wall. Remember that the cell surface membrane is present in all cells and is partially permeable, while the cell wall is external to it in plants/bacteria and is freely permeable.
protoplasm — All the living material inside a cell (cytoplasm plus nucleus).
Protoplasm encompasses all the living components of a cell, including the nucleus and the cytoplasm. It represents the active, functional substance of the cell, where all metabolic processes occur. Protoplasm is like the entire 'living content' of a house, including the furniture, people, and air, as opposed to just the empty structure.
cytoplasm — The contents of a cell, excluding the nucleus.
Cytoplasm is the jelly-like substance that fills the cell and surrounds the organelles, providing a medium for many biochemical reactions. It consists of the cytosol (the fluid portion) and the organelles suspended within it, playing a crucial role in cell metabolism and transport. If the cell is a house, the cytoplasm is all the space and contents within the walls, excluding the main office (nucleus).
nucleus — A relatively large organelle found in eukaryotic cells, but absent from prokaryotic cells; the nucleus contains the cell’s DNA and therefore controls the activities of the cell; it is surrounded by two membranes which together form the nuclear envelope.
The nucleus is the control centre of eukaryotic cells, housing the genetic material (DNA) organised into chromosomes. Its double membrane, the nuclear envelope, regulates the passage of molecules between the nucleus and cytoplasm through nuclear pores, ensuring precise control over cell activities. The nucleus is like the main office or control room of a factory, where all the blueprints (DNA) and instructions for running the entire operation are stored and managed.
nuclear envelope — The two membranes, situated close together, that surround the nucleus; the envelope is perforated with nuclear pores.
The nuclear envelope is a double membrane that encloses the nucleus in eukaryotic cells, separating the genetic material from the cytoplasm. Its continuity with the endoplasmic reticulum and the presence of nuclear pores are crucial for regulating molecular traffic between the nucleus and cytoplasm. The nuclear envelope is like the double-layered wall of the control room (nucleus), with guarded doorways (nuclear pores) controlling who and what enters and leaves.
nuclear pores — Pores found in the nuclear envelope which control the exchange of materials, e.g. mRNA, between the nucleus and the cytoplasm.
Nuclear pores are complex protein structures embedded in the nuclear envelope that regulate the bidirectional transport of macromolecules, such as proteins and RNA, between the nucleus and the cytoplasm. This controlled exchange is vital for gene expression and cellular function. Nuclear pores are like the security checkpoints or gates in the wall of the control room (nucleus), allowing specific authorised personnel (molecules) to pass through.
chromatin — The material of which chromosomes are made, consisting of DNA, proteins and small amounts of RNA; visible as patches or fibres within the nucleus when stained.
Chromatin is the complex of DNA and proteins (primarily histones) that forms chromosomes within the nucleus of eukaryotic cells. Its coiled structure allows the long DNA molecules to be compactly stored and organised, preventing tangling and facilitating gene regulation. Imagine chromatin as a very long thread (DNA) wound around spools (proteins) and then further coiled into a ball, making it manageable and organised within a small space.
chromosome — In the nucleus of the cells of eukaryotes, a structure made of tightly coiled chromatin (DNA, proteins and RNA) visible during cell division; the term ‘circular DNA’ is now also commonly used for the circular strand of DNA present in a prokaryotic cell.
Chromosomes are highly organised structures containing the cell's genetic material, DNA, along with proteins. In eukaryotes, they become visible as distinct structures during cell division when chromatin condenses, ensuring accurate segregation of genetic information to daughter cells. Prokaryotes have a single circular chromosome. If DNA is a long string of instructions, a chromosome is like a neatly bound and organised book containing those instructions, ready to be copied and distributed.
nucleolus — A small structure, one or more of which is found inside the nucleus; the nucleolus is usually visible as a densely stained body; its function is to manufacture ribosomes using the information in its own DNA.
The nucleolus is a prominent region within the nucleus responsible for synthesising ribosomal RNA (rRNA) and assembling ribosomal subunits. Its size often correlates with the cell's protein synthesis activity, as ribosomes are essential for this process. The nucleolus is like a small ribosome factory within the main control room (nucleus), producing the essential machinery for protein production.
endoplasmic reticulum (ER) — A network of flattened sacs running through the cytoplasm of eukaryotic cells; molecules, particularly proteins, can be transported through the cell inside the sacs separate from the rest of the cytoplasm; ER is continuous with the outer membrane of the nuclear envelope.
The ER is an extensive network of interconnected membranes that forms flattened sacs (cisternae) and tubules throughout the cytoplasm. It serves as a transport system and is involved in protein synthesis (rough ER) and lipid synthesis, detoxification, and calcium storage (smooth ER). The ER is like a network of interconnected highways and workshops within the cell, where materials are transported and processed in a segregated environment.
ribosome — A tiny organelle found in large numbers in all cells; prokaryotic ribosomes are about 20 nm in diameter while eukaryotic ribosomes are about 25 nm in diameter.
Ribosomes are essential organelles responsible for protein synthesis, translating messenger RNA into polypeptide chains. They consist of two subunits (large and small) and are found free in the cytoplasm or attached to the rough endoplasmic reticulum, with prokaryotic ribosomes (70S) being slightly smaller than eukaryotic ones (80S). Ribosomes are like the small assembly machines or 3D printers in a factory, taking instructions (mRNA) and building products (proteins).
Students often think all organelles are membrane-bound. Remember that some, like ribosomes and centrioles, are not.
Golgi apparatus (Golgi body, Golgi complex) — An organelle found in eukaryotic cells; the Golgi apparatus consists of a stack of flattened sacs, constantly forming at one end and breaking up into Golgi vesicles at the other end.
The Golgi apparatus is a stack of flattened membrane-bound sacs (cisternae) that modifies, sorts, and packages proteins and lipids synthesised in the ER. It processes molecules, adds sugars to proteins (glycoproteins) and lipids (glycolipids), and then dispatches them in Golgi vesicles to other cellular destinations or for secretion. The Golgi apparatus is like the cell's post office or packaging and distribution centre, receiving, modifying, sorting, and shipping out products (proteins, lipids) in packages (vesicles).
Golgi vesicles — Carry their contents to other parts of the cell, often to the cell surface membrane for secretion; the Golgi apparatus chemically modifies the molecules it transports, e.g. sugars may be added to proteins to make glycoproteins.
Golgi vesicles are small, membrane-bound sacs that bud off from the Golgi apparatus, transporting processed molecules to various destinations within the cell or to the cell surface membrane for secretion outside the cell. They are crucial for the secretory pathway and for forming other organelles like lysosomes. Golgi vesicles are like delivery trucks or packages leaving the post office (Golgi apparatus), carrying their contents to different addresses inside or outside the city (cell).
lysosome — A spherical organelle found in eukaryotic cells; it contains digestive (hydrolytic) enzymes and has a variety of destructive functions, such as removal of old cell organelles.
Lysosomes are membrane-bound organelles containing digestive enzymes that break down waste materials and cellular debris. They are involved in recycling old organelles, digesting foreign particles (e.g., in phagocytosis), and can even trigger programmed cell death. Lysosomes are like the recycling and waste disposal units of the cell, breaking down unwanted materials.
mitochondrion — The organelle in eukaryotes in which aerobic respiration takes place.
Mitochondria are often called the 'powerhouses' of the cell because they are responsible for generating most of the cell's supply of adenosine triphosphate (ATP) through aerobic respiration. They have a double membrane, with the inner membrane folded into cristae to increase surface area for these reactions. Mitochondria are like the power plants of a city (the cell), constantly generating energy (ATP) to fuel all its activities.
cristae — Folds of the inner membrane of the mitochondrial envelope on which are found stalked particles of ATP synthase and electron transport chains associated with aerobic respiration.
Cristae are the numerous folds of the inner mitochondrial membrane, significantly increasing its surface area. This extensive surface is crucial for housing the enzymes and electron transport chains necessary for aerobic respiration and efficient ATP production. Cristae are like the folded internal walls of a power plant, maximising the space for energy-generating machinery.
ATP (adenosine triphosphate) — The molecule that is the universal energy currency in all living cells; the purpose of respiration is to make ATP.
ATP is the primary energy currency of the cell, providing the energy required for almost all cellular processes, including muscle contraction, active transport, and synthesis of macromolecules. It stores energy in its phosphate bonds, which is released upon hydrolysis to ADP. ATP is like the rechargeable battery of the cell, storing and releasing energy as needed.
ADP (adenosine diphosphate) — The molecule that is converted to ATP by addition of phosphate (a reaction known as phosphorylation) during cell respiration; the enzyme responsible is ATP synthase; the reaction requires energy.
ADP is a lower-energy molecule that is phosphorylated to form ATP during cellular respiration, a process that captures energy released from nutrient breakdown. This reversible conversion between ADP and ATP is central to energy transfer within the cell. ADP is like a partially discharged battery, ready to be recharged into ATP.
Always link mitochondria to aerobic respiration and ATP production, and mention the cristae for increased surface area, as these are key structural-functional relationships.
microtubules — Tiny tubes made of a protein called tubulin and found in most eukaryotic cells; microtubules have a large variety of functions, including cell support and determining cell shape; the ‘spindle’ on which chromatids and chromosomes separate during nuclear division is made of microtubules.
Microtubules are hollow cylinders made of tubulin protein, forming part of the cytoskeleton. They provide structural support, maintain cell shape, and are involved in intracellular transport, cell division (forming the spindle fibres), and the movement of cilia and flagella. Microtubules are like the scaffolding and railway tracks of the cell, providing structure and guiding movement.
centriole — One of two small, cylindrical structures, made from microtubules, found just outside the nucleus in animal cells, in a region known as the centrosome; they are also found at the bases of cilia and flagella.
Centrioles are cylindrical structures composed of microtubules, typically found in pairs within the centrosome of animal cells. They play a crucial role in cell division by organising spindle fibres and are also involved in the formation of cilia and flagella. Centrioles are like the anchors or organisers for the cell's internal transport and division machinery.
centrosome — The main microtubule organising centre (MTOC) in animal cells.
The centrosome is a region in animal cells that serves as the main microtubule-organising centre (MTOC). It contains two centrioles and is responsible for initiating microtubule growth and organising the mitotic spindle during cell division. The centrosome is like the central hub from which the cell's internal structural elements are built and managed.
cilia — Whip-like structures projecting from the surface of many animal cells and the cells of many unicellular organisms; they beat, causing locomotion or the movement of fluid across the cell surface.
Cilia are short, hair-like appendages that project from the cell surface, composed of microtubules. They beat in a coordinated fashion to move fluids over the cell surface (e.g., in the respiratory tract) or to propel unicellular organisms. Cilia are like tiny oars on the surface of a boat, moving fluid or the cell itself.
flagella — Whip-like structures projecting from the surface of some animal cells and the cells of many unicellular organisms; they beat, causing locomotion or the movement of fluid across the cell surface; they are identical in structure to cilia, but longer.
Flagella are longer, whip-like appendages similar in structure to cilia, also composed of microtubules. They are primarily involved in cell locomotion, propelling cells through fluid environments (e.g., sperm cells). Flagella are like a single, powerful propeller on a boat, driving it forward.
While sharing many eukaryotic features, plant and animal cells exhibit key structural differences. Plant cells possess a rigid cell wall, a large central vacuole, and chloroplasts, which are absent in animal cells. These unique structures enable plants to perform photosynthesis, maintain turgor, and provide structural support.
cell wall — A wall surrounding prokaryote, plant and fungal cells; the wall contains a strengthening material which protects the cell from mechanical damage, supports it and prevents it from bursting by osmosis if the cell is surrounded by a solution with a higher water potential.
The cell wall is a rigid outer layer that provides structural support and protection to plant, fungal, and prokaryotic cells. Unlike the cell surface membrane, it is freely permeable, allowing water and solutes to pass through, and its strength helps maintain cell shape and turgor pressure. The cell wall is like the strong, rigid outer brick wall of a building, providing structural support and protection, while the cell membrane is the flexible inner lining.
plasmodesma — A pore-like structure found in plant cell walls; plasmodesmata of neighbouring plant cells line up to form tube-like pores through the cell walls, allowing the controlled passage of materials from one cell to the other; the pores contain ER and are lined with the cell surface membrane.
Plasmodesmata are microscopic channels that traverse the cell walls of adjacent plant cells, connecting their cytoplasm and endoplasmic reticulum. They facilitate intercellular communication and transport of water, nutrients, and signalling molecules, forming the symplast pathway. Plasmodesmata are like small tunnels or bridges connecting neighbouring houses (plant cells), allowing residents to share resources and communicate directly without going outside.
vacuole — An organelle found in eukaryotic cells; a large, permanent central vacuole is a typical feature of plant cells, where it has a variety of functions, including storage of biochemicals such as salts, sugars and waste products; temporary vacuoles, such as phagocytic vacuoles (also known as phagocytic vesicles), may form in animal cells.
In plant cells, the large central vacuole plays a critical role in maintaining turgor pressure, storing water, nutrients, and waste products, and can also act as a lysosome. Animal cells may have smaller, temporary vacuoles involved in processes like phagocytosis. The plant vacuole is like a multi-purpose storage tank and pressure regulator in a house, holding water, supplies, and waste, and helping to keep the house rigid.
Students often think vacuoles are exclusive to plant cells. Remember that animal cells can have small, temporary vacuoles, though they lack the large, permanent central vacuole characteristic of plants.
tonoplast — The partially permeable membrane that surrounds plant vacuoles.
The tonoplast is a single membrane that encloses the central vacuole in plant cells, controlling the movement of substances between the cytoplasm and the vacuolar sap. Its selective permeability is essential for maintaining the vacuole's internal environment and regulating turgor pressure. The tonoplast is the specific 'skin' or boundary of the plant cell's storage tank (vacuole), regulating what goes in and out of it.
chloroplast — An organelle, bounded by an envelope (i.e. two membranes), in which photosynthesis takes place in eukaryotes.
Chloroplasts are the sites of photosynthesis in plant and algal cells, converting light energy into chemical energy. They contain photosynthetic pigments like chlorophyll within their internal membrane system of thylakoids, which are often stacked into grana, increasing the surface area for light absorption. Chloroplasts are like the solar panels of a plant cell, capturing sunlight and converting it into usable energy (sugars).
photosynthesis — The production of organic substances from inorganic ones, using energy from light.
Photosynthesis is the fundamental process by which green plants, algae, and some bacteria convert light energy, water, and carbon dioxide into glucose (an organic substance) and oxygen. This process is vital for sustaining most life on Earth, forming the base of many food chains. Photosynthesis is like a chef using sunlight as energy to cook raw ingredients (carbon dioxide and water) into food (sugars) for the plant.
grana — Stacks of membranes inside a chloroplast.
Grana (singular: granum) are stacks of flattened, disc-like thylakoids within the chloroplast stroma. These stacks increase the surface area for the light-dependent reactions of photosynthesis, where chlorophyll and other pigments capture light energy. Grana are like stacks of pancakes (thylakoids) inside the chloroplast, where each pancake's surface is covered with light-absorbing ingredients.
thylakoid — A flattened, membrane-bound, fluid-filled sac which is the site of the light-dependent reactions of photosynthesis in a chloroplast.
Thylakoids are flattened, membrane-bound sacs within chloroplasts, where the light-dependent reactions of photosynthesis occur. They contain chlorophyll and other photosynthetic pigments embedded in their membranes, capturing light energy. Thylakoids are like individual solar panels within the chloroplast, absorbing sunlight.
phospholipid — A lipid to which phosphate is added; the molecule is made up of a glycerol molecule, two fatty acids and a phosphate group; a double layer (a bilayer) of phospholipids forms the basic structure of all cell membranes.
Phospholipids are a major component of cell membranes, forming a bilayer due to their amphipathic nature (hydrophilic head and hydrophobic tails). This structure creates a selectively permeable barrier essential for cell function. Phospholipids are like the building blocks of a flexible, self-sealing wall, forming the boundary of the cell.
Bacteria are single-celled prokaryotic microorganisms, characterised by their simpler cellular organisation compared to eukaryotes. They lack a membrane-bound nucleus and other membrane-bound organelles, with their genetic material (circular DNA) free in the cytoplasm. Despite their simplicity, bacteria possess essential structures like a cell wall, cell surface membrane, and ribosomes, and may have additional features such as a flagellum, capsule, or plasmids.
bacteria — A group of single-celled prokaryotic microorganisms; they have a number of characteristics, such as the ability to form spores, which distinguish them from the other group of prokaryotes known as Archaea.
Bacteria are a diverse group of prokaryotic organisms, meaning their cells lack a membrane-bound nucleus and other membrane-bound organelles. They are ubiquitous and play crucial roles in various ecosystems, from nutrient cycling to disease. Bacteria are like the ancient, efficient single-room workshops of the biological world, capable of performing all life functions without complex internal compartments.
peptidoglycan — A polysaccharide combined with amino acids; it is also known as murein; it makes the bacterial cell wall more rigid.
Peptidoglycan, also known as murein, is a unique polymer found in the cell walls of most bacteria. Its strong, mesh-like structure provides rigidity and protection to the bacterial cell, preventing osmotic lysis. Peptidoglycan is like the chainmail armour of a bacterium, providing strength and protection.
plasmid — A small circular piece of DNA in a bacterium (not its main chromosome); plasmids often contain genes that provide resistance to antibiotics.
Plasmids are small, circular, extrachromosomal DNA molecules found in bacteria, separate from the main circular chromosome. They often carry genes that confer advantageous traits, such as antibiotic resistance, and can be transferred between bacteria, contributing to genetic diversity. Plasmids are like small, optional instruction manuals that a bacterium can pick up, giving it extra abilities.
pili (singular: pilus) — Hair-like appendages on the surface of many bacteria; they are involved in attachment to surfaces and other cells, and in bacterial conjugation (transfer of genetic material).
Pili are short, hair-like protein structures projecting from the surface of many bacteria. They are primarily involved in adhesion to host cells or other surfaces, and some specialized pili (sex pili) facilitate the transfer of genetic material between bacteria during conjugation. Pili are like tiny grappling hooks or sticky hairs that help bacteria attach and interact with their environment.
Students often think bacterial flagella have the same complex '9 + 2' microtubule structure as eukaryotic flagella. Remember that bacterial flagella are much simpler and rotate like a propeller.
Viruses are distinct from both prokaryotic and eukaryotic cells as they are acellular, meaning they are not composed of cells. They are obligate intracellular parasites, requiring a living host cell to replicate. A virus typically consists of genetic material (DNA or RNA) enclosed within a protein coat, and sometimes an outer lipid envelope.
virus — A very small (20–300 nm) infectious particle which can replicate only inside living cells; it consists of a molecule of DNA or RNA (the genome) surrounded by a protein coat; an outer lipid envelope may also be present.
Viruses are non-cellular infectious agents that can only replicate by infecting living host cells. They consist of genetic material (DNA or RNA) encased in a protein coat (capsid), and sometimes an outer lipid envelope derived from the host cell membrane. Viruses are like tiny, parasitic instruction manuals that hijack a factory (host cell) to make more copies of themselves.
Students often think viruses are living cells. Remember that they are acellular and obligate intracellular parasites, meaning they can only replicate inside living host cells.
When asked to 'outline' structure and function, provide concise descriptions for each organelle, linking structure to its role.
Be precise when comparing prokaryotic and eukaryotic cells; focus on the presence/absence of a nucleus, membrane-bound organelles, and DNA form.
For microscopy questions, clearly state the advantages of electron microscopes (higher resolution, greater magnification) over light microscopes.
When drawing cells, include clear labels for all visible structures and indicate magnification if provided.
Remember the role of ATP as the universal energy currency, produced primarily by mitochondria in eukaryotes.
Definitions Bank
cell
The basic unit of all living organisms; it is surrounded by a cell surface membrane and contains genetic material (DNA) and cytoplasm containing organelles.
organelle
A functionally and structurally distinct part of a cell, e.g. a ribosome or mitochondrion.
nucleus
A relatively large organelle found in eukaryotic cells, but absent from prokaryotic cells; the nucleus contains the cell’s DNA and therefore controls the activities of the cell; it is surrounded by two membranes which together form the nuclear envelope.
eukaryote
An organism whose cells contain a nucleus and other membrane-bound organelles.
prokaryote
An organism whose cells do not contain a nucleus or any other membrane-bound organelles.
+40 more definitions
View all →Command Word Guide
| Describe | Provide a detailed account of the structure or features of a cell or organelle, without explaining why or how it functions. |
| Explain | Give reasons or mechanisms for a particular structure or function, linking cause and effect. For example, explain how the structure of mitochondria relates to its function in ATP production. |
| Compare | Identify both similarities and differences between two or more entities, such as prokaryotic and eukaryotic cells, or plant and animal cells. Use comparative language (e.g., 'whereas', 'both'). |
| Outline | Give a brief summary of the main points, without extensive detail. For example, outline the role of ATP in cells. |
+1 more
View all →Common Mistakes
Students often confuse magnification with resolution.
Magnification is about making an image larger, while resolution is about seeing more detail and distinguishing between two close points.
Students often think all organelles are membrane-bound.
Some organelles, like ribosomes and centrioles, do not have membranes.
Students often confuse the cell wall with the cell surface membrane.
The cell wall is external, rigid, and freely permeable (in plants/bacteria), while the cell surface membrane is internal, thin, partially permeable, and present in all cells.
+3 more
View all →This chapter explores the essential biological molecules: carbohydrates, lipids, proteins, and water. It details their structures, functions, and how large macromolecules are built from smaller monomers via condensation reactions and broken down by hydrolysis. Key properties of water and biochemical identification tests are also covered.
macromolecule — A large molecule such as a polysaccharide, protein or nucleic acid.
Macromolecules are giant molecules formed from smaller repeating subunits. They are essential for life, performing diverse functions from energy storage to structural support and genetic information. Think of a macromolecule like a long train, where each carriage is a smaller subunit (monomer) linked together to form a much larger structure.
Students often think all large molecules are macromolecules, but actually the term specifically refers to polymers like polysaccharides, proteins, and nucleic acids, which are built from repeating monomer units.
polymer — A giant molecule made from many similar repeating subunits joined together in a chain; the subunits are much smaller and simpler molecules known as monomers; examples of biological polymers are polysaccharides, proteins and nucleic acids.
Polymers are formed through condensation reactions where monomers are linked by covalent bonds, with the removal of water. This repeating process allows for the creation of complex structures from simple building blocks. A polymer is like a pearl necklace, where each pearl is a monomer, and the entire necklace is the polymer.
When identifying polymers, remember to state that they are made of 'repeating subunits' (monomers) and give specific biological examples like starch or protein, not just 'large molecules'.
monomer — A relatively simple molecule which is used as a basic building block for the synthesis of a polymer; many monomers are joined together by covalent bonds to make the polymer, usually by condensation reactions; common examples of monomers are monosaccharides, amino acids and nucleotides.
Monomers are the fundamental units that link together to form polymers. The specific type of monomer determines the properties and function of the resulting polymer. A monomer is like a single LEGO brick; many identical or similar bricks can be joined together to build a larger, more complex LEGO model (the polymer).
When asked to name monomers, be precise: 'monosaccharides' for carbohydrates, 'amino acids' for proteins, and 'nucleotides' for nucleic acids. Avoid generic terms.
condensation reaction — A chemical reaction involving the joining together of two molecules by removal of a water molecule.
This reaction is crucial for building larger biological molecules (polymers) from smaller ones (monomers). The formation of glycosidic, ester, and peptide bonds are all examples of condensation reactions. Imagine two people holding hands; a condensation reaction is like them joining hands and a drop of water falling away as they connect.
Always mention the 'removal of a water molecule' when describing condensation reactions, as this is a key part of the definition and mechanism.
hydrolysis — A chemical reaction in which a chemical bond is broken by the addition of a water molecule; commonly used to break down complex molecules into simpler molecules.
Hydrolysis is the reverse of a condensation reaction, essential for digestion and breaking down polymers into their constituent monomers, making nutrients available for absorption or recycling. If condensation is like two people holding hands and a drop of water falling away, hydrolysis is like adding a drop of water to their hands, causing them to let go.
Large biological molecules, or macromolecules, are fundamental to life. These include carbohydrates, lipids, and proteins. They are typically polymers, meaning they are constructed from smaller, repeating monomer units. The synthesis of these polymers from monomers occurs through condensation reactions, where a water molecule is removed. Conversely, these polymers can be broken down into their constituent monomers via hydrolysis, a reaction that involves the addition of a water molecule.
monosaccharide — A molecule consisting of a single sugar unit and with the general formula (CH2O)n.
Monosaccharides are the simplest carbohydrates, serving as primary energy sources (e.g., glucose) and building blocks for disaccharides and polysaccharides. They are typically sweet-tasting and water-soluble. A monosaccharide is like a single sugar cube; it's the smallest, most basic unit of sugar.
When asked for examples of monosaccharides, name specific ones like glucose, fructose, galactose (hexoses) or ribose, deoxyribose (pentoses), and remember their general formula (CH2O)n.
disaccharide — A sugar molecule consisting of two monosaccharides joined together by a glycosidic bond.
Disaccharides are formed by a condensation reaction between two monosaccharides, such as maltose (glucose + glucose) or sucrose (glucose + fructose). They are soluble sugars with roles in energy transport and diet. A disaccharide is like two sugar cubes stuck together; it's still sweet and soluble, but a bit larger than a single cube.
Students often think all disaccharides are reducing sugars, but actually sucrose is a common non-reducing sugar.
glycosidic bond — A C–O–C link between two sugar molecules, formed by a condensation reaction; it is a covalent bond.
This covalent bond is formed when two hydroxyl groups from monosaccharides react, releasing a water molecule. It is the fundamental linkage in disaccharides and polysaccharides. It's like a small oxygen bridge connecting two sugar molecules, holding them firmly together.
Benedict’s test — A test for the presence of reducing sugars; the unknown substance is heated with Benedict’s reagent, and a change from a clear blue solution to the production of a yellow, red or brown precipitate indicates the presence of reducing sugars such as glucose.
Reducing sugars donate electrons to reduce the blue copper(II) ions in Benedict's reagent to brick-red copper(I) oxide precipitate. The intensity of the colour change can be used for semi-quantitative estimation. It's like a chemical 'traffic light' for sugar: blue means no sugar, and green, yellow, orange, or red means increasing amounts of reducing sugar are present.
For Benedict's test, remember to state 'heat in a water bath' and describe the colour change from 'blue to green/yellow/orange/brick-red precipitate' for a positive result. Mentioning 'excess Benedict's reagent' is key for semi-quantitative use.
polysaccharide — A polymer whose subunits are monosaccharides joined together by glycosidic bonds.
Polysaccharides are large, complex carbohydrates formed from many monosaccharide units. They serve as energy stores (starch, glycogen) or structural components (cellulose) and are generally insoluble and unreactive. A polysaccharide is like a long, complex chain made of many identical or similar beads (monosaccharides).
Students often think polysaccharides are sweet like sugars, but actually they are not sugars and do not taste sweet.
glycogen — A polysaccharide made of many glucose molecules linked together, that acts as a glucose store in liver and muscle cells.
Glycogen is the primary energy storage carbohydrate in animals, highly branched like amylopectin but more so, allowing for rapid glucose release when needed. It forms granules in liver and muscle cells. Glycogen is like a highly branched tree, where each leaf is a glucose molecule, allowing for quick access to many leaves (glucose) from many points.
Highlight that glycogen is the 'animal storage carbohydrate' and mention its 'highly branched structure' (1,4 and 1,6 linkages of α-glucose) for efficient glucose release.
cellulose — A polysaccharide made from beta-glucose subunits; used as a strengthening material in plant cell walls.
Cellulose is a structural polysaccharide in plants, formed from β-glucose monomers linked by 1,4 glycosidic bonds, with alternate glucose units inverted. This arrangement allows extensive hydrogen bonding, forming strong microfibrils and fibres. Cellulose is like a strong, interwoven fabric made of many long, parallel threads (cellulose molecules), giving plant cell walls immense strength.
Emphasise that cellulose is a polymer of 'β-glucose' and that the 'alternating 180° rotation' of glucose units enables extensive 'hydrogen bonding' between parallel chains, leading to high 'tensile strength' in plant cell walls.
Carbohydrates are essential biological molecules, ranging from simple monosaccharides like glucose, which serve as immediate energy sources, to complex polysaccharides. Monosaccharides can join together via condensation reactions to form disaccharides, such as maltose or sucrose, linked by glycosidic bonds. Polysaccharides, like starch and glycogen, are large polymers of glucose used for energy storage in plants and animals respectively. Cellulose, another polysaccharide, provides structural support in plant cell walls due to its unique β-glucose linkages and extensive hydrogen bonding.
hydrogen bond — A relatively weak bond formed by the attraction between a group with a small positive charge on a hydrogen atom (Hδ+) and another group carrying a small negative charge (δ−), e.g. between two –Oδ– Hδ+ groups.
Hydrogen bonds are crucial intermolecular forces, weaker than covalent bonds but collectively strong. They are responsible for many unique properties of water and play vital roles in maintaining the structure of large biological molecules like proteins and nucleic acids. Think of hydrogen bonds as weak magnetic attractions between molecules; individually they are easily broken, but many together can create a strong overall force.
Students often think hydrogen bonds are covalent bonds, but actually they are intermolecular forces of attraction, much weaker than the intramolecular covalent bonds that hold atoms together within a molecule.
ester bond / ester linkage — A chemical bond, represented as –COO– , formed when an acid reacts with an alcohol.
Ester bonds are formed via condensation reactions, typically between a carboxyl group of a fatty acid and a hydroxyl group of an alcohol (like glycerol). They are characteristic linkages in lipids, particularly triglycerides. An ester bond is like a chemical 'clasp' that joins a fatty acid 'chain' to a glycerol 'backbone', forming a lipid molecule.
triglyceride — A type of lipid formed when three fatty acid molecules combine with glycerol, an alcohol with three hydroxyl (−OH) groups.
Triglycerides are the most common lipids, serving as efficient energy stores due to their high number of C-H bonds. They are hydrophobic and insoluble in water, also providing insulation and buoyancy. A triglyceride is like a three-pronged fork (glycerol) with three long spaghetti strands (fatty acids) attached to it.
For triglycerides, remember 'three fatty acids' and 'one glycerol' joined by 'ester bonds' via 'condensation reactions'. Emphasize their 'hydrophobic' nature and role as 'energy stores' and 'insulators'.
Lipids are a diverse group of biological molecules, characterized by their insolubility in water. Triglycerides, formed from one glycerol molecule and three fatty acids linked by ester bonds through condensation reactions, are primary energy storage molecules. Their hydrophobic nature also makes them excellent insulators. Phospholipids, with their hydrophilic head and hydrophobic tails, are crucial components of cell membranes, forming a bilayer that regulates substance passage.
peptide bond — The covalent bond joining neighbouring amino acids together in proteins; it is a C–N link between two amino acid molecules, formed by a condensation reaction.
Peptide bonds are the fundamental linkages in proteins, formed between the carboxyl group of one amino acid and the amino group of another, with the release of water. A chain of amino acids linked by peptide bonds is a polypeptide. A peptide bond is like a strong clasp connecting two beads (amino acids) on a necklace (polypeptide chain).
When describing peptide bond formation, specify the reaction between the 'carboxyl group' of one amino acid and the 'amino group' of another, and mention it's a 'condensation reaction' forming a 'C–N link'.
polypeptide — A long chain of amino acids formed by condensation reactions between the individual amino acids; proteins are made of one or more polypeptide chains; see peptide bond.
Polypeptides are linear sequences of amino acids linked by peptide bonds. They are the primary structural component of proteins, which may consist of one or more polypeptide chains folded into a specific 3D shape. A polypeptide is like a string of different coloured beads, where each bead is an amino acid, and the string itself is the chain.
primary structure — The sequence of amino acids in a polypeptide or protein.
The primary structure is the most fundamental level of protein organization, determined by the genetic code. It dictates all subsequent levels of folding and ultimately the protein's final 3D shape and function. The primary structure is like the specific order of letters in a word; changing even one letter can completely change the meaning (function) of the word.
secondary structure — The structure of a protein molecule resulting from the regular coiling or folding of the chain of amino acids (an α-helix or β-pleated sheet).
Secondary structures are local, repeating patterns formed by hydrogen bonding between the backbone atoms (C=O and N-H groups) of amino acids. The two main types are the α-helix and β-pleated sheet. Secondary structure is like the basic patterns you can make with a flexible wire, such as coiling it into a spring (α-helix) or folding it into zig-zags (β-pleated sheet).
Students often think R groups are involved in secondary structure, but actually secondary structure is formed by hydrogen bonds between the 'backbone' atoms (C=O and N-H) of the polypeptide chain, not the R groups.
α-helix — A helical structure formed by a polypeptide chain, held in place by hydrogen bonds; an α-helix is an example of secondary structure in a protein.
In an α-helix, the polypeptide chain coils into a corkscrew shape, stabilized by hydrogen bonds between the oxygen of a C=O group and the hydrogen of an N-H group four amino acids ahead in the chain. All backbone C=O and N-H groups are involved. An α-helix is like a coiled telephone cord, where the coils are held in shape by invisible 'sticky spots' (hydrogen bonds) along the cord.
β-pleated sheet — A loose, sheet-like structure formed by hydrogen bonding between parallel polypeptide chains; a β-pleated sheet is an example of secondary structure in a protein.
In a β-pleated sheet, polypeptide chains lie side-by-side in a zig-zag pattern, forming a sheet-like structure. This is stabilized by hydrogen bonds between C=O and N-H groups of adjacent parallel segments of the polypeptide. A β-pleated sheet is like a folded fan or a corrugated cardboard sheet, where the folds are held in place by hydrogen bonds.
tertiary structure — The compact structure of a protein molecule resulting from the three-dimensional coiling of the chain of amino acids.
Tertiary structure is the overall 3D shape of a single polypeptide chain, formed by interactions between R groups of amino acids. These interactions include hydrogen bonds, disulfide bonds, ionic bonds, and hydrophobic interactions, which stabilize the precise shape. Tertiary structure is like taking a coiled spring (secondary structure) and then bending, twisting, and knotting it into a specific, complex sculpture.
When explaining tertiary structure, list the four types of bonds/interactions (hydrogen, disulfide, ionic, hydrophobic) and state that they occur between 'R groups' to maintain the 'precise 3D shape'.
quaternary structure — The three-dimensional arrangement of two or more polypeptides, or of a polypeptide and a non-protein component such as haem, in a protein molecule.
Quaternary structure applies to proteins composed of multiple polypeptide chains (subunits) or those with non-protein prosthetic groups. It describes how these subunits are arranged and interact to form the complete functional protein. Quaternary structure is like several individual sculptures (tertiary structures of polypeptide chains) coming together and arranging themselves into a larger, functional art installation.
haemoglobin — The red pigment found in red blood cells, whose molecules contain four iron atoms within a globular protein made up of four polypeptides; it combines reversibly with oxygen.
Haemoglobin is a globular protein with a quaternary structure, consisting of two α-globin and two β-globin polypeptide chains, each associated with a haem prosthetic group containing an iron atom. It is responsible for oxygen transport in the blood. Haemoglobin is like a tiny oxygen taxi, with four seats (haem groups) that can each pick up and drop off one oxygen molecule.
When describing haemoglobin, mention its 'globular' nature, 'quaternary structure' (four polypeptide chains), and the presence of 'four haem groups' each with an 'iron atom' for 'reversible oxygen binding'.
globular protein — A protein whose molecules are folded into a relatively spherical shape, often has physiological roles and is often water-soluble and metabolically active, e.g. insulin, haemoglobin and enzymes.
Globular proteins typically have hydrophobic R groups folded into the interior and hydrophilic R groups on the exterior, making them soluble in water. Their precise 3D shape is critical for their diverse physiological functions, such as catalysis (enzymes) or transport (haemoglobin). A globular protein is like a tightly wound ball of yarn, with the 'water-hating' parts tucked inside and the 'water-loving' parts on the surface, allowing it to dissolve in water.
sickle cell anaemia — A genetic disease caused by a faulty gene coding for haemoglobin, in which haemoglobin tends to precipitate when oxygen concentrations are low.
This condition results from a single amino acid substitution (valine for glutamic acid) on the surface of the β-chain of haemoglobin. This makes haemoglobin less soluble, causing red blood cells to become sickle-shaped, leading to blockages and reduced oxygen transport. It's like a tiny, crucial bolt on a machine being replaced with a slightly wrong one, causing the whole machine (red blood cell) to malfunction and change shape under stress.
collagen — The main structural protein of animals; known as ‘white fibres’, the fundamental unit of the fibre consists of three helical polypeptide chains wound around each other, forming a ‘triple helix’ with high tensile strength.
Collagen is an insoluble fibrous protein, highly abundant in animals, providing strength and flexibility to tissues like skin, tendons, and bones. Its triple helix structure, stabilized by hydrogen and covalent bonds, and staggered arrangement of fibrils contribute to its immense tensile strength. Collagen is like a super-strong, braided rope made of three smaller ropes (polypeptide helices), which are then woven together with other ropes to form an even stronger cable.
fibrous protein — A protein whose molecules have a relatively long, thin structure that is generally insoluble and metabolically inactive, and whose function is usually structural, e.g. keratin and collagen.
Fibrous proteins are elongated and often form strong, insoluble fibres, making them ideal for structural roles in organisms. Their insolubility arises from having many hydrophobic R groups exposed on their surface. A fibrous protein is like a long, strong piece of string or a hair strand, designed for strength and support rather than for dissolving or reacting quickly.
biuret test — A test for the presence of amine groups and thus for the presence of protein; biuret reagent is added to the unknown substance, and a change from pale blue to purple indicates the presence of protein.
The biuret test detects peptide bonds (which contain nitrogen atoms in amine groups). Copper(II) ions in the alkaline biuret reagent form a purple complex with these bonds. No heating is required. It's like a chemical 'purple alert' for protein; if the solution turns purple, protein is present.
Students often think the biuret test detects amino acids, but actually it specifically detects the 'peptide bonds' between amino acids, meaning it tests for polypeptides or proteins.
Proteins are polymers of amino acids linked by peptide bonds, formed through condensation reactions. Their function is intimately tied to their complex three-dimensional structure, which is described at four levels. The primary structure is the unique sequence of amino acids. This sequence dictates the secondary structure, which involves regular coiling (α-helix) or folding (β-pleated sheet) stabilized by hydrogen bonds in the polypeptide backbone. The tertiary structure is the overall 3D shape of a single polypeptide, maintained by interactions between R groups, including hydrogen, ionic, disulfide bonds, and hydrophobic interactions. Some proteins, like haemoglobin, exhibit quaternary structure, involving the arrangement of multiple polypeptide chains or non-protein components.
Water is an indispensable biological molecule, crucial for life due to its unique properties. Its polarity, arising from hydrogen bonding, makes it an excellent solvent, allowing many substances to dissolve and be transported within organisms. Water also possesses a high specific heat capacity, meaning it can absorb or release large amounts of heat with only a small change in its own temperature, helping to stabilize internal body temperatures. Furthermore, its high latent heat of vaporisation allows organisms to cool effectively through evaporation.
When comparing molecules (e.g., starch vs. cellulose), always link their structural differences (e.g., α/β-glucose, branching) directly to their functional differences (e.g., energy storage vs. structural support).
Practice drawing and labelling: α-glucose, a triglyceride (showing ester bonds), and the general structure of an amino acid. Be able to show how a peptide or glycosidic bond is formed.
For proteins like haemoglobin and collagen, explicitly relate their structure (globular vs. fibrous, quaternary structure, prosthetic groups) to their specific biological role.
Definitions Bank
macromolecule
A large molecule such as a polysaccharide, protein or nucleic acid.
polymer
A giant molecule made from many similar repeating subunits joined together in a chain; the subunits are much smaller and simpler molecules known as monomers; examples of biological polymers are polysaccharides, proteins and nucleic acids.
monomer
A relatively simple molecule which is used as a basic building block for the synthesis of a polymer; many monomers are joined together by covalent bonds to make the polymer, usually by condensation reactions; common examples of monomers are monosaccharides, amino acids and nucleotides.
condensation reaction
A chemical reaction involving the joining together of two molecules by removal of a water molecule.
hydrolysis
A chemical reaction in which a chemical bond is broken by the addition of a water molecule; commonly used to break down complex molecules into simpler molecules.
+24 more definitions
View all →Command Word Guide
| Describe | For structures, describe specific features (e.g., 'alpha-glucose with hydroxyl group on carbon 1 below the ring'). For tests, describe the reagent, conditions, and exact colour changes (e.g., 'Benedict's reagent, heat in water bath, blue to brick-red precipitate'). |
| Explain | Provide reasons and mechanisms. For example, 'Explain how cellulose provides strength' requires linking β-glucose, 180° rotation, hydrogen bonding, microfibrils, and tensile strength. For water properties, link hydrogen bonding to specific heat capacity or latent heat of vaporisation. |
| Compare | State both similarities and differences, often linking structure to function. For example, comparing starch and cellulose requires mentioning α vs β glucose, branching vs linear, and energy storage vs structural roles. |
| Suggest | Apply knowledge to a novel situation. For example, 'Suggest why a particular molecule is soluble' would require referring to polar groups and hydrogen bonding with water. |
Common Mistakes
Thinking all large molecules are macromolecules.
Macromolecules specifically refer to polymers like polysaccharides, proteins, and nucleic acids, which are built from repeating monomer units.
Believing all disaccharides are reducing sugars.
Sucrose is a common non-reducing sugar and requires prior hydrolysis to be detected by Benedict's test.
Thinking polysaccharides are sweet like sugars.
Polysaccharides are not sugars and do not taste sweet; they are large, complex carbohydrates.
+3 more
View all →Enzymes are biological catalysts, typically globular proteins, that accelerate reactions by lowering activation energy. Their activity is highly specific due to their active sites and is influenced by various environmental factors. Understanding enzyme kinetics and inhibition is crucial for comprehending their roles in biological systems and industrial applications.
enzyme — a protein produced by a living organism that acts as a biological catalyst in a chemical reaction by reducing activation energy
Enzymes are essential for life, catalysing almost all metabolic reactions. They are globular proteins with precise 3D shapes, and their names often end in '-ase'. Think of an enzyme as a specific tool, like a key, that perfectly fits and operates on a particular lock (the substrate) to perform a task (the reaction) much faster than it would otherwise happen.
active site — an area on an enzyme molecule where the substrate can bind
The active site has a specific shape complementary to the substrate, allowing temporary bonds to form. This binding is crucial for the enzyme's catalytic activity. Imagine the active site as a glove designed to fit only one specific hand (the substrate). Only when the hand fits perfectly can the glove perform its function.
activation energy — the energy that must be provided to make a reaction take place; enzymes reduce the activation energy required for a substrate to change into a product
Without enzymes, many biological reactions would require high temperatures to proceed, which is incompatible with life. Enzymes lower this energy barrier by holding substrates in a way that facilitates reaction. Imagine pushing a ball over a hill. The activation energy is the effort needed to get the ball to the top. An enzyme is like digging a tunnel through the hill, making it much easier to get the ball to the other side.
Students often think enzymes are used up in reactions, but actually they remain unchanged and can be reused after converting substrate to product.
When asked to define 'enzyme', ensure you include 'protein', 'biological catalyst', and 'reduces activation energy' for full marks.
Enzymes function by binding to specific substrate molecules at their active site, forming an enzyme-substrate complex. This binding facilitates the conversion of substrate into product. The enzyme then releases the product and is regenerated, ready to catalyse another reaction. This process significantly lowers the activation energy required for the reaction to proceed.
lock-and-key hypothesis — a hypothesis for enzyme action; the substrate is a complementary shape to the active site of the enzyme, and fits exactly into the site; the enzyme shows specificity for the substrate
This hypothesis explains enzyme specificity, where each enzyme acts on only one type of substrate due to the precise fit. Temporary bonds form between the substrate and R groups in the active site. Just as a specific key fits only one lock, a specific substrate fits only one enzyme's active site.
induced-fit hypothesis — a hypothesis for enzyme action; the substrate is a complementary shape to the active site of the enzyme, but not an exact fit – the enzyme, or sometimes the substrate, can change shape slightly to ensure a perfect fit, but it is still described as showing specificity
This modern hypothesis refines the lock-and-key model by suggesting flexibility in the enzyme and/or substrate. This slight shape change optimizes the fit, making catalysis even more efficient. Instead of a rigid lock and key, think of a glove (enzyme) that slightly molds to the hand (substrate) as it's put on, ensuring a snug and effective fit.
Students often think the active site is rigid, but actually the induced-fit hypothesis suggests it can change shape slightly upon substrate binding for a better fit.
When describing enzyme action, always refer to the 'active site' and its complementary shape to the substrate.
The progress of enzyme-controlled reactions can be monitored by measuring either the rate of product formation or the rate of substrate disappearance. This allows for the determination of initial reaction rates, which are crucial for studying the effects of various factors on enzyme activity. A colorimeter is often used for quantitative analysis.
colorimeter — an instrument that measures the colour of a solution by measuring the absorption of different wavelengths of light
Colorimeters provide quantitative data for reactions involving colour changes, allowing for precise measurement of product formation or substrate disappearance over time. Greater absorption indicates higher concentration of the coloured substance. Think of a colorimeter as a sophisticated eye that can precisely quantify how much light of a specific colour is being blocked by a solution, telling you how much of a coloured substance is present.
When describing colorimeter use, specify that it measures 'absorption' and how this relates to 'concentration' of the substance causing the colour.
Enzyme activity is highly sensitive to environmental conditions. Key factors include temperature, pH, enzyme concentration, and substrate concentration. Each of these factors can significantly influence the rate of an enzyme-catalysed reaction, often exhibiting an optimum point beyond which activity decreases.
When explaining temperature effects, always mention both the increase in kinetic energy (more frequent collisions) up to the optimum, and the breaking of bonds causing denaturation beyond the optimum.
When explaining pH effects, refer specifically to the alteration of charges on amino acid R-groups in the active site, which disrupts bonds and changes the tertiary structure.
The efficiency of an enzyme can be quantified using kinetic parameters such as Vmax and Km. Vmax represents the maximum reaction rate when all active sites are saturated with substrate. Km, the Michaelis-Menten constant, indicates the substrate concentration at which the enzyme works at half its maximum rate, providing a measure of substrate affinity.
Vmax — the theoretical maximum rate of an enzyme-controlled reaction, obtained when all the active sites of the enzyme are occupied
At Vmax, the enzyme is saturated with substrate, meaning all active sites are continuously busy converting substrate to product. Increasing substrate concentration further will not increase the reaction rate. Imagine a factory with a fixed number of machines. Vmax is the maximum output when all machines are running at full capacity, with a constant supply of raw materials.
Michaelis–Menten constant (Km) — the substrate concentration at which an enzyme works at half its maximum rate (½Vmax), used as a measure of the efficiency of an enzyme; the lower the value of Km, the more efficient the enzyme
Km is an indicator of an enzyme's affinity for its substrate. A low Km means the enzyme achieves half its maximum rate at a low substrate concentration, indicating high affinity and efficiency. If Vmax is how fast a taxi driver can drive, Km is how easily they find passengers. A low Km means they find passengers quickly even in a quiet area, indicating high efficiency.
Students often think a high Km means high affinity, but actually a lower Km indicates a higher affinity of the enzyme for its substrate.
Relate Km directly to 'affinity' and 'efficiency'; a lower Km signifies higher affinity and thus a more efficient enzyme.
Enzyme activity can be reduced by inhibitors, which are substances that interfere with the enzyme's function. These can be classified as reversible competitive or non-competitive inhibitors, each with distinct mechanisms of action and effects on reaction kinetics. Understanding inhibition is vital for drug development and metabolic regulation.
competitive inhibition — when a substance reduces the rate of activity of an enzyme by competing with the substrate molecules for the enzyme’s active site; increasing substrate concentration reduces the degree of inhibition; increasing inhibitor concentration increases the degree of inhibition
Competitive inhibitors are structurally similar to the substrate and bind reversibly to the active site. The effect can be overcome by increasing substrate concentration, which outcompetes the inhibitor. Imagine two different keys that can both fit into the same lock, but only one (the substrate) can open it. The other key (the inhibitor) just blocks the lock temporarily.
non-competitive inhibition — when a substance reduces the rate of activity of an enzyme, but increasing the concentration of the substrate does not reduce the degree of inhibition; many non-competitive inhibitors bind to areas of the enzyme molecule other than the active site itself
Non-competitive inhibitors bind to an allosteric site, causing a conformational change in the enzyme that distorts the active site, making it less effective or unable to bind substrate. This inhibition cannot be overcome by increasing substrate concentration. Imagine a machine (enzyme) that needs a specific part (substrate) to work. A non-competitive inhibitor is like someone bending a crucial lever on the machine, making it unable to function properly, regardless of how many parts you feed it.
Students often confuse non-competitive inhibition with competitive inhibition, but actually the key difference is that non-competitive inhibition is not overcome by increasing substrate concentration.
To distinguish inhibitors, remember: competitive inhibition can be overcome by increasing substrate concentration (Vmax is unchanged, Km increases), while non-competitive cannot (Vmax is lowered, Km is unchanged).
Immobilised enzymes are enzymes that have been fixed to a surface or trapped within inert materials, such as alginate beads. This technique offers significant commercial advantages, including the ability to reuse enzymes, ensure product purity by preventing enzyme contamination, and often enhance enzyme stability against adverse conditions like temperature and pH fluctuations.
immobilised enzymes — enzymes that have been fixed to a surface or trapped inside beads of agar gel
Immobilised enzymes are used commercially to allow for enzyme reuse, prevent product contamination, and often increase enzyme stability against temperature and pH changes. They are commonly trapped in alginate beads. Think of a chef who wants to reuse a special cooking tool without it getting mixed into the food. Immobilising the enzyme is like attaching the tool to a fixed stand, so it can be used repeatedly and the food remains pure.
When asked for advantages of immobilised enzymes, focus on 'reusability', 'product purity (enzyme-free)', and 'increased stability to temperature/pH'.
In practical investigation questions, always state that the 'initial rate' of reaction should be measured, as substrate concentration is not yet a limiting factor.
Definitions Bank
enzyme
a protein produced by a living organism that acts as a biological catalyst in a chemical reaction by reducing activation energy
active site
an area on an enzyme molecule where the substrate can bind
lock-and-key hypothesis
a hypothesis for enzyme action; the substrate is a complementary shape to the active site of the enzyme, and fits exactly into the site; the enzyme shows specificity for the substrate
induced-fit hypothesis
a hypothesis for enzyme action; the substrate is a complementary shape to the active site of the enzyme, but not an exact fit – the enzyme, or sometimes the substrate, can change shape slightly to ensure a perfect fit, but it is still described as showing specificity
activation energy
the energy that must be provided to make a reaction take place; enzymes reduce the activation energy required for a substrate to change into a product
+6 more definitions
View all →Command Word Guide
| Define | Provide the precise, mark-scheme definition for terms like 'enzyme', 'active site', 'activation energy', 'Vmax', and 'Km', ensuring all key components are included. |
| Explain | For enzyme action, explain the induced-fit hypothesis, mentioning the slight change in shape for a perfect fit. For factors affecting activity, explain the underlying molecular reasons (e.g., kinetic energy, denaturation, R-group charges). For inhibition, explain the mechanism of binding and its effect on the active site. |
| Describe | Describe the process of investigating enzyme reactions, including how to use a colorimeter and what measurements are taken (e.g., initial rate, product formation/substrate disappearance). |
| Compare | When comparing lock-and-key and induced-fit, highlight the flexibility aspect of the latter. When comparing competitive and non-competitive inhibition, focus on the binding site and whether increasing substrate concentration overcomes the inhibition. |
+1 more
View all →Common Mistakes
Thinking enzymes are used up during a reaction.
Enzymes are biological catalysts that remain unchanged and are regenerated after converting substrate to product, allowing them to be reused.
Believing the active site is a rigid structure.
The induced-fit hypothesis suggests the active site is flexible and can change shape slightly upon substrate binding to achieve a more perfect fit.
Assuming enzymes provide energy for reactions.
Enzymes do not provide energy; instead, they lower the activation energy barrier required for a reaction to start, thereby speeding it up.
+3 more
View all →This chapter explores the structure and function of cell membranes, focusing on the fluid mosaic model and the roles of its components. It details various transport mechanisms across membranes, including passive and active processes, and introduces cell signalling as a crucial communication method.
fluid mosaic model — The currently accepted model of membrane structure, proposed by Singer and Nicolson in 1972, in which protein molecules are free to move about in a fluid bilayer of phospholipid molecules.
This model describes the cell membrane as a dynamic structure where phospholipids form a fluid bilayer, allowing proteins to move laterally within it, creating a mosaic-like pattern. This fluidity is crucial for membrane function, including cell growth, movement, and signalling. Imagine a sea of olive oil (phospholipids) with icebergs (proteins) floating and moving within it, some anchored, others drifting freely.
Students often think the membrane is a rigid, static structure, but actually it is fluid and dynamic, allowing components to move.
When describing the fluid mosaic model, ensure you mention both 'fluid' (phospholipid and protein movement) and 'mosaic' (scattered protein pattern) aspects for full marks.
cholesterol — A small, lipid-related molecule with a hydrophilic head and a hydrophobic tail which is an essential constituent of membranes; it is particularly common in animal cells and gives flexibility and stability to the membrane as well as reducing fluidity.
Cholesterol molecules insert between phospholipids, reducing membrane fluidity at higher temperatures and preventing close packing at lower temperatures, thus maintaining optimal membrane consistency. Its hydrophobic regions also help prevent the passage of ions and polar molecules. Think of cholesterol as the 'temperature regulator' and 'stabilizer' of the membrane, like adding a thickener to a sauce to control its consistency.
Students often think cholesterol only makes membranes more rigid, but actually it also prevents phospholipids from packing too closely at low temperatures, maintaining fluidity.
Remember to state both the 'stability' and 'fluidity regulation' roles of cholesterol, especially its importance in animal cells and its absence in prokaryotes.
Cell surface membranes are composed of phospholipids, cholesterol, and proteins, along with glycolipids and glycoproteins. Phospholipids form the basic bilayer structure, providing a barrier. Cholesterol regulates membrane fluidity and stability. Proteins embedded within or associated with the membrane perform diverse functions, including transport, cell signalling, and enzymatic activity. Glycolipids and glycoproteins are involved in cell-to-cell recognition.
cell signalling — The molecular mechanisms by which cells detect and respond to external stimuli, including communication between cells.
Cell signalling involves a ligand binding to a specific receptor, triggering a cascade of events inside the cell, often involving second messengers, to bring about a cellular response. This process allows cells to coordinate activities and respond to their environment. It's like a secret message system: a messenger (ligand) delivers a coded note to a specific mailbox (receptor) on a house (cell), which then triggers a series of actions inside the house.
Students often think signalling only involves direct contact, but actually it frequently involves chemical messengers transported over distances.
ligand — A biological molecule which binds specifically to another molecule, such as a cell surface membrane receptor, during cell signalling.
Ligands are the 'messenger molecules' in cell signalling, initiating a response by binding to a complementary receptor protein. This binding causes a change in the receptor's shape, transmitting the signal into the cell. A ligand is like a specific key that fits into a particular lock (the receptor) to open a door (initiate a cellular response).
Emphasize the 'specificity' of ligand-receptor binding when explaining its role in cell signalling.
transduction — Occurs during cell signalling and is the process of converting a signal from one method of transmission to another.
After a ligand binds to a receptor, the external signal is converted into an intracellular message, often involving a series of molecular changes or the production of second messengers. This allows the signal to be relayed and amplified within the cell. It's like a translator converting a message from one language (external signal) into another (internal cellular signal) so the cell can understand and act on it.
When outlining cell signalling, ensure you include the stages: stimulus, ligand secretion, transport, binding to receptor, transduction, and cellular response, mentioning amplification.
Substances move across cell membranes through various mechanisms, categorized as passive or active. Passive transport, including diffusion, facilitated diffusion, and osmosis, does not require metabolic energy and moves substances down their concentration or water potential gradients. Active transport and bulk transport, however, require energy (ATP) to move substances against gradients or to transport large quantities of material.
diffusion — The net movement of molecules or ions from a region of higher concentration to a region of lower concentration down a concentration gradient, as a result of the random movements of particles.
Diffusion is a passive process driven by the kinetic energy of molecules, leading to an even distribution of substances over time. It is effective over short distances and is how small, non-polar molecules like oxygen and carbon dioxide cross cell membranes. Imagine opening a bottle of perfume in a room; the scent molecules will gradually spread out until they are evenly distributed throughout the room.
Always include 'net movement' and 'down a concentration gradient' in your definition of diffusion for accuracy.
facilitated diffusion — The diffusion of a substance through a transport protein (channel protein or carrier protein) in a cell membrane; the protein provides hydrophilic areas that allow the molecule or ion to pass through the membrane, which would otherwise be less permeable to it.
This is a passive process that allows larger polar molecules and ions, which cannot easily cross the hydrophobic lipid bilayer, to move down their concentration gradient with the help of specific membrane proteins. It does not require metabolic energy. It's like having a special gate or a revolving door (transport protein) in a wall (membrane) that only allows specific people (molecules) to pass through, but they still move from a crowded side to a less crowded side.
Students often confuse facilitated diffusion with active transport, but actually facilitated diffusion is passive and moves substances down a concentration gradient, while active transport is active and moves against it.
channel protein — A membrane protein of fixed shape which has a water-filled pore through which selected hydrophilic ions or molecules can pass by facilitating diffusion or active transport.
Channel proteins provide a hydrophilic pathway for charged or polar substances to cross the membrane. Many are 'gated', meaning they can open or close to control the passage of ions, playing a crucial role in nerve impulses. Think of a channel protein as a tunnel through a mountain (the membrane) that only allows specific types of vehicles (ions/molecules) to pass through, sometimes with a gatekeeper controlling access.
carrier protein — A membrane protein which changes shape to allow the passage into or out of the cell of specific ions or molecules by facilitated diffusion or active transport.
Carrier proteins bind to specific molecules or ions and then undergo a conformational change to transport them across the membrane. They can be involved in both passive (facilitated diffusion) and active transport (pumps). Imagine a revolving door or a shuttle bus (carrier protein) that picks up a specific passenger (molecule) on one side of a building (membrane) and drops them off on the other side by changing its orientation.
Emphasize the 'change in shape' as a key characteristic distinguishing carrier proteins from channel proteins.
osmosis — The net diffusion of water molecules from a region of higher water potential to a region of lower water potential, through a partially permeable membrane.
Osmosis is a specific type of diffusion for water, crucial for maintaining cell volume and turgor in living organisms. It is a passive process that does not require metabolic energy. It's like water trying to 'dilute' a concentrated solution by moving across a barrier that only lets water through, until the concentration is equal on both sides.
Students often think osmosis is the movement of solute, but actually it is specifically the net movement of water molecules.
Always include 'net diffusion of water molecules', 'partially permeable membrane', and 'down a water potential gradient' in your definition of osmosis.
water potential — A measure of the tendency of water to move from one place to another; water moves from a solution with higher water potential to one with lower water potential; water potential is decreased by the addition of solute, and increased by the application of pressure; the symbol for water potential is ψ or ψw.
Water potential quantifies the 'free energy' of water, indicating its tendency to move. Pure water has the highest water potential (0 kPa), and adding solutes lowers it (makes it more negative). Pressure increases water potential. Think of water potential as the 'pressure' or 'desire' of water to move. Water always 'wants' to move from where it has more 'desire' (higher potential) to where it has less 'desire' (lower potential).
Students often confuse higher water potential with a more negative value, but actually a higher water potential means a less negative value (closer to 0 kPa).
Remember that pure water has a water potential of 0 kPa, and all solutions have negative water potentials. A less negative value indicates a higher water potential.
The effects of osmosis differ between animal and plant cells due to the presence of a cell wall in plants. Animal cells, lacking a cell wall, can swell and burst (lyse) in hypotonic solutions or shrink (crenate) in hypertonic solutions. Plant cells, however, become turgid in hypotonic solutions as the cell wall prevents bursting, and undergo plasmolysis in hypertonic solutions where the protoplast shrinks away from the cell wall.
protoplast — The living contents of a plant cell, including the cell surface membrane but excluding the cell wall.
The protoplast is the functional, living part of a plant cell that undergoes osmotic changes. In plasmolysis, it is the protoplast that shrinks away from the cell wall. If a plant cell is a house with a strong brick wall, the protoplast is everything inside the house, including the inner skin (cell surface membrane) and all its contents.
plasmolysis — The loss of water from a plant or prokaryote cell to the point where the protoplast shrinks away from the cell wall.
Plasmolysis occurs when a plant cell is placed in a solution with a lower water potential, causing water to leave the cell by osmosis. The shrinking protoplast pulls away from the rigid cell wall, leading to loss of turgor. Imagine a deflated balloon (protoplast) inside a rigid box (cell wall); as the balloon loses air, it pulls away from the box's sides.
Students often think plasmolysis means the cell bursts, but actually it means the protoplast shrinks away from the cell wall due to water loss.
incipient plasmolysis — The point at which plasmolysis is about to occur when a plant cell or a prokaryote cell is losing water; at this point the protoplast is exerting no pressure on the cell wall.
This is the critical stage where the cell has lost enough water that its protoplast is no longer pressing against the cell wall, but has not yet visibly pulled away. It represents the transition from turgid to plasmolysed. It's like the moment a balloon inside a box has just enough air that it's touching all sides, but if you let out even a tiny bit more air, it will start to pull away.
active transport — The movement of molecules or ions through transport proteins across a cell membrane, against their concentration gradient, using energy from ATP.
Active transport allows cells to accumulate substances or remove waste products, maintaining specific internal concentrations different from the external environment. It is an energy-consuming process, typically powered by ATP hydrolysis. It's like pushing a ball uphill (against the concentration gradient) which requires energy, unlike letting it roll downhill (diffusion).
Key elements for active transport are 'against concentration gradient', 'uses carrier proteins (pumps)', and 'requires ATP/energy'.
sodium–potassium pump (Na+–K+ pump) — A membrane protein (or proteins) that moves sodium ions out of a cell and potassium ions into it, using ATP.
This specific carrier protein is vital in animal cells for maintaining ion gradients across the cell membrane, crucial for nerve impulse transmission and osmotic balance. It pumps three Na+ out and two K+ in for each ATP molecule used, creating a potential difference. Imagine a bouncer at a club who lets two specific people (K+) in for every three specific people (Na+) he pushes out, and this requires constant effort (ATP).
endocytosis — The bulk movement of liquids (pinocytosis) or solids (phagocytosis) into a cell, by the infolding of the cell surface membrane to form vesicles containing the substance; endocytosis is an active process requiring ATP.
This process allows cells to take in large molecules, particles, or even other cells that are too big to pass through membrane proteins. It involves the formation of a vesicle from the cell surface membrane and requires energy. It's like a cell 'eating' or 'drinking' by wrapping its membrane around a large item and pulling it inside in a bubble.
exocytosis — The bulk movement of liquids or solids out of a cell, by the fusion of vesicles containing the substance with the cell surface membrane; exocytosis is an active process requiring ATP.
Exocytosis is the reverse of endocytosis, used by cells to secrete substances like hormones or enzymes, or to remove waste products. Vesicles containing the material fuse with the cell surface membrane, releasing their contents outside. It's like a cell 'spitting out' or 'secreting' substances by having a bubble containing the material merge with its outer skin and release the contents.
phagocyte — A type of cell that ingests (eats) and destroys pathogens or damaged body cells by the process of phagocytosis; some phagocytes are white blood cells.
Phagocytes are specialized cells, part of the immune system, that protect the body by engulfing harmful foreign particles, bacteria, and dead or dying cells. This process is a form of endocytosis called phagocytosis. Think of phagocytes as the 'clean-up crew' or 'pac-men' of the body, constantly engulfing and digesting unwanted invaders or debris.
Be ready to compare transport mechanisms. Use a table to revise the differences between simple diffusion, facilitated diffusion, and active transport (Gradient? Protein needed? ATP needed? Specificity?).
Always link a larger surface area to volume ratio with a faster rate of diffusion in application questions.
Definitions Bank
fluid mosaic model
The currently accepted model of membrane structure, proposed by Singer and Nicolson in 1972, in which protein molecules are free to move about in a fluid bilayer of phospholipid molecules.
cholesterol
A small, lipid-related molecule with a hydrophilic head and a hydrophobic tail which is an essential constituent of membranes; it is particularly common in animal cells and gives flexibility and stability to the membrane as well as reducing fluidity.
cell signalling
The molecular mechanisms by which cells detect and respond to external stimuli, including communication between cells.
ligand
A biological molecule which binds specifically to another molecule, such as a cell surface membrane receptor, during cell signalling.
transduction
Occurs during cell signalling and is the process of converting a signal from one method of transmission to another.
+14 more definitions
View all →Command Word Guide
| Describe | For the fluid mosaic model, describe both the 'fluid' nature (phospholipid and protein movement) and 'mosaic' pattern (scattered proteins). For transport mechanisms, describe the direction of movement, energy requirement, and any proteins involved. |
| Explain | For osmosis, explain the movement of water in terms of water potential gradients and the role of the partially permeable membrane. For cholesterol, explain how it regulates fluidity at both high and low temperatures. For cell signalling, explain the sequence of events from ligand binding to cellular response. |
| Compare | When comparing transport mechanisms (e.g., facilitated diffusion vs. active transport), clearly state similarities and differences regarding concentration gradient, energy requirement, and protein involvement. |
| Illustrate | When illustrating the principle of surface area to volume ratios, explain how increasing size leads to a decrease in the ratio and its implications for transport efficiency. |
Common Mistakes
Thinking the cell membrane is a rigid, static structure.
The cell membrane is fluid and dynamic, with components able to move laterally.
Believing cholesterol only makes membranes more rigid.
Cholesterol also prevents phospholipids from packing too closely at low temperatures, maintaining fluidity.
Confusing facilitated diffusion with active transport.
Facilitated diffusion is passive (no ATP) and moves substances down a concentration gradient, while active transport is active (uses ATP) and moves against it.
+3 more
View all →This chapter explores the mitotic cell cycle, a precisely regulated series of events enabling body cells to grow and divide. It details chromosome behaviour during mitosis, highlighting its importance for growth, repair, and asexual reproduction. The chapter also covers the protective role of telomeres, the significance of stem cells, and how uncontrolled cell division can lead to cancer.
cell cycle — The sequence of events that takes place from one cell division until the next; it is made up of interphase, mitosis and cytokinesis.
The cell cycle is a precisely controlled series of events that allows cells to grow, replicate their DNA, and divide. It ensures that new cells are produced accurately and efficiently, maintaining tissue integrity and organismal growth, much like a factory production line gathers raw materials, duplicates blueprints, performs quality checks, and then assembles identical products.
Students often think interphase is a 'resting phase', but actually it is a period of intense growth and metabolic activity, including DNA replication.
chromatid — One of two identical parts of a chromosome, held together by a centromere, formed during interphase by the replication of the DNA strand.
During the S phase of interphase, DNA replicates, resulting in each chromosome consisting of two sister chromatids. These identical structures ensure that when the cell divides, each daughter cell receives an exact copy of the genetic material, similar to each side of a copied zipper joined at the pull-tab.
Students often think a chromatid is a whole chromosome, but actually a chromosome before cell division consists of two sister chromatids.
mitosis — The division of a nucleus into two so that the two daughter cells have exactly the same number and type of chromosomes as the parent cell.
Mitosis is a crucial process for growth, repair, and asexual reproduction in multicellular organisms. It ensures genetic continuity by producing two genetically identical daughter nuclei from a single parent nucleus, acting like a photocopier for cells to make identical copies of genetic material.
Students often think mitosis is the entire cell division process, but actually mitosis refers specifically to nuclear division, which is then followed by cytokinesis (cell division).
The cell cycle is a precisely controlled sequence of events by which body cells grow and divide. It consists of two main phases: Interphase and the Mitotic (M) phase. Interphase is a period of intense growth and metabolic activity, including DNA replication, and is subdivided into G1, S, and G2 phases. The M phase includes mitosis, the nuclear division, followed by cytokinesis, the division of the cytoplasm.
Be able to outline the events in each phase (G1, S, G2, M) and their relative durations, as this is frequently tested.
Before cell division, each chromosome consists of two identical sister chromatids joined at a centromere. During mitosis, these chromosomes undergo a series of organised movements. The nuclear envelope breaks down, and a spindle forms. Microtubules attach to kinetochores, protein structures at the centromeres, to facilitate the precise segregation of chromatids to daughter cells.
kinetochore — A protein structure found at the centromere of a chromatid to which microtubules attach during nuclear division.
Kinetochores are essential for the accurate segregation of sister chromatids during anaphase. They act as the attachment points for spindle microtubules, which pull the chromatids towards opposite poles of the cell, much like a coupling mechanism connects a train car to its engine.
When asked to describe chromosome structure before division, ensure you mention 'two identical chromatids joined by a centromere'.
When describing anaphase, mention the role of kinetochores in microtubule attachment and the pulling of chromatids towards the poles.
Mitosis is divided into four main stages: Prophase, Metaphase, Anaphase, and Telophase (PMAT). During prophase, chromosomes condense and become visible. In metaphase, chromosomes align at the metaphase plate. Anaphase involves the separation of sister chromatids, which are pulled to opposite poles. Finally, in telophase, new nuclear envelopes form around the separated chromosomes, and the cell begins to divide.
When asked to describe the stages of mitosis, focus on the behaviour and arrangement of the chromosomes at each stage.
Mitosis is fundamental for several biological processes. It is essential for the growth of multicellular organisms, allowing for an increase in cell number. It also plays a crucial role in the repair of damaged tissues by replacing dead or injured cells. Furthermore, mitosis is the basis of asexual reproduction, producing genetically identical offspring from a single parent.
asexual reproduction — The production of new individuals of a species by a single parent organism.
Asexual reproduction relies on mitosis to produce offspring that are genetically identical to the parent. This method is common in unicellular organisms and many plants, allowing for rapid population growth in stable environments, similar to making a clone of a plant by taking a cutting.
When explaining the importance of mitosis, link it directly to growth, repair, asexual reproduction, and immune response, as these are common mark points.
telomere — Repetitive sequence of DNA at the end of a chromosome that protects genes from the chromosome shortening that happens at each cell division.
Telomeres are crucial because DNA replication enzymes cannot fully copy the very ends of DNA strands, leading to shortening with each division. Telomeres, being non-coding, absorb this shortening, protecting vital genetic information from being lost, much like the plastic tips on shoelaces protect the main lace from fraying.
Explain that telomeres prevent gene loss during DNA replication because the copying enzyme cannot replicate the very end of the DNA strand.
Students often think telomeres contain genetic information, but actually they are made of short, repeated base sequences that have no useful information, serving purely a protective role.
stem cell — A relatively unspecialised cell that retains the ability to divide an unlimited number of times, and which has the potential to become a specialised cell (such as a blood cell or muscle cell).
Stem cells are vital for growth, development, and tissue repair. Their ability to self-renew and differentiate into various cell types makes them crucial for replacing damaged or dead cells throughout an organism's life, acting like blank canvases that can be painted into any type of specialised cell.
Students often think all stem cells can become any cell type, but actually their potency varies (totipotent, pluripotent, multipotent).
Distinguish between totipotent, pluripotent, and multipotent stem cells, providing examples for each type if possible.
The precise control of the cell cycle is vital. A breakdown in these control mechanisms can lead to uncontrolled cell division, a hallmark of cancers. This often results from mutations in genes that regulate the cell cycle, causing cells to divide uncontrollably and form tumours. These tumours can then spread to other parts of the body through a process called metastasis.
cancers — A group of diseases that result from a breakdown in the usual control mechanisms that regulate cell division; certain cells divide uncontrollably and form tumours, from which cells may break away and form secondary tumours in other areas of the body (metastasis).
Cancers arise from uncontrolled mitosis, often initiated by mutations in genes that regulate the cell cycle. This leads to the formation of tumours, which can interfere with normal tissue function and spread throughout the body, much like rogue cars ignoring traffic signals and crashing into others.
mutation — A random change in the base sequence (structure) of DNA (a gene mutation), or in the structure and/or number of chromosomes (a chromosome mutation).
Mutations are the underlying cause of many diseases, including cancer, as they can alter the function of genes that control critical cellular processes like cell division. While most mutations are harmless or lead to cell death, some can lead to uncontrolled growth, similar to a typo in a recipe that causes an endless, harmful amount of one ingredient.
carcinogen — A substance or environmental factor that can cause cancer.
Carcinogens induce mutations in DNA, particularly in oncogenes or tumour suppressor genes, which can disrupt normal cell cycle control and lead to uncontrolled cell division and tumour formation. They act like saboteurs introducing errors into blueprints, causing the production line to malfunction.
oncogene — The term for a mutated gene that causes cancer.
Oncogenes are typically derived from proto-oncogenes, which normally regulate cell growth and division. When mutated, oncogenes can promote uncontrolled cell proliferation, contributing to cancer development, much like an accelerator pedal stuck in the 'on' position.
metastasis — The spread of cancers in this way is called metastasis.
Metastasis is the process by which cancer cells break away from a primary tumour, travel through the blood or lymphatic system, and form secondary tumours in distant parts of the body. This makes cancer much harder to treat and is a dangerous characteristic of malignant tumours, akin to enemy soldiers escaping a main base to establish new ones.
When discussing cancer, ensure you mention uncontrolled mitosis, tumour formation, and metastasis as key characteristics.
Students often think all tumours are cancerous, but actually only malignant tumours are cancerous; benign tumours do not spread.
In cancer-related questions, always link the disease to a loss of control of the cell cycle, leading to the formation of a tumour.
Use precise terminology. For example, state that 'sister chromatids' are separated during anaphase, not 'chromosomes'.
Definitions Bank
chromatid
One of two identical parts of a chromosome, held together by a centromere, formed during interphase by the replication of the DNA strand.
mitosis
The division of a nucleus into two so that the two daughter cells have exactly the same number and type of chromosomes as the parent cell.
cell cycle
The sequence of events that takes place from one cell division until the next; it is made up of interphase, mitosis and cytokinesis.
kinetochore
A protein structure found at the centromere of a chromatid to which microtubules attach during nuclear division.
asexual reproduction
The production of new individuals of a species by a single parent organism.
+7 more definitions
View all →Command Word Guide
| Describe | For mitosis stages, describe the behaviour and arrangement of chromosomes, nuclear envelope, cell surface membrane, and spindle at each specific stage (Prophase, Metaphase, Anaphase, Telophase). |
| Explain | When explaining the importance of mitosis, link it to specific biological processes like growth, repair, and asexual reproduction, detailing how genetically identical cells are produced. For telomeres, explain how they prevent gene loss due to incomplete DNA replication. |
| Outline | For the cell cycle, outline the main events in interphase (G1, S, G2) and the mitotic phase (mitosis and cytokinesis). For stem cells, outline their key characteristics: unspecialised, ability to divide unlimited times, and potential to differentiate. |
| Identify | Be able to identify the different stages of mitosis (Prophase, Metaphase, Anaphase, Telophase) in photomicrographs, diagrams, or microscope slides based on chromosome appearance and location. |
Common Mistakes
Interphase is considered a 'resting phase'.
Interphase is a period of intense metabolic activity, growth, and DNA synthesis, not rest.
Confusing a chromatid with a whole chromosome.
A replicated chromosome consists of two identical sister chromatids joined by a centromere. A chromatid is one of these identical halves.
Believing mitosis is the entire cell division process.
Mitosis is nuclear division only. The entire process, including cytoplasmic division, is called cell division (mitosis followed by cytokinesis).
+3 more
View all →This chapter explores nucleic acids, DNA and RNA, as the fundamental molecules of life, detailing their nucleotide structure and how they form polynucleotide chains. It covers semi-conservative DNA replication and the process of protein synthesis through transcription and translation, along with mRNA modification and the impact of gene mutations.
nucleotide — a molecule consisting of a nitrogen-containing base, a pentose sugar and a phosphate group
Nucleotides are the monomer units that link together to form the long chains of nucleic acids like DNA and RNA. The specific sequence of bases in these nucleotides carries genetic information. ATP is also a nucleotide, but functions as an energy currency rather than a genetic building block, much like individual LEGO bricks assembled into complex structures.
Students often think that nucleotides are only found in DNA and RNA, but actually ATP (adenosine triphosphate) is also a nucleotide that plays a crucial role in energy transfer.
When asked to describe a nucleotide, ensure you mention all three components: base, sugar, and phosphate group. Distinguish between ribose and deoxyribose sugars for RNA and DNA respectively.
dinucleotide — two nucleotides joined together by a phosphodiester bond
A dinucleotide is an intermediate step in the formation of a longer polynucleotide chain. The phosphodiester bond is a key covalent linkage in nucleic acid structure, formed through a condensation reaction, much like two train cars coupled together as a small segment of a longer train.
Students often think a dinucleotide is a functional genetic molecule, but actually it's just a short chain of two nucleotides, typically not biologically active on its own in genetic coding.
phosphodiester bond — a bond joining two nucleotides together; there are two ester bonds, one from the shared phosphate group to each of the sugars either side of it
This covalent bond forms the sugar-phosphate backbone of nucleic acids, providing structural integrity to the DNA and RNA molecules. It is formed via a condensation reaction, releasing a water molecule. The phosphate group acts like a double-sided connector, linking sugars on either side to form a continuous chain.
Students often think phosphodiester bonds are weak like hydrogen bonds, but actually they are strong covalent bonds that form the stable backbone of nucleic acids.
When asked about the formation of phosphodiester bonds, remember to mention that it's a condensation reaction and that it links the phosphate of one nucleotide to the sugar of another.
polynucleotide — a chain of nucleotides joined together by phosphodiester bonds
Polynucleotides form the backbone of DNA and RNA molecules. The phosphodiester bonds create a strong, stable chain, with the bases projecting outwards, ready for pairing or coding. If a nucleotide is a single bead, a polynucleotide is a string of many beads, with the string itself (sugar-phosphate backbone) being strong.
Students often think that polynucleotides are always double-stranded, but actually RNA is a single polynucleotide strand, and even DNA is made of two separate polynucleotide strands.
When describing polynucleotides, remember to mention the phosphodiester bonds that link the nucleotides, forming the sugar-phosphate backbone.
DNA is a double helix of two anti-parallel polynucleotide strands, while RNA is typically a single polynucleotide strand. Both are composed of nucleotides, but DNA contains deoxyribose sugar and bases Adenine (A), Thymine (T), Cytosine (C), and Guanine (G). RNA contains ribose sugar and bases Adenine (A), Uracil (U), Cytosine (C), and Guanine (G). The specific sequence of bases carries genetic information.
complementary base pairing — the hydrogen bonding of A with T or U and of C with G in nucleic acids
This specific pairing (A with T/U, C with G) is fundamental to the structure of DNA (double helix) and RNA (tRNA folding, mRNA-tRNA interaction). It ensures accurate DNA replication and transcription, and precise translation during protein synthesis. A-T pairs have two hydrogen bonds, while C-G pairs have three, making C-G bonds stronger, much like specific puzzle pieces fitting together perfectly.
Always specify which bases pair with which (A-T/U, C-G) and the number of hydrogen bonds involved (2 for A-T/U, 3 for C-G). This detail is often required for full marks.
Students often think that any base can pair with any other base, but actually specific purine-pyrimidine pairings (A-T/U, C-G) are dictated by hydrogen bonding and molecular shape.
DNA replication is the process by which a DNA molecule is copied to form two identical molecules. This process is semi-conservative, meaning each new DNA molecule contains one strand from the original molecule and one newly synthesised strand. This mechanism ensures that genetic information is accurately passed from parent to daughter cells.
semi-conservative replication — the method by which a DNA molecule is copied to form two identical molecules, each containing one strand from the original molecule and one newly synthesised strand
This mechanism ensures that genetic information is accurately passed from parent to daughter cells. Each new DNA molecule is a hybrid, conserving half of the original molecule. This is like making two copies of a book by splitting the original, and using each half as a template to print a new complementary half.
Students often think 'conservative' replication means the original DNA molecule stays completely intact, but actually semi-conservative means each new molecule has one old and one new strand.
When explaining semi-conservative replication, clearly state that each new DNA molecule consists of one original (parent) strand and one newly synthesized strand.
DNA polymerase — an enzyme that copies DNA; it runs along the separated DNA strands lining up one complementary nucleotide at a time ready for joining by DNA ligase
DNA polymerase is crucial for DNA replication, synthesizing new DNA strands by adding nucleotides complementary to the template strand. It can only synthesize in the 5' to 3' direction, leading to the formation of leading and lagging strands. It acts like a skilled builder, selecting correct new bricks for a growing structure.
When describing DNA polymerase, mention its role in adding complementary nucleotides and its 5' to 3' synthesis direction, which explains the leading and lagging strands.
leading strand — during DNA replication, the parent strand that runs in the 3′ to 5′ direction is copied to produce the leading strand
The leading strand is synthesized continuously because DNA polymerase can move in the same direction as the unwinding replication fork. This continuous synthesis simplifies the process compared to the lagging strand, much like a car driving smoothly forward on a continuously opening road.
lagging strand — during DNA replication, the parent strand that runs in the 5′ to 3′ direction is copied to produce the lagging strand
The lagging strand is synthesized discontinuously in short segments called Okazaki fragments because DNA polymerase must work backwards relative to the unwinding fork. These fragments are later joined by DNA ligase. This is like a car that has to drive forward a bit, then reverse, then drive forward again on a road opening in the opposite direction.
Students often think both new strands are synthesized in the same way, but actually the leading strand is continuous while the lagging strand is discontinuous due to the enzyme's directionality.
When discussing the lagging strand, remember to mention Okazaki fragments and the role of DNA ligase in joining them, explaining why it's synthesized discontinuously.
DNA ligase — an enzyme that catalyses the joining together of two nucleotides with covalent phosphodiester bonds during DNA replication
DNA ligase is essential for completing DNA replication, particularly on the lagging strand where it joins Okazaki fragments. It forms phosphodiester bonds, ensuring the integrity and continuity of the newly synthesized DNA backbone. It acts like a molecular 'glue' that seals gaps, creating a continuous strand.
Students often think DNA polymerase does all the joining, but actually DNA ligase is specifically responsible for forming the final phosphodiester bonds between newly synthesized fragments.
The genetic code in DNA dictates the sequence of amino acids in a polypeptide. A gene, a specific length of DNA, codes for a particular polypeptide or protein. This code is universal, meaning it is largely the same across all organisms, and is read in triplets of bases called codons, which specify amino acids or start/stop signals.
gene — a length of DNA that codes for a particular polypeptide or protein
Genes are the fundamental units of heredity, carrying the instructions for building and maintaining an organism. The sequence of bases within a gene determines the sequence of amino acids in a polypeptide, which in turn dictates the protein's structure and function. Think of a gene as a specific recipe in a cookbook (the entire DNA molecule).
Students often think a gene codes directly for a trait, but actually a gene codes for a polypeptide, which then contributes to a trait.
Define a gene as coding for a polypeptide or protein, not directly for a characteristic. Emphasize that it's a 'length of DNA'.
Transcription is the first stage of protein synthesis, where the genetic information in a DNA molecule is copied into a complementary strand of messenger RNA (mRNA). This process occurs in the nucleus of eukaryotic cells, with RNA polymerase using only one DNA strand (the template strand) as a guide. Uracil replaces thymine in the mRNA molecule.
transcription — copying the genetic information in a molecule of DNA into a complementary strand of mRNA; a single strand of the DNA is used as a template (this is called the template or transcribed strand) – the enzyme responsible is RNA polymerase
Transcription is the first stage of protein synthesis, occurring in the nucleus of eukaryotic cells. It allows the genetic information to be carried from the DNA in the nucleus to the ribosomes in the cytoplasm, where proteins are made. Only one DNA strand, the template strand, is used for this process, much like making a temporary working copy of a master blueprint.
Students often think both DNA strands are copied during transcription, but actually only the template strand is used.
When describing transcription, specify that it occurs in the nucleus, uses RNA polymerase, and produces mRNA from a DNA template strand, remembering that uracil replaces thymine in RNA.
After transcription in eukaryotes, the initial mRNA transcript undergoes modification. This involves the removal of non-coding regions called introns, while the coding regions, exons, are spliced together. This processing ensures that only the relevant genetic information is translated into protein.
Translation is the second stage of protein synthesis, where the sequence of nucleotides in an mRNA molecule is converted into a corresponding sequence of amino acids in a polypeptide chain. This process takes place at ribosomes in the cytoplasm, with transfer RNA (tRNA) molecules bringing the correct amino acids based on complementary base pairing with mRNA codons.
translation — a stage in protein synthesis during which a sequence of nucleotides in a molecule of messenger RNA (mRNA) is converted (translated) into a corresponding sequence of amino acids in a polypeptide chain; it takes place at ribosomes
Translation is the second stage of protein synthesis, where the genetic code carried by mRNA is used to assemble a specific sequence of amino acids into a polypeptide. This process occurs on ribosomes in the cytoplasm and involves tRNA molecules bringing the correct amino acids. It is the decoding of the mRNA message into a protein, much like a chef reading a recipe and assembling a dish.
Students often think translation happens in the nucleus, but actually it occurs in the cytoplasm at the ribosomes.
For translation, remember to mention the ribosome as the site, mRNA as the template, tRNA as the amino acid carrier, and the product as a polypeptide chain.
codon — sequence of three bases on an mRNA molecule that codes for a specific amino acid or for a stop signal
Codons are the fundamental units of the genetic code in mRNA, read by ribosomes during translation. Each codon specifies either a particular amino acid to be added to the polypeptide chain or a signal to terminate protein synthesis. The degeneracy of the genetic code means some amino acids are coded by multiple codons, like a three-letter word specifying an action.
Students often think codons are found on DNA, but actually codons are specifically sequences of three bases on mRNA.
Distinguish between a DNA triplet and an mRNA codon. Codons are specifically on mRNA and are read during translation.
anticodon — sequence of three unpaired bases on a tRNA molecule that binds with a codon on mRNA
The anticodon is crucial for ensuring that the correct amino acid is delivered to the ribosome during translation. Its complementary pairing with the mRNA codon ensures the accurate sequencing of amino acids in the growing polypeptide chain. If a codon is a lock, the anticodon is the specific key that fits it.
Students often think the anticodon codes for the amino acid, but actually it's the codon on the mRNA that codes for the amino acid, and the anticodon simply ensures the correct tRNA (carrying that amino acid) binds.
Gene mutations are changes in the base sequence in part of a DNA molecule. These random events can alter the genetic code, potentially leading to changes in the amino acid sequence of a polypeptide. While often harmful, they are also the source of genetic variation. Mutations can arise from errors during DNA replication or damage from mutagens.
gene mutation — a change in the base sequence in part of a DNA molecule
Gene mutations are random events that can alter the genetic code, potentially leading to changes in the amino acid sequence of a polypeptide. These changes can arise from errors during DNA replication or damage from mutagens. While often harmful, they are also the source of genetic variation, much like a typo in a recipe.
Students often think all gene mutations are harmful, but actually some can be neutral or even beneficial, though harmful ones are more likely.
When defining gene mutation, specify that it's a change in the 'base sequence' of DNA, not just any change in DNA.
Gene mutations include substitution, insertion, and deletion. A substitution mutation involves the replacement of one base with another. Insertion and deletion mutations involve the addition or removal of one or more nucleotides, respectively. These latter two types often lead to more severe consequences due to frame-shift effects.
frame-shift mutation — a type of gene mutation caused by insertion or deletion of one or more nucleotides, resulting in incorrect reading of the sequence of triplets in the genetic code due to a shift in the reading frame
Frame-shift mutations are typically more severe than substitutions because they alter every subsequent codon downstream from the mutation. This usually leads to a completely different amino acid sequence, often resulting in a non-functional protein or a premature stop codon. This is like deleting a letter in a sentence without spaces, completely changing the meaning from that point onwards.
When explaining frame-shift mutations, clearly state that they are caused by insertions or deletions (not substitutions) and that they alter the 'reading frame', affecting all subsequent codons and amino acids.
chromosome mutation — a random and unpredictable change in the structure or number of chromosomes in a cell
Chromosome mutations are larger-scale changes than gene mutations, affecting entire chromosomes or significant parts of them. These can involve deletions, duplications, inversions, or translocations of chromosomal segments, or changes in the total number of chromosomes. They often have more severe phenotypic consequences than gene mutations, like deleting an entire chapter of a book rather than just a typo.
Students often confuse gene mutations with chromosome mutations, but actually gene mutations affect the base sequence within a gene, while chromosome mutations involve larger structural or numerical changes to chromosomes.
For descriptions of transcription or translation, always state the precise location (nucleus/cytoplasm), the key molecules involved (e.g., mRNA, tRNA, ribosome), and the final product.
Use sickle cell anaemia as a specific example of a substitution mutation, explaining how a single base change alters the polypeptide and protein function.
Definitions Bank
nucleotide
a molecule consisting of a nitrogen-containing base, a pentose sugar and a phosphate group
polynucleotide
a chain of nucleotides joined together by phosphodiester bonds
dinucleotide
two nucleotides joined together by a phosphodiester bond
phosphodiester bond
a bond joining two nucleotides together; there are two ester bonds, one from the shared phosphate group to each of the sugars either side of it
complementary base pairing
the hydrogen bonding of A with T or U and of C with G in nucleic acids
+13 more definitions
View all →Command Word Guide
| Describe | For nucleotides, describe all three components (base, sugar, phosphate). For DNA/RNA structure, describe the double helix, anti-parallel strands, complementary base pairing, and phosphodiester bonds. For replication/synthesis, describe the steps and molecules involved. |
| Explain | For semi-conservative replication, explain how each new molecule gets one old and one new strand. For protein synthesis, explain the roles of DNA, mRNA, tRNA, ribosomes, and enzymes in transcription and translation. For mutations, explain how a change in base sequence leads to altered polypeptide/protein function. |
| Compare | When comparing DNA and RNA, use a table and state clear differences: pentose sugar (deoxyribose vs. ribose), bases (T vs. U), and structure (double vs. single strand). |
Common Mistakes
Students often think that proteins are the genetic material.
DNA is the genetic molecule, carrying the hereditary information.
Students often confuse adenine with adenosine, or thymine with thiamine.
Be precise with base names: Adenine, Thymine, Cytosine, Guanine, Uracil.
Students often think both strands of DNA are copied during transcription.
Only one DNA strand, the template strand, is used for transcription.
+3 more
View all →This chapter details the essential transport systems in plants, focusing on xylem for water and mineral transport and phloem for organic solute transport. It covers the structure of dicotyledonous stems, roots, and leaves, explaining passive water movement via transpiration and active assimilate transport through mass flow.
vascular system — a system of fluid-filled tubes, vessels or spaces, most commonly used for long-distance transport in living organisms; examples are the blood vascular system in animals and the vascular system of xylem and phloem in plants
This system enables efficient transport of materials over long distances in multicellular organisms, overcoming the limitations of diffusion. In plants, it comprises xylem for water and minerals, and phloem for organic solutes, much like a city's plumbing system with separate pipes for fresh water and waste/recycled materials.
vascular — a term referring to tubes or vessels (from the Latin ‘vascul’, meaning vessel)
This term is broadly used in biology to describe structures that facilitate fluid transport. In plants, it specifically refers to the xylem and phloem tissues, similar to how 'tubular' refers to tubes.
xylem — a tissue containing tubes called vessels and other types of cell, responsible for the transport of water and mineral salts through a plant and for support
Xylem vessels are dead, lignified tubes that form a continuous pathway for water and mineral ions from roots to leaves. Lignin also provides crucial structural support to the plant, much like water pipes in a building bringing fresh water up from the ground.
Students often think xylem is only for transport, but actually its lignified walls also provide significant structural support to the plant.
When describing xylem's function, remember to include both water/mineral transport and mechanical support, as both are key adaptations.
phloem — a tissue containing tubes called sieve tubes and other types of cell, responsible for the transport through the plant of organic solutes (assimilates) such as sucrose
Phloem consists of living sieve tube elements and companion cells, transporting sugars and amino acids from sources (e.g., leaves) to sinks (e.g., roots, fruits) throughout the plant. It acts like a food delivery system, taking prepared meals from the kitchen (leaves) to all hungry parts of the house.
Students often think phloem sap moves only downwards, but actually it can move in any direction, from source to sink, depending on the plant's needs.
Distinguish clearly between xylem (water, unidirectional) and phloem (assimilates, bidirectional) in your explanations, especially when comparing them.
vascular tissue — a tissue in plants consisting mainly of xylem and phloem but also containing sclerenchyma and parenchyma cells
This collective term refers to the primary transport tissues in plants, forming vascular bundles in stems and leaves, and a central core in roots. It includes supportive and packing cells alongside the main transport vessels, much like a utility corridor containing main lines, structural supports, and maintenance access.
dicotyledon — flowering plants can be classified as monocotyledons or dicotyledons; the seeds of dicotyledonous plants contain an embryo with two cotyledons (seed leaves) in their seeds and the adult plant typically has leaves with a blade (lamina) and a stalk (petiole)
Dicotyledons are a major group of flowering plants characterized by having two embryonic leaves in their seeds. Their vascular tissue arrangement in stems, roots, and leaves differs from monocotyledons, similar to classifying cars into sedans and SUVs.
eyepiece graticule — small scale that is placed in a microscope eyepiece
This scale is used in conjunction with a stage micrometer to calibrate the microscope and measure the size of specimens viewed through the eyepiece. It's like a transparent ruler placed inside binoculars, but it needs calibration.
stage micrometer — very small, accurately drawn scale of known dimensions, engraved on a microscope slide
The stage micrometer is used to calibrate the eyepiece graticule, allowing accurate measurement of specimens under the microscope at various magnifications. It serves as the 'master ruler' to set the scale of the eyepiece graticule.
Be prepared to describe the calibration process of an eyepiece graticule using a stage micrometer in practical questions.
vascular bundle — a strand of vascular tissue running longitudinally in a plant; within the bundle, the arrangement of tissues like xylem, phloem and sclerenchyma varies in different plants and organs
Vascular bundles are discrete units containing xylem, phloem, and often sclerenchyma fibers, responsible for transport and support. Their arrangement is characteristic of different plant organs, much like a utility cable containing multiple wires and protective sheathing.
parenchyma — a basic plant tissue typically used as packing tissue between more specialised structures; it is metabolically active and may have a variety of functions such as food storage and support; parenchyma cells also play an important role in the movement of water and food products in the xylem and phloem
Parenchyma cells are versatile, thin-walled cells that make up the bulk of plant tissues, including the cortex and pith. They are involved in storage, photosynthesis, and short-distance transport, acting as the general-purpose 'filler' or 'support staff' in a plant.
collenchyma — a modified form of parenchyma in which the corners of the cells have extra cellulose thickening, providing extra support, as in the midrib of leaves and at the corners of square stems; in three dimensions the tissue occurs in strands (as in celery petioles)
Collenchyma provides flexible support to growing parts of the plant, unlike sclerenchyma which provides rigid support. Its thickened cellulose walls are characteristic, similar to flexible plastic tubing providing support.
epidermis — the outer layer of cells covering the body of a plant or animal; in plants it is usually one cell thick and may be covered with a cuticle which provides additional protection against loss of water and disease
The epidermis forms the protective outermost layer of plant organs, regulating gas exchange and water loss, and defending against pathogens. It is typically a single cell layer thick, much like the skin of a plant.
endodermis — the layer of cells surrounding the vascular tissue of plants; it is most clearly visible in roots
The endodermis, particularly in roots, contains the Casparian strip, a waxy band that blocks the apoplast pathway, forcing water and solutes into the symplast pathway and allowing the plant to regulate uptake. It acts like a security checkpoint around the plant's central transport hub.
Students often think water can freely enter the xylem from the cortex, but actually the endodermis with its Casparian strip regulates this movement.
When explaining water movement across the root, explicitly mention the Casparian strip and its role in blocking the apoplast pathway at the endodermis.
sclerenchyma — a plant tissue consisting of thick-walled cells with a purely mechanical function (strength and support); the cell walls have usually become impregnated with lignin and the mature cells are dead with no visible contents; many sclerenchyma cells take the form of fibres
Sclerenchyma provides rigid, non-stretching support to mature plant parts. Its cells are dead at maturity, with heavily lignified walls, forming fibers or sclereids, much like the steel beams or concrete in a building.
lignin — a hard material made by plants and used to strengthen the cell walls of certain types of cell, particularly xylem vessel elements and sclerenchyma cells; it is the main material in wood
Lignin is a complex polymer that makes cell walls waterproof and rigid, providing mechanical strength and preventing collapse of xylem vessels under tension. It is a key component of wood, similar to concrete or rebar in a building's structure.
Plants require efficient transport systems to move water and mineral ions absorbed by the roots to all parts of the plant, especially the leaves for photosynthesis. Simultaneously, organic compounds, or assimilates, produced during photosynthesis in the leaves need to be transported to other areas for growth, storage, or metabolic activity. Both mineral ions and organic compounds are transported dissolved in water, necessitating a robust vascular system for long-distance movement.
Dicotyledonous plants exhibit distinct arrangements of vascular tissues in their organs. In stems, vascular bundles, containing both xylem and phloem, are typically arranged in a ring. Roots feature a central core of vascular tissue, with xylem often forming a star shape. Leaves, on the other hand, have vascular bundles forming veins, which are distributed throughout the mesophyll tissue. These arrangements are crucial for efficient transport and support.
When drawing low-power plans, ensure you correctly identify and label the vascular tissue and its constituent parts within the organ.
transpiration — the loss of water vapour from a plant to its environment; it mostly takes place through the stomata in the leaves
Transpiration is the evaporative loss of water from plant surfaces, primarily leaves, driven by the sun's energy. This process creates a water potential gradient that pulls water up the plant, much like a plant 'sweating' to cool itself and create suction.
When explaining transpiration, ensure you mention the role of stomata, the evaporation from mesophyll cell walls, and the resulting water potential gradient.
mesophyll — the region of a leaf between the upper and lower epidermis; in dicotyledonous plants the mesophyll has an upper palisade layer and a lower mesophyll layer; the palisade mesophyll cells are column-shaped and form the main photosynthetic layer, whereas the spongy mesophyll has large air spaces between the cells for gas exchange
Mesophyll tissue is the primary site of photosynthesis in leaves, with specialized palisade cells for light absorption and spongy cells with air spaces for efficient gas exchange and water vapor accumulation. It acts as the 'engine room' of the leaf, where most energy production occurs.
stoma — a pore in the epidermis of a leaf, bounded by two guard cells and needed for efficient gas exchange
Stomata regulate the exchange of carbon dioxide, oxygen, and water vapor between the plant and the atmosphere. Their opening and closing are controlled by guard cells in response to environmental cues, acting like tiny adjustable windows on the leaf surface.
Students often think stomata are always open, but actually they close at night or during water stress to conserve water.
xerophyte — a plant adapted to survive in conditions where water is in short supply
Xerophytes possess various structural and physiological adaptations, such as thick cuticles, sunken stomata, rolled leaves, or reduced leaf surface area, to minimize water loss in dry environments. A xerophyte is like a desert survivalist, equipped with special strategies to conserve water.
When asked to describe xerophytic adaptations, provide specific examples (e.g., sunken stomata, hairs) and explain *how* each feature reduces water loss.
cuticle — a layer covering, and secreted by, the epidermis; in plants it is made of a fatty substance called cutin, which helps to provide protection against water loss and infection
The cuticle is a waxy, waterproof layer on the outer surface of leaves and stems, reducing uncontrolled water evaporation and providing a barrier against pathogens. It's like a clear, waterproof raincoat covering the plant.
Water and mineral ions enter the root from the soil, primarily through root hairs. They then move across the root cortex towards the central xylem via two main pathways: the apoplast pathway and the symplast pathway. The apoplast pathway involves movement through the non-living cell walls and intercellular spaces. The symplast pathway involves movement through the cytoplasm of living cells, connected by plasmodesmata. At the endodermis, the Casparian strip blocks the apoplast pathway, forcing water and solutes into the symplast, allowing the plant to regulate uptake before entering the xylem.
symplast pathway — the living system of interconnected protoplasts extending through a plant, used as a transport pathway for the movement of water and solutes; individual protoplasts are connected via plasmodesmata
In the symplast pathway, water and solutes move through the cytoplasm of living cells, passing from one cell to the next through plasmodesmata, which are cytoplasmic connections. This is like water moving through interconnected rooms in a house, passing directly from one room's interior to the next.
apoplast pathway — the non-living system of interconnected cell walls extending throughout a plant, used as a transport pathway for the movement of water and mineral ions
In the apoplast pathway, water and mineral ions move through the non-living spaces of the cell walls and intercellular spaces, without entering the cytoplasm of the cells. This is similar to water soaking through a sponge or moving through the gaps between bricks in a wall.
Water moves up the xylem from root to leaf passively, driven by the transpiration pull. As water evaporates from the leaves (transpiration), it creates a tension, or negative pressure, in the xylem. This tension pulls the continuous column of water upwards. The cohesive forces between water molecules (cohesion) prevent the water column from breaking, while adhesive forces between water molecules and the lignified xylem walls (adhesion) prevent the column from pulling away from the walls. This combined effect of cohesion, adhesion, and tension is known as the cohesion-tension theory.
When explaining water movement up the xylem, you must use and explain the terms cohesion (water-water attraction), adhesion (water-xylem wall attraction), and tension (the pull from transpiration).
xylem vessel element — a dead, lignified cell found in xylem specialised for transporting water and for support; the ends of the cells break down and join with neighbouring elements to form long tubes called xylem vessels
These individual cells, after lignification and loss of protoplast, form continuous, hollow tubes. Their structure is highly adapted for efficient, unidirectional water transport and mechanical support. Each element is like a single, hollow pipe segment that, when joined, forms a long pipeline.
Students often think xylem vessel elements are living cells, but actually they are dead at maturity, leaving an empty lumen for water flow.
xylem vessel — a dead, empty tube with lignified walls, through which water is transported in plants; it is formed by xylem vessel elements lined up end to end
Xylem vessels are the primary conduits for long-distance water and mineral transport in plants. Their continuous, lignified structure allows for mass flow under tension and provides structural integrity. A xylem vessel is the complete pipeline running through the plant.
Assimilates, such as sucrose and amino acids, are transported through phloem sieve tubes from areas of production (sources) to areas of utilization or storage (sinks). This movement occurs via mass flow, driven by a hydrostatic pressure gradient. Active loading of sucrose at the source creates a high solute concentration, causing water to move in by osmosis and build up high hydrostatic pressure. At the sink, unloading of sucrose reduces the solute concentration, leading to water moving out by osmosis and a lower hydrostatic pressure. This pressure difference drives the bulk flow of phloem sap.
Structure your mass flow explanation logically: 1. Active loading of sucrose at source lowers water potential. 2. Water enters by osmosis, creating high hydrostatic pressure. 3. Unloading at sink raises water potential. 4. Water leaves by osmosis, creating low hydrostatic pressure. 5. Sap flows down the pressure gradient.
source — a site in a plant which provides food for another part of the plant, the sink
Sources are typically photosynthetic leaves where sugars are produced, or storage organs when they are releasing stored food. They are characterized by a high concentration of assimilates, acting like a food factory or pantry.
sink — a site in a plant which receives food from another part of the plant, the source
Sinks are regions of growth, development, or storage, such as roots, fruits, flowers, or young leaves, where assimilates are consumed or stored. They have a lower concentration of assimilates, similar to a construction site or storage warehouse needing food.
sieve tube element — a cell found in phloem tissue, with non-thickened cellulose walls, very little cytoplasm, no nucleus and end walls perforated to form sieve plates, through which sap containing sucrose is transported
Sieve tube elements are living cells, but lack a nucleus and most organelles at maturity, forming continuous tubes for efficient assimilate transport. They rely on companion cells for metabolic support, much like a segment of a food delivery tube that is mostly hollowed out but still alive.
Students often think sieve tube elements are dead like xylem vessels, but actually they are living cells, albeit highly modified with reduced organelles.
companion cell — a cell with an unthickened cellulose wall and dense cytoplasm that is found in close association with a phloem sieve tube element to which it is directly linked via many plasmodesmata; the companion cell and the sieve tube element form a functional unit
Companion cells are metabolically active cells that support the sieve tube elements, providing ATP for active loading of assimilates and maintaining their cellular functions. They are connected by numerous plasmodesmata, acting like the control room or support staff for the sieve tube element.
When explaining active loading, describe the role of companion cells in pumping H+ ions and co-transporting sucrose into the sieve tube elements.
sieve tube — tube formed from sieve tube elements lined up end to end
Sieve tubes are the continuous conduits within the phloem tissue through which organic solutes, primarily sucrose, are transported throughout the plant via mass flow. A sieve tube is the complete food delivery pipeline, made up of many connected sieve tube elements.
For low-power plan diagrams, draw the distribution of tissues with clear, un-shaded outlines. Do NOT draw individual cells. In high-power drawings, draw only a few representative cells accurately. Use a sharp pencil for single lines and include clear labels pointing to the specific structure.
Definitions Bank
vascular system
a system of fluid-filled tubes, vessels or spaces, most commonly used for long-distance transport in living organisms; examples are the blood vascular system in animals and the vascular system of xylem and phloem in plants
vascular
a term referring to tubes or vessels (from the Latin ‘vascul’, meaning vessel)
xylem
a tissue containing tubes called vessels and other types of cell, responsible for the transport of water and mineral salts through a plant and for support
phloem
a tissue containing tubes called sieve tubes and other types of cell, responsible for the transport through the plant of organic solutes (assimilates) such as sucrose
vascular tissue
a tissue in plants consisting mainly of xylem and phloem but also containing sclerenchyma and parenchyma cells
+24 more definitions
View all →Command Word Guide
| Outline | When asked to 'outline' the transport needs, ensure you mention both water/minerals from roots and organic food from leaves, and the need for long-distance transport. |
| Describe | When asked to 'describe' xerophytic adaptations, provide specific examples (e.g., sunken stomata, hairs) and explain *how* each feature reduces water loss. |
| Explain | When asked to 'explain' water movement up the xylem, you must use and explain the terms cohesion (water-water attraction), adhesion (water-xylem wall attraction), and tension (the pull from transpiration). |
| Draw | For low-power plan diagrams, draw the distribution of tissues with clear, un-shaded outlines. Do NOT draw individual cells. For high-power drawings, draw only a few representative cells accurately. Use a sharp pencil for single lines and include clear labels pointing to the specific structure. |
Common Mistakes
Students often think xylem is only for transport.
Remember that xylem's lignified walls also provide significant structural support to the plant.
Students often think phloem sap moves only downwards.
Phloem sap can move in any direction, from source to sink, depending on the plant's needs.
Students often think stomata are always open.
Stomata close at night or during water stress to conserve water.
+3 more
View all →This chapter explores the mammalian circulatory system, a closed double circulation involving the heart and blood vessels. It details the structure-function relationships of various blood vessels, the formation and role of tissue fluid, and the composition and functions of blood, including gas transport and the Bohr shift. Finally, it covers the heart's structure, the cardiac cycle, and its myogenic control.
circulatory system — A system that carries fluids around an organism’s body.
In mammals, this is a closed double circulatory system involving the heart, blood vessels, and blood. Its primary role is to transport oxygen, nutrients, hormones, and waste products, much like a city's plumbing system with a central pump maintaining flow.
closed blood system — A circulatory system made up of vessels containing blood.
In a closed system, blood always remains within the blood vessels, ensuring efficient transport and maintaining high pressure. This is similar to a sealed central heating system where fluid stays within pipes, never spilling out, though exchange still occurs across capillary walls.
Students often think 'closed' means it's a sealed loop with no exchange, but actually it means blood is contained within vessels, while exchange still occurs across capillary walls.
double circulation — A circulatory system in which the blood passes through the heart twice on one complete circuit of the body.
This system consists of two loops: the pulmonary circulation (heart to lungs and back) and the systemic circulation (heart to body and back). This allows for higher blood pressure in the systemic circuit for efficient delivery to tissues, like two separate train lines starting and ending at the same station.
Students often think 'double' means two hearts, but actually it refers to the blood passing through the single heart twice per full body circuit.
When describing the circulatory system, ensure you mention both the 'closed' and 'double' aspects, as these are key distinguishing features of mammals.
systemic circulation — The part of the circulatory system that carries blood from the heart to all of the body except the gas exchange surface, and then back to the heart.
This circuit delivers oxygenated blood and nutrients to body tissues and collects deoxygenated blood and waste products. It operates at a higher pressure than the pulmonary circulation, acting like the main highway system of a country.
pulmonary circulation — The part of the circulatory system that carries blood from the heart to the gas exchange surface and then back to the heart.
This circuit transports deoxygenated blood from the right ventricle to the lungs to pick up oxygen and release carbon dioxide, then returns oxygenated blood to the left atrium. It operates at a lower pressure to protect delicate lung capillaries, much like a short shuttle bus route to an airport.
Clearly distinguish between the pulmonary and systemic circulations when explaining double circulation, mentioning the path and purpose of each.
artery — Vessel with thick, strong walls that carries high-pressure blood away from the heart.
Arteries have thick, elastic, and muscular walls to withstand and smooth out the high-pressure, pulsed flow of blood from the heart. Their structure allows them to stretch and recoil, maintaining blood flow, similar to a high-pressure fire hose.
vein — Vessel with relatively thin walls that carries low-pressure blood back to the heart.
Veins have thinner walls and larger lumens compared to arteries, as they carry blood at much lower pressure. They contain semilunar valves to prevent backflow of blood, especially against gravity, functioning like a drainage pipe.
Students often think arteries always carry oxygenated blood and veins always carry deoxygenated blood, forgetting the pulmonary circulation exceptions.
When comparing arteries and veins, focus on the thickness and composition of the wall (elastic and muscle fibres) and the pressure of the blood they carry, not just the direction of flow.
arteriole — Small artery.
Arterioles contain a lot of smooth muscle in their walls, allowing them to constrict (vasoconstriction) or dilate (vasodilation) to control blood flow to capillary beds and reduce blood pressure before it enters capillaries, acting like a tap controlling water flow.
venule — Small vein.
Venules are formed when capillaries join together, collecting blood from the capillary beds and gradually merging to form larger veins. They have thin walls, reflecting the low blood pressure, much like small streams joining to form larger rivers.
capillary — The smallest blood vessel, whose role is to deliver oxygen and nutrients to body tissues, and to remove their waste products.
Capillaries have extremely thin walls, only one cell thick (endothelium), and a very narrow lumen, just wide enough for red blood cells to pass in single file. This structure facilitates rapid exchange of substances between blood and tissue fluid, like a fine mesh net.
When describing capillary function, always link its structural features (thin walls, narrow lumen, extensive network) directly to its role in efficient substance exchange.
endothelium — A tissue that lines the inner surface of a structure such as a blood vessel.
It consists of a single layer of flat cells (squamous epithelium) that provides a very smooth surface, minimizing friction with moving blood. In capillaries, this single layer forms the entire wall, facilitating diffusion, much like a smooth, non-stick lining of a pipe.
squamous epithelium — One or more layers of thin, flat cells forming the lining of some hollow structures, e.g. blood vessels and alveoli.
In blood vessels, a single layer of squamous epithelium forms the endothelium, providing a smooth surface and a short diffusion distance for substance exchange in capillaries. Its thinness is key to its function, like a single layer of thin, flat tiles.
smooth muscle — A type of muscle that can contract steadily over long periods of time.
Found in the walls of arteries and arterioles, smooth muscle allows for sustained contraction or relaxation, altering the diameter of the vessels. This controls blood flow and pressure to different tissues, responding to nerve impulses and hormones, similar to the slow contraction of a sphincter muscle.
elastic arteries — Relatively large arteries, which have a lot of elastic tissue and little muscle tissue in their walls.
These arteries, like the aorta, are close to the heart and stretch to accommodate the high-pressure blood surge from the ventricles. Their recoil helps to 'even out' blood flow and maintain pressure during diastole, like a large, strong rubber band.
muscular arteries — Arteries that are closer to the final destination of the blood inside them than elastic arteries, with more smooth muscle in their walls which allows them to constrict and dilate.
These arteries regulate blood flow to specific regions of the body by contracting or relaxing their smooth muscle. This control is vital for diverting blood to active tissues or reducing flow to less active ones, like adjustable pipes in a plumbing system.
vasoconstriction — The narrowing of a muscular artery or arteriole, caused by the contraction of the smooth muscle in its walls.
This process reduces blood flow to a particular area and increases blood pressure upstream. It is controlled by nerve impulses and hormones, allowing the body to divert blood to other tissues, similar to squeezing a garden hose.
vasodilation — The widening of a muscular artery or arteriole, caused by the relaxation of the smooth muscle in its walls.
This process increases blood flow to a particular area and decreases blood pressure upstream. It is also controlled by nerve impulses and hormones, allowing for increased supply to active tissues, like opening a tap fully.
semilunar valve — A half-moon shaped valve, such as the ones in the veins and between the ventricles and arteries.
These valves prevent the backflow of blood. In veins, they ensure unidirectional flow towards the heart, especially against gravity. In the heart, they are found at the exits of the ventricles (aortic and pulmonary valves) to prevent blood from flowing back into the ventricles during diastole, acting like a one-way gate.
Blood pressure is highest in the arteries, particularly the elastic arteries near the heart, due to the direct pumping action of the ventricles. As blood flows through arterioles and into the extensive capillary networks, resistance increases and the total cross-sectional area expands, leading to a significant drop in pressure. Pressure continues to decrease in venules and veins, becoming very low by the time blood returns to the heart, necessitating valves to prevent backflow.
plasma — The liquid component of blood, in which the blood cells float; it carries a very large range of different substances in solution.
Plasma is mostly water (about 95%) and contains dissolved nutrients, waste products, hormones, and plasma proteins. It plays a crucial role in transport and heat distribution throughout the body, much like the water in a river carrying various dissolved substances.
plasma proteins — A range of several different proteins dissolved in the blood plasma, each with their own function; many of them are made in the liver.
These proteins, such as albumin, are too large to easily escape capillaries and are crucial for maintaining the water potential of the blood. They contribute to osmotic pressure, drawing water back into capillaries from tissue fluid, like large, non-diffusible molecules creating an osmotic pull.
tissue fluid — The almost colourless fluid that fills the spaces between body cells; it forms from the fluid that leaks from blood capillaries.
Tissue fluid is similar to blood plasma but contains far fewer protein molecules and no red blood cells. It acts as the medium for exchange of materials (oxygen, nutrients, waste) between blood and body cells, like a 'soup' surrounding individual cells.
Students often think tissue fluid is identical to blood plasma, overlooking the significant difference in protein content and absence of red blood cells.
Use precise terminology for tissue fluid formation. Refer to 'hydrostatic pressure' forcing fluid out and a 'water potential gradient' (due to plasma proteins) drawing fluid back in.
Tissue fluid forms at the arteriole end of capillaries where high hydrostatic pressure forces fluid, containing dissolved nutrients and oxygen, out of the blood and into the interstitial spaces. Large plasma proteins and red blood cells remain in the capillaries. This fluid bathes the body cells, facilitating the exchange of substances. At the venule end of the capillaries, the hydrostatic pressure is lower, and the lower water potential of the blood (due to plasma proteins) draws most of the tissue fluid back into the capillaries by osmosis.
Blood is a vital transport medium composed of plasma and various blood cells. Plasma transports dissolved substances like nutrients, hormones, and waste products. Red blood cells are specialised for oxygen transport, while white blood cells are crucial for immunity. Blood also plays a role in thermoregulation and clotting.
neutrophil — One type of phagocytic white blood cell; it has a lobed nucleus and granular cytoplasm.
Neutrophils are the commonest type of phagocyte, destroying invading microorganisms by phagocytosis. They are a key component of the innate immune system, rapidly responding to infection, like 'first responder' police officers.
monocyte — The largest type of white blood cell; it has a bean-shaped nucleus; monocytes can leave the blood and develop into a type of phagocytic cell called a macrophage.
Monocytes circulate in the blood and then migrate into tissues, where they differentiate into macrophages. Macrophages are powerful phagocytes and also act as antigen-presenting cells, linking innate and adaptive immunity, like a 'reserve' police officer transforming into a detective.
macrophage — Phagocytic cell found in tissues throughout the body; they act as antigen-presenting cells (APCs).
Macrophages engulf and digest pathogens and cellular debris. They also present antigens from pathogens to lymphocytes, initiating specific immune responses. They are crucial for both innate immunity and activating adaptive immunity, like a 'clean-up crew' that also collects evidence.
lymphocyte — A white blood cell with a nucleus that almost fills the cell, which responds to antigens and helps to destroy the antigens or the structure that is carrying them.
Lymphocytes are central to adaptive immunity, recognising specific antigens. They include B cells (producing antibodies) and T cells (cell-mediated immunity). They are smaller than most phagocytes and have a large, round nucleus, acting like 'special forces' soldiers.
haemoglobin — A protein with a quaternary structure, found in red blood cells, that transports oxygen.
Haemoglobin is a respiratory pigment containing iron, which reversibly binds to oxygen. Its structure allows for cooperative binding, meaning the binding of one oxygen molecule increases the affinity for subsequent oxygen molecules, making it highly efficient for oxygen transport.
Haemoglobin-oxygen binding
Represents the reversible binding of four oxygen molecules (eight oxygen atoms) to one haemoglobin molecule.
partial pressure — A measure of the concentration of a gas.
In a mixture of gases, the partial pressure of a specific gas is the pressure it would exert if it alone occupied the volume. It is used to describe gas concentrations in blood and alveoli, influencing gas diffusion, like how much of the total noise in a room is made by one group.
percentage saturation — The degree to which the haemoglobin in the blood is combined with oxygen, calculated as a percentage of the maximum amount with which it can combine.
This value indicates how much oxygen haemoglobin is carrying relative to its full capacity. It varies with the partial pressure of oxygen, as shown by the dissociation curve, and is crucial for efficient oxygen transport, like a battery charge indicator.
dissociation curve — A graph showing the percentage saturation of a pigment (such as haemoglobin) with oxygen, plotted against the partial pressure of oxygen.
The S-shaped curve illustrates haemoglobin's affinity for oxygen: high affinity at high partial pressures (lungs) for loading, and lower affinity at low partial pressures (tissues) for unloading. The shape reflects cooperative binding, like a graph showing how many seats on a bus are filled at different stops.
The haemoglobin dissociation curve is sigmoidal (S-shaped) due to cooperative binding. When the first oxygen molecule binds to haemoglobin, it causes a conformational change that increases the affinity of the remaining binding sites for oxygen. This ensures efficient loading of oxygen in the high partial pressure environment of the lungs and efficient unloading in the low partial pressure environment of respiring tissues.
Bohr shift — The decrease in affinity of haemoglobin for oxygen that occurs when carbon dioxide is present.
High partial pressures of carbon dioxide (and thus lower pH) cause haemoglobin to release oxygen more readily. This is advantageous in actively respiring tissues where CO2 is high and oxygen is needed, shifting the dissociation curve to the right, like a taxi driver eager to drop off passengers in a busy area.
For full marks on the Bohr shift, state the full sequence: ↑CO₂ → ↑H⁺ (lower pH) → change in haemoglobin's tertiary structure → reduced affinity for O₂ → more O₂ released to respiring tissues.
Students often think carbon dioxide binds to the same site on haemoglobin as oxygen, rather than to different amine groups.
carbonic anhydrase — An enzyme found in the cytoplasm of red blood cells that catalyses the reaction between carbon dioxide and water to form carbonic acid.
This enzyme rapidly converts CO2 into carbonic acid, which then dissociates into hydrogen ions and hydrogencarbonate ions. This reaction is crucial for efficient carbon dioxide transport and buffering blood pH, acting like a super-fast chemical 'mixer'.
Carbon dioxide to carbonic acid
Catalysed by carbonic anhydrase in red blood cells, this reaction is the first step in CO2 transport as hydrogencarbonate ions.
Carbonic acid dissociation
Carbonic acid dissociates into hydrogen ions and hydrogencarbonate ions, contributing to the Bohr shift and CO2 transport.
chloride shift — The movement of chloride ions into red blood cells from blood plasma, to balance the movement of hydrogencarbonate ions into the plasma from the red blood cells.
As hydrogencarbonate ions move out of red blood cells into the plasma for transport, chloride ions move into the red blood cells to maintain electrical neutrality across the cell membrane. This is essential for continuous CO2 transport.
Students often forget that the chloride shift's primary purpose is to maintain electrical neutrality in red blood cells.
carbaminohaemoglobin — A compound formed when carbon dioxide binds with haemoglobin.
A small proportion of carbon dioxide is transported in the blood by binding directly to the amine groups of haemoglobin, forming carbaminohaemoglobin. This binding is reversible and contributes to the overall transport of CO2.
Carbon dioxide is transported in the blood in three main ways: dissolved in plasma, bound to haemoglobin as carbaminohaemoglobin, and most significantly, as hydrogencarbonate ions. In red blood cells, carbonic anhydrase rapidly converts CO2 and water into carbonic acid, which then dissociates into hydrogen ions and hydrogencarbonate ions. The hydrogencarbonate ions diffuse into the plasma, with chloride ions moving into the red blood cells to maintain electrical neutrality (chloride shift).
In explanations of CO₂ transport, explicitly name the enzyme 'carbonic anhydrase' and the process of the 'chloride shift' to score maximum marks.
cardiac muscle — The type of muscle that makes up the walls of the heart.
Cardiac muscle is a specialised type of muscle found only in the heart. It is myogenic, meaning it can contract and relax rhythmically without nervous stimulation, and its continuous, involuntary contractions are essential for pumping blood throughout the body.
coronary arteries — Arteries that branch from the aorta and spread over the walls of the heart, supplying the cardiac muscle with nutrients and oxygen.
The heart muscle itself requires a constant supply of oxygen and nutrients to sustain its continuous pumping action. The coronary arteries ensure this vital supply, branching directly from the aorta to deliver oxygenated blood to the cardiac muscle tissue.
Students often believe the heart muscle gets its oxygen directly from the blood flowing through its chambers, rather than via the coronary arteries.
septum — The layer of tissue that separates the left and right sides of the heart.
The septum is a muscular wall that divides the heart into two distinct halves, preventing the mixing of oxygenated and deoxygenated blood. This separation is crucial for maintaining the efficiency of the double circulatory system in mammals.
atrium — One of the chambers of the heart that receives low-pressure blood from the veins.
The atria are the upper chambers of the heart. The right atrium receives deoxygenated blood from the body, and the left atrium receives oxygenated blood from the lungs. They act as collecting chambers before pumping blood into the ventricles.
ventricle — One of the chambers of the heart that receives blood from the atria and then pushes it into the arteries.
The ventricles are the lower, more muscular chambers of the heart. The right ventricle pumps deoxygenated blood to the lungs, while the left ventricle pumps oxygenated blood to the rest of the body. The left ventricle has a thicker muscular wall to generate higher pressure for systemic circulation.
atrioventricular valve — A valve between the atria and ventricles that closes when the ventricles contract and stops backflow of blood into the atria.
These valves, the bicuspid (mitral) on the left and tricuspid on the right, ensure unidirectional blood flow from the atria to the ventricles. They prevent blood from being forced back into the atria during ventricular contraction.
bicuspid valve — The atrioventricular valve on the left side of the heart.
Also known as the mitral valve, the bicuspid valve is located between the left atrium and the left ventricle. It has two cusps and prevents the backflow of oxygenated blood into the left atrium when the left ventricle contracts.
tricuspid valve — The atrioventricular valve on the right side of the heart.
The tricuspid valve is located between the right atrium and the right ventricle. It has three cusps and prevents the backflow of deoxygenated blood into the right atrium when the right ventricle contracts.
The mammalian heart is a four-chambered muscular organ, divided by a septum into left and right sides. The atria receive blood, while the ventricles pump it out. The left ventricle has a thicker wall to pump blood around the systemic circulation at high pressure, while the right ventricle pumps blood to the pulmonary circulation at lower pressure. Atrioventricular and semilunar valves ensure unidirectional blood flow, preventing backflow during the cardiac cycle.
cardiac cycle — The sequence of events that takes place during one heartbeat.
The cardiac cycle involves a coordinated series of contractions (systole) and relaxations (diastole) of the atria and ventricles. This cycle ensures efficient filling and emptying of the heart chambers, maintaining continuous blood flow throughout the body.
atrial systole — The stage of the cardiac cycle when the muscle in the walls of the atria contracts.
During atrial systole, the atria contract, pushing the remaining blood into the ventricles. This ensures that the ventricles are fully filled before they begin to contract, maximising the efficiency of blood ejection.
ventricular systole — The stage of the cardiac cycle when the muscle in the walls of the ventricles contracts.
Ventricular systole involves the powerful contraction of the ventricular walls, forcing blood out into the pulmonary artery (from the right ventricle) and the aorta (from the left ventricle). During this phase, the atrioventricular valves close to prevent backflow into the atria, and the semilunar valves open.
diastole — The stage of the cardiac cycle when the muscle in the walls of the heart relaxes.
During diastole, all chambers of the heart relax, allowing blood to flow from the veins into the atria and passively into the ventricles. This relaxation phase is crucial for the heart to refill with blood before the next contraction cycle.
When describing the cardiac cycle, be systematic. For each stage, state which chambers are contracting (systole) or relaxing (diastole) and the status (open/closed) of both the atrioventricular and semilunar valves.
myogenic — A word used to describe muscle tissue that contracts and relaxes even when there is no stimulation from a nerve.
The heart's cardiac muscle is myogenic, meaning it generates its own electrical impulses that initiate contraction. This intrinsic ability allows the heart to beat rhythmically and continuously without direct nervous input, though nerve signals can modify the rate.
sinoatrial node (SAN) — A patch of cardiac muscle in the right atrium of the heart which contracts and relaxes in a rhythm that sets the pattern for the rest of the heart muscle.
The SAN is the natural pacemaker of the heart. It generates electrical impulses that spread across the atria, causing them to contract. This rhythmic impulse generation is fundamental to the heart's myogenic activity.
atrioventricular node (AVN) — A patch of tissue in the septum of the heart which transmits the wave of excitation from the walls of the atria and transmits it to the Purkyne tissue.
The AVN receives the electrical impulse from the SAN and introduces a crucial delay before transmitting it to the ventricles. This delay ensures that the atria complete their contraction and empty blood into the ventricles before ventricular contraction begins.
Purkyne tissue — A bundle of fibres that conduct the wave of excitation down through the septum of the heart to the base (apex) of the ventricles.
The Purkyne tissue (also known as the Bundle of His and Purkyne fibres) rapidly conducts the electrical impulse from the AVN down the septum and then upwards through the ventricular walls. This ensures that the ventricles contract from the apex upwards, efficiently pushing blood into the arteries.
Students often confuse the roles of the SAN and AVN, particularly the AVN's crucial delay function.
The AVN does not initiate the heartbeat. The SAN is the pacemaker; the AVN's key role is to introduce a delay before ventricular contraction.
The heartbeat is myogenic, initiated by the sinoatrial node (SAN) in the right atrium, which acts as the pacemaker. The electrical impulse spreads across the atria, causing atrial systole. The impulse then reaches the atrioventricular node (AVN) in the septum, which delays the impulse. This delay allows the atria to fully empty before the impulse is transmitted via the Purkyne tissue down the septum and into the ventricular walls, causing ventricular systole from the apex upwards.
When describing blood vessels, always link structure to function. E.g., 'The thick elastic wall of the artery allows it to stretch and recoil to maintain high pressure'.
Definitions Bank
circulatory system
A system that carries fluids around an organism’s body.
closed blood system
A circulatory system made up of vessels containing blood.
double circulation
A circulatory system in which the blood passes through the heart twice on one complete circuit of the body.
systemic circulation
The part of the circulatory system that carries blood from the heart to all of the body except the gas exchange surface, and then back to the heart.
pulmonary circulation
The part of the circulatory system that carries blood from the heart to the gas exchange surface and then back to the heart.
+43 more definitions
View all →Command Word Guide
| Describe | For blood vessels, describe specific structural features (e.g., wall thickness, presence of elastic/muscle fibres, lumen size, valves) and directly link them to their function. For the cardiac cycle, describe the sequence of events in atria and ventricles, including valve status. |
| Explain | For the Bohr shift, explain the causal chain from increased CO2 to reduced haemoglobin affinity for oxygen. For tissue fluid formation, explain the roles of hydrostatic pressure and water potential gradient. For heartbeat control, explain the roles of SAN, AVN, and Purkyne tissue in sequence. |
| Compare | When comparing arteries and veins, focus on differences in wall structure (thickness, elastic/muscle content), lumen size, and presence of valves, relating these to the pressure of blood carried and direction of flow. |
Common Mistakes
Students often think arteries always carry oxygenated blood and veins always carry deoxygenated blood.
Remember the pulmonary circulation: pulmonary arteries carry deoxygenated blood to the lungs, and pulmonary veins carry oxygenated blood from the lungs to the heart.
Students often confuse the roles of the SAN and AVN.
The SAN initiates the heartbeat (pacemaker), while the AVN's crucial role is to introduce a delay before ventricular contraction, ensuring atria contract first.
Students often think tissue fluid is identical to blood plasma.
Tissue fluid is similar to plasma but contains far fewer large protein molecules and no red blood cells.
+3 more
View all →This chapter details the human gas exchange system, focusing on the structure and function of the airways and alveoli. It explains how air is warmed and cleaned, and how efficient gas exchange occurs in the lungs.
gas exchange surface — Any part of an organism that allows the movement of gases between the surroundings and the body.
A gas exchange surface is like a permeable window, specifically designed for the efficient movement of oxygen into the body and carbon dioxide out. Organisms with small surface area:volume ratios, such as mammals, require specialised surfaces like the lungs to facilitate this process.
alveolus — A small air sac in the lungs composed of a single layer of squamous epithelium and some elastic fibres.
Alveoli are tiny, thin-walled balloons clustered at the end of the airways, each surrounded by a net of capillaries. This structure allows gases to easily pass through their thin walls, providing a huge collective surface area for efficient diffusion between alveolar air and blood.
trachea — The tube-like structure that extends from the larynx to the bronchi.
Also known as the windpipe, the trachea is the main highway for air, ensuring a clear path to the lungs. It is supported by C-shaped rings of cartilage, which prevent it from collapsing while allowing some flexibility.
bronchus — A major branch of the trachea that extends into the lungs.
The trachea divides into two bronchi, which are like major exit ramps leading into the lungs. These then subdivide extensively, forming a bronchial 'tree' and containing irregular blocks of cartilage for support.
bronchiole — A microscopic branch of a bronchus that leads to the alveoli.
Bronchioles are smaller than bronchi and lack cartilage, acting like small residential streets leading directly to the alveoli. They are surrounded by smooth muscle, which can adjust their diameter to regulate airflow.
Students often confuse bronchi with bronchioles. Remember that bronchi are larger, have cartilage blocks, and branch directly from the trachea, while bronchioles are smaller and lack cartilage.
The human gas exchange system begins with the trachea, which branches into two bronchi. These bronchi further subdivide into progressively smaller bronchioles, ultimately leading to the alveoli. This branching structure ensures air reaches the vast surface area required for efficient gas exchange.
cartilage — A type of skeletal tissue that is strong and flexible and supports the larynx, trachea and bronchi in the gas exchange system.
Cartilage acts like a flexible but sturdy scaffolding for the main airways, ensuring they remain open. In the trachea, it forms C-shaped rings, and in the bronchi, irregular blocks, preventing collapse or bursting due to pressure changes during breathing.
Distinguish between the cartilage structure in the trachea (C-shaped rings) and bronchi (irregular blocks) in your descriptions.
goblet cell — A cell shaped like a drinking goblet that secretes mucus.
Goblet cells are tiny, specialised factories that continuously produce and release sticky mucus. This mucus, containing mucin droplets, traps inhaled particles like dust, pollen, bacteria, and viruses, preventing them from reaching the delicate lung tissues.
mucin — Any glycoprotein that forms part of the mucus secreted by goblet cells and mucous cells.
Mucin is the key sticky ingredient in mucus, composed of glycoproteins with many carbohydrate chains. This property enables mucus to effectively trap inhaled particles, preventing them from reaching the delicate lung tissues.
ciliated epithelium — An epithelium that consists mainly of ciliated cells.
Ciliated epithelium, often containing goblet cells, acts like a moving carpet or escalator. The cilia beat continually to sweep the sticky mucus layer, along with trapped particles, upwards towards the larynx to be swallowed, thus cleaning the airways.
The airways are adapted for warming and cleaning inhaled air. Goblet cells and mucous glands produce sticky mucus, which traps foreign particles. Ciliated epithelial cells then use their coordinated beating action to move this mucus, along with the trapped debris, out of the lungs, maintaining the health of the gas exchange system.
Students often think goblet cells are primarily for protection against pathogens. Remember their main role is to produce mucus that traps particles, which are then removed by cilia.
Students often think cilia are only for movement. In the gas exchange system, their primary role is to move mucus, not to propel the cell itself.
elastic fibres — Bundles of the fibrous protein elastin which can stretch and recoil like elastic bands.
Elastic fibres are like tiny rubber bands embedded in the walls of the alveoli. They allow the alveoli to stretch during inspiration and then passively recoil during expiration, helping to force air out of the lungs.
Students often think elastic fibres actively contract. Remember they passively recoil after being stretched, similar to a stretched rubber band returning to its original shape.
Students often think alveoli are rigid structures. Actually, their elastic fibres allow them to stretch during inspiration and recoil during expiration.
Gas exchange occurs efficiently in the alveoli. These tiny air sacs are adapted with a large collective surface area, very thin walls (composed of a single layer of squamous epithelium), and a rich blood supply from surrounding capillaries. These features work together to maintain steep concentration gradients for oxygen and carbon dioxide, facilitating rapid diffusion.
Students often think gas exchange surfaces are only for oxygen intake. Remember they are also crucial for carbon dioxide removal.
When asked about alveolar adaptations, mention the thin walls (squamous epithelium), large collective surface area, and rich blood supply (capillaries).
Always link structure to function. For example, 'The single layer of squamous epithelium in the alveoli provides a short diffusion path for gases'.
When describing gas exchange, always mention the maintenance of a steep concentration gradient by breathing and blood flow.
For microscope drawing questions, be sure to accurately place tissues. Show cartilage in the trachea/bronchus but NOT in a bronchiole.
Use precise terminology: 'squamous epithelium', 'ciliated epithelium', 'goblet cells', 'elastic recoil' will score more marks than vague descriptions.
Be ready for 'compare and contrast' questions. Know the key structural differences between the trachea, bronchi, and bronchioles regarding cartilage, smooth muscle, and epithelial cells.
Definitions Bank
gas exchange surface
Any part of an organism that allows the movement of gases between the surroundings and the body.
alveolus
A small air sac in the lungs composed of a single layer of squamous epithelium and some elastic fibres.
trachea
The tube-like structure that extends from the larynx to the bronchi.
bronchus
A major branch of the trachea that extends into the lungs.
bronchiole
A microscopic branch of a bronchus that leads to the alveoli.
+5 more definitions
View all →Command Word Guide
| Describe | Provide detailed structural features, e.g., 'Describe the structure of the trachea' requires mentioning C-shaped cartilage rings, ciliated epithelium, and goblet cells. |
| Explain | Give reasons for observed structures or processes, linking structure to function. E.g., 'Explain how the alveoli are adapted for gas exchange' requires linking thin walls to short diffusion distance, large surface area to efficient diffusion, and rich blood supply to maintaining concentration gradients. |
| Compare | Identify similarities and differences between two or more structures. E.g., 'Compare the structure of a bronchus and a bronchiole' requires mentioning cartilage presence/absence, size, and smooth muscle. |
Common Mistakes
Thinking gas exchange surfaces are only for oxygen intake.
Gas exchange surfaces are crucial for both oxygen intake and carbon dioxide removal.
Thinking alveoli are rigid structures.
Alveoli are highly elastic due to elastic fibres, allowing them to stretch during inspiration and recoil during expiration.
Confusing bronchi with bronchioles.
Bronchi are larger, have cartilage blocks, and branch directly from the trachea. Bronchioles are smaller, lack cartilage, and have smooth muscle for diameter regulation.
+2 more
View all →This chapter explores infectious diseases, which are caused by pathogens transmitted between individuals. It details specific diseases like cholera, malaria, HIV/AIDS, and tuberculosis, covering their pathogens, transmission, and control measures, while also examining the action of antibiotics and the critical issue of antibiotic resistance.
infectious disease — A disease caused by an organism such as a protoctist, bacterium or virus.
These diseases, also known as communicable diseases, can spread from an infected person to an uninfected person, or from animals to humans. They are distinct from non-infectious diseases, which are not caused by pathogens. Like a computer virus spreading from one computer to another, an infectious disease spreads from one host to another.
Students often think 'disease' always implies something very serious, but actually many mild conditions like the common cold are also infectious diseases.
pathogen — An organism that causes disease.
Pathogens can be viruses, bacteria, protoctists, or fungi. They invade a host organism and disrupt its normal physiological functions, leading to illness. Understanding the specific pathogen is crucial for effective treatment and control. Think of a pathogen as a saboteur that infiltrates a system (the body) and causes it to malfunction.
Students often think all microorganisms are pathogens, but actually most microorganisms are harmless or beneficial; only those that cause disease are pathogens.
When asked to define 'infectious disease', ensure you include 'caused by an organism' or 'pathogen' and 'transmitted' to gain full marks.
disease transmission — The transfer of a pathogen from a person infected with that pathogen to an uninfected person; transmission may occur by direct contact, through the air or water or by animal vectors, such as insects.
Transmission can be direct (e.g., sexual contact, airborne droplets) or indirect (e.g., contaminated food/water, vectors). Breaking the transmission cycle is a key strategy in disease control and eradication efforts. Imagine a relay race where the baton (pathogen) is passed from one runner (host) to the next, either directly or via an intermediary.
Students often think transmission always requires direct contact, but actually many pathogens can survive in the environment or be carried by vectors, leading to indirect transmission.
When describing transmission, specify the method (e.g., 'water-borne', 'insect vector', 'direct contact') and the specific agent involved (e.g., 'female Anopheles mosquito', 'faeces').
disease carrier — Person infected with a pathogen who shows no symptoms, but can be the source of infection in other people (not carrier of an inherited disease).
Carriers are particularly problematic for disease control because their lack of symptoms makes them difficult to identify and isolate. They can unknowingly spread the pathogen, contributing to wider outbreaks. A disease carrier is like a hidden messenger, delivering a harmful package (pathogen) without showing any outward signs of carrying it.
disease vector — An organism which carries a pathogen from one person to another or from an animal to a human.
Vectors are typically arthropods like mosquitoes or ticks, which transmit pathogens during feeding. They are not necessarily infected themselves but act as intermediaries. Controlling vectors is a crucial strategy for preventing vector-borne diseases. A disease vector is like a delivery service that transports a package (pathogen) from one sender (infected host) to a recipient (uninfected host).
Distinguish between a disease carrier (infectious but asymptomatic) and a vector (an organism that transmits the pathogen but is not necessarily infected itself).
transmission cycle — The passage of a pathogen from one host to another is continually repeated as the pathogen infects new hosts.
This cycle describes the continuous process of a pathogen moving between hosts, ensuring its survival and spread. Control methods aim to interrupt this cycle at various points, such as by vaccinating hosts or eliminating vectors. Think of a continuous loop or a chain reaction where each infection leads to the potential for more infections, perpetuating the pathogen's existence.
disease eradication — The complete breakage of the transmission cycle of a pathogen so that there are no more cases of the disease caused by the pathogen anywhere in the world.
Eradication is the ultimate goal for some diseases, meaning the pathogen no longer exists in nature. This is a rare achievement, requiring highly effective control measures and global coordination. Eradication is like completely deleting a harmful file from every computer system globally, ensuring it can never reappear.
endemic disease — A disease that is always in a population.
An endemic disease is consistently present at a predictable level within a specific geographical area or population. Its continued presence is maintained by ongoing transmission within that population. Malaria in tropical regions is an example. Like a background noise that is always present in a particular environment, an endemic disease is a constant presence in a population.
Human pathogens can be viruses, bacteria, and protoctists. Cholera, caused by the bacterium *Vibrio cholerae*, is transmitted via contaminated water. Malaria, caused by the protoctist *Plasmodium*, is transmitted by a vector, specifically the female *Anopheles* mosquito. Tuberculosis (TB), caused by the bacterium *Mycobacterium tuberculosis*, spreads through airborne droplets. HIV, a virus, is transmitted through the exchange of body fluids.
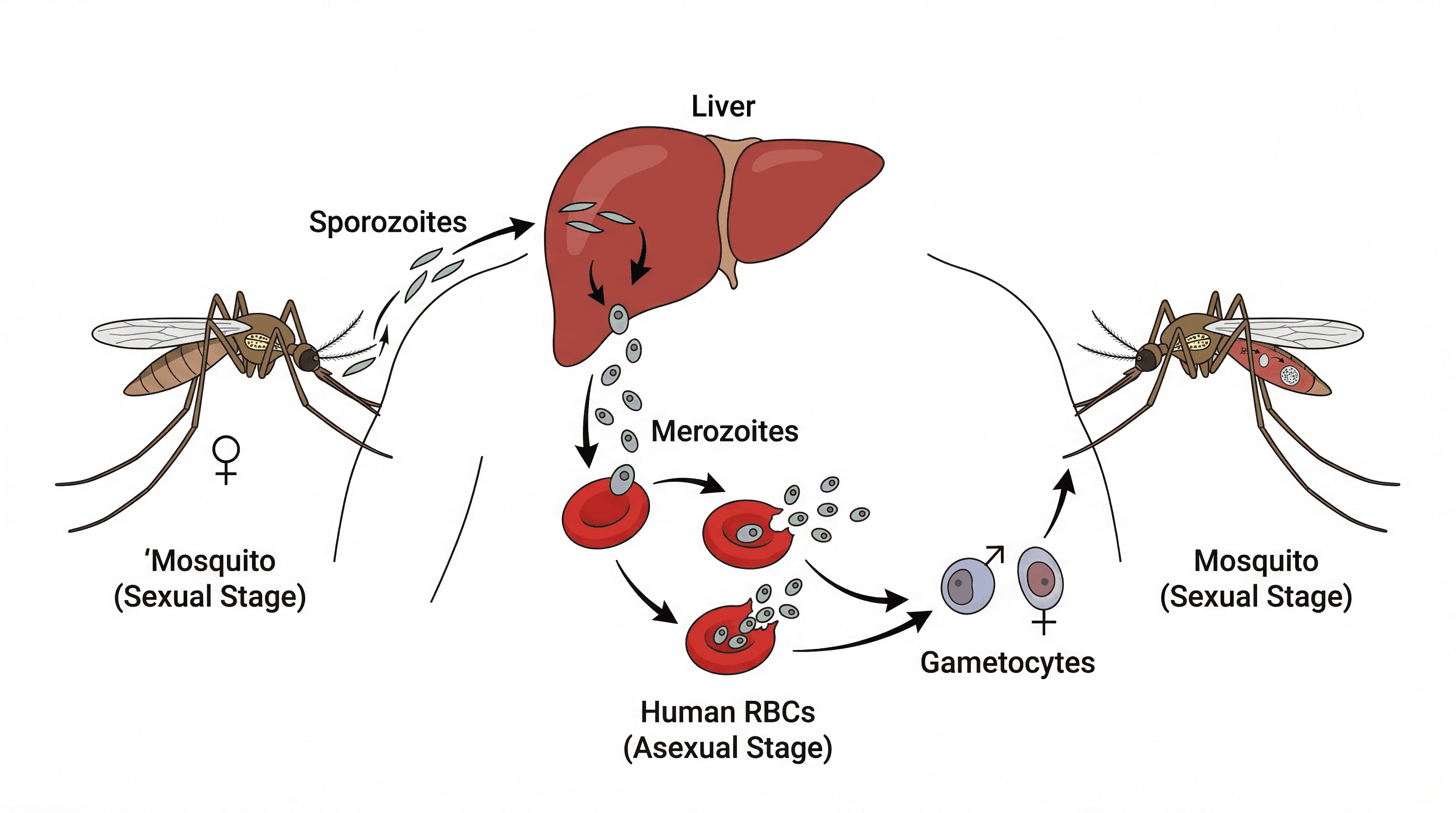
HIV — Human immunodeficiency virus.
HIV is a retrovirus that primarily infects and destroys T-helper lymphocytes, weakening the immune system. It is the causative agent of AIDS. Its genetic material is RNA, which is reverse transcribed into DNA upon infection. HIV is like a saboteur that specifically targets and dismantles the command center (T-helper lymphocytes) of the body's defense system.
AIDS — Acquired immunodeficiency syndrome.
AIDS is a collection of opportunistic diseases and conditions that occur when the immune system is severely weakened by HIV infection. It is not a single disease but a syndrome characterized by a low count of T-helper lymphocytes, making the body vulnerable to various infections and cancers. AIDS is like a castle whose walls have been so severely damaged (by HIV) that it can no longer defend itself against various invaders (opportunistic infections).
Students often think HIV and AIDS are the same thing, but actually HIV is the virus that causes the infection, and AIDS is the syndrome (collection of opportunistic infections) that develops as a result of advanced HIV infection.
Ensure you differentiate between HIV (the virus) and AIDS (the syndrome) and understand that HIV infection leads to AIDS over time as the immune system deteriorates.
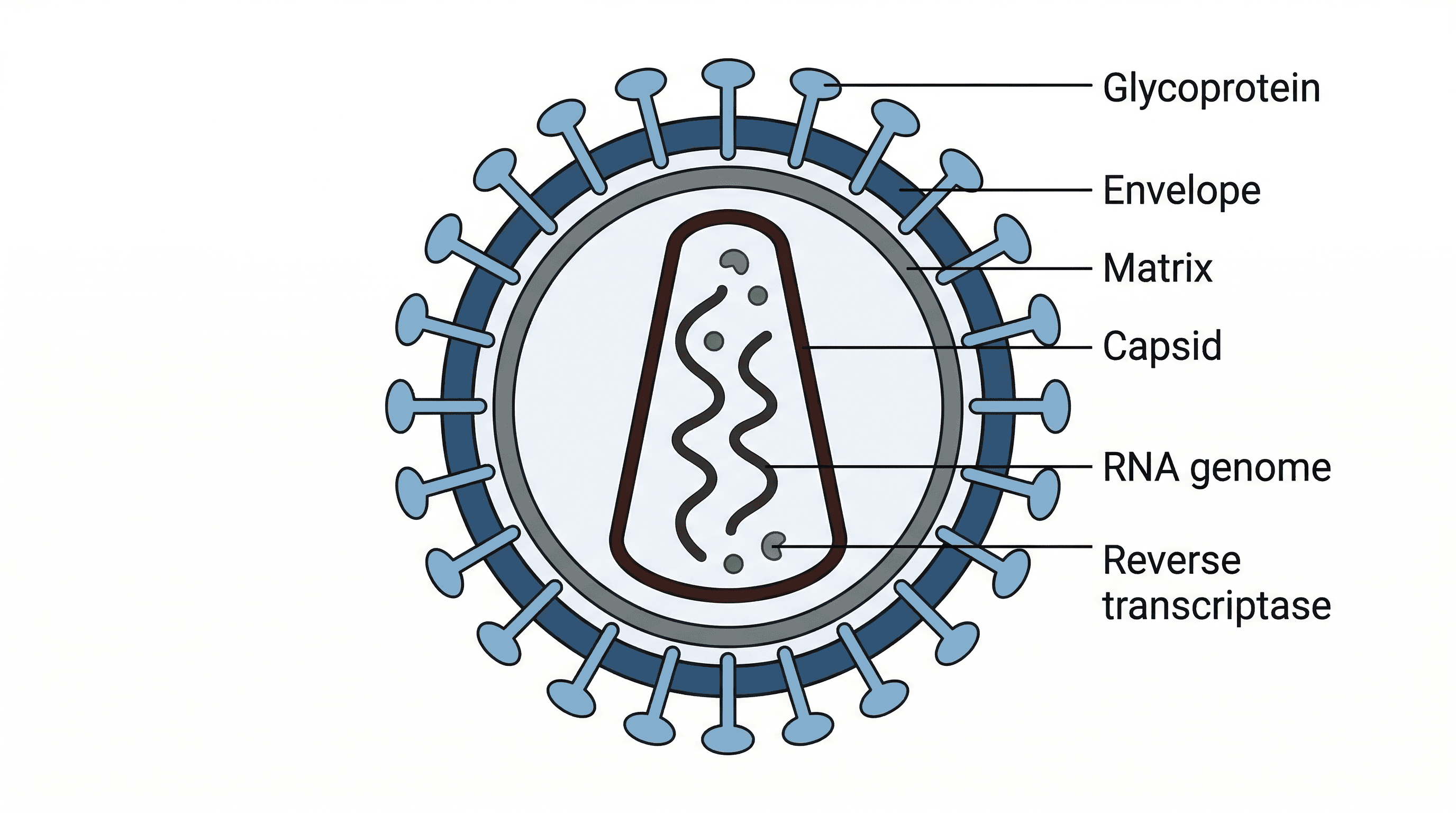
opportunistic infection — An infection caused by pathogens that take advantage of a host with a weakened immune system, as may happen in someone with an HIV infection.
These infections are typically harmless in individuals with healthy immune systems but can become severe or life-threatening when the host's defenses are compromised. They are a hallmark of AIDS, as HIV destroys T-helper lymphocytes, leaving the body vulnerable. Imagine a city with strong defenses (immune system) that normally keeps petty criminals (opportunistic pathogens) in check. If the defenses collapse, these criminals can run rampant and cause serious damage.
The effectiveness of control measures for diseases like cholera, malaria, TB, and HIV is influenced by biological, social, and economic factors. Biological factors include pathogen characteristics and drug resistance. Social factors involve public health education and community engagement. Economic factors relate to the cost of treatments, preventative infrastructure, and access to healthcare. Control measures aim to interrupt the transmission cycle, for example, by providing clean water for cholera, vector control for malaria, vaccination for TB, and safe practices for HIV.
antibiotic — A substance derived from a living organism that is capable of killing or inhibiting the growth of a microorganism.
Antibiotics are drugs specifically targeting bacteria by interfering with their unique cellular processes, such as cell wall synthesis or protein synthesis. They are ineffective against viruses because viruses lack these bacterial targets. An antibiotic is like a specialized weapon that targets specific vulnerabilities in bacterial cells, leaving human cells unharmed.
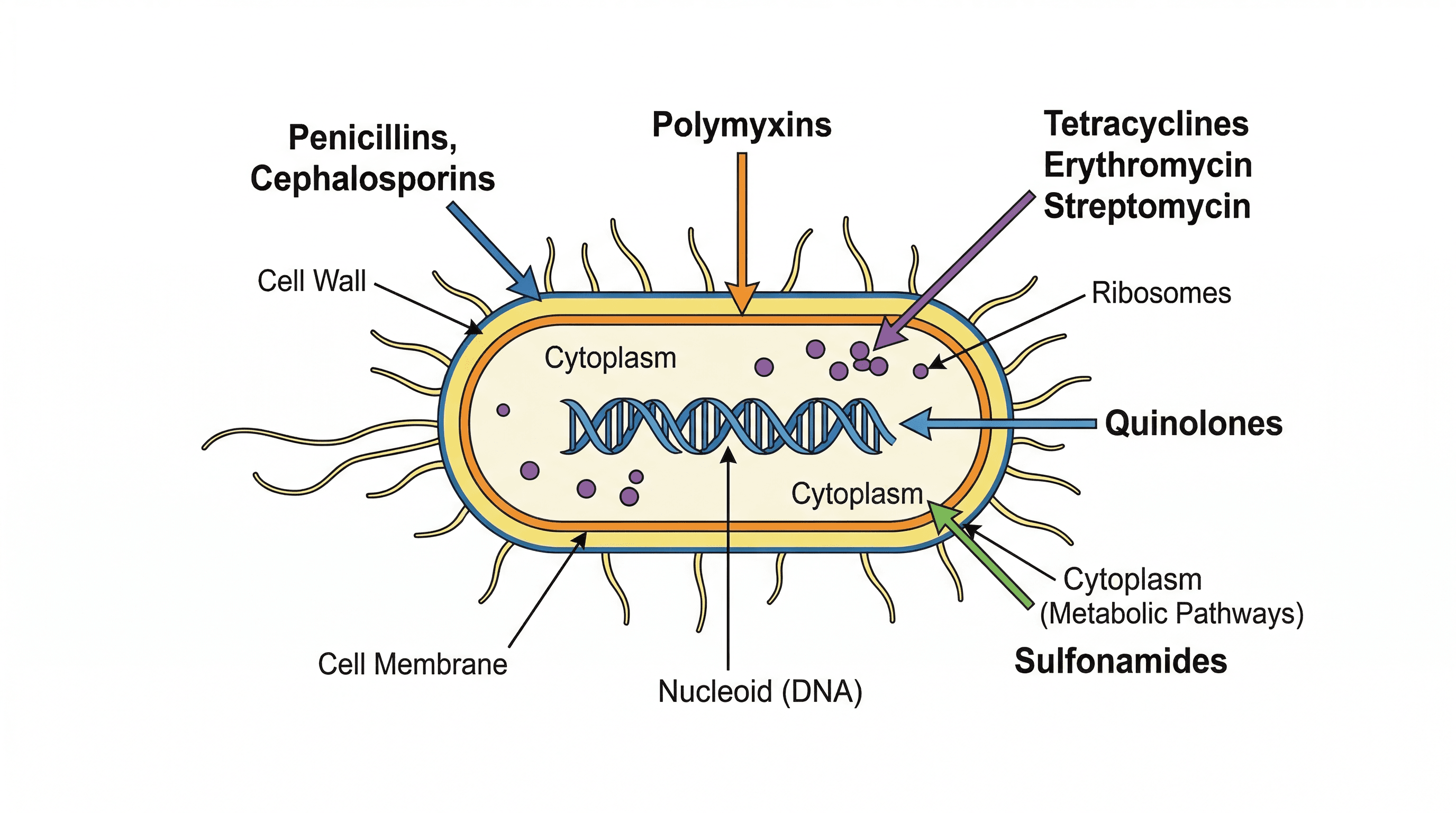
Antibiotics, such as penicillin, act on bacteria by interfering with specific bacterial structures or metabolic pathways. For instance, penicillin inhibits the synthesis of bacterial cell walls, leading to cell lysis. Viruses, however, lack cell walls and other metabolic machinery targeted by antibiotics, as they rely on host cells for replication. Therefore, antibiotics have no effect on viruses.
Students often think antibiotics kill all microorganisms, but actually they are primarily effective against bacteria and have no effect on viruses or fungi.
When explaining antibiotic action, specify the bacterial target (e.g., cell wall synthesis, protein synthesis) and explicitly state why they do not affect human cells or viruses.
antibiotic resistance — The ability of bacteria or fungi to grow in the presence of an antibiotic that would normally slow their growth or kill them; antibiotic resistance arises by mutation and becomes widespread when antibiotics are overused.
Resistance develops through genetic mutations in bacteria, allowing them to survive exposure to antibiotics. These resistant bacteria then reproduce, leading to a population dominated by resistant strains. Overuse and misuse of antibiotics accelerate this process, posing a significant public health threat. Antibiotic resistance is like a lock-picking skill that some bacteria develop, allowing them to bypass the 'lock' (antibiotic) that was designed to stop them.
Students often think individuals become resistant to antibiotics, but actually it is the bacteria that become resistant, not the human host.
The consequences of antibiotic resistance are severe, leading to infections that are harder to treat, longer hospital stays, and increased mortality. Drug-resistant TB is a significant concern. To reduce the impact of antibiotic resistance, measures include reducing the overuse and misuse of antibiotics, completing full courses of antibiotics, developing new antibiotics, and improving hygiene to prevent infections in the first place. This process is driven by natural selection, where antibiotics act as a selective pressure, favoring resistant mutants.
Explain antibiotic resistance using the principles of natural selection: variation (random mutation), selection pressure (the antibiotic), and inheritance (resistant bacteria survive and reproduce).
For any named disease, structure your answer using: Pathogen, Mode of Transmission, Treatment, and Prevention methods. When discussing prevention, always link the method directly to breaking the pathogen's transmission cycle.
Definitions Bank
infectious disease
A disease caused by an organism such as a protoctist, bacterium or virus.
pathogen
An organism that causes disease.
disease transmission
The transfer of a pathogen from a person infected with that pathogen to an uninfected person; transmission may occur by direct contact, through the air or water or by animal vectors, such as insects.
disease carrier
Person infected with a pathogen who shows no symptoms, but can be the source of infection in other people (not carrier of an inherited disease).
transmission cycle
The passage of a pathogen from one host to another is continually repeated as the pathogen infects new hosts.
+8 more definitions
View all →Command Word Guide
| Explain | Provide reasons or mechanisms. For example, 'Explain how antibiotics work' requires detailing their specific bacterial targets. 'Explain why antibiotics don't affect viruses' requires mentioning the absence of bacterial targets in viruses. |
| Discuss | Present a balanced argument or explore various aspects of a topic. For example, 'Discuss the biological, social, and economic factors influencing disease control' requires addressing each type of factor with relevant examples. |
| Outline | Give a brief summary of the main points. For example, 'Outline measures to reduce antibiotic resistance' requires listing key strategies without extensive detail. |
| Give the names of | State the specific scientific names of pathogens or vectors. For example, *Vibrio cholerae* for cholera, *Plasmodium* for malaria, *Anopheles* mosquito as the vector. |
Common Mistakes
Confusing HIV and AIDS.
HIV is the virus that causes the infection, and AIDS is the syndrome (collection of opportunistic infections) that develops as a result of advanced HIV infection.
Assuming antibiotics work on all microorganisms.
Antibiotics are primarily effective against bacteria and have no effect on viruses or fungi.
Thinking all disease transmission requires direct contact.
Many pathogens can survive in the environment or be carried by vectors, leading to indirect transmission (e.g., water-borne cholera, insect-borne malaria).
+3 more
View all →This chapter explores the human immune system, detailing cellular defences like phagocytes and lymphocytes, and how antigens trigger specific primary and secondary immune responses. It covers the production of antibodies and memory cells for long-term immunity, differentiates various types of immunity, and explains the role of vaccination and monoclonal antibodies in public health and medicine.
immune system — The body’s internal defence system.
This system comprises various cells and molecules that work together to protect the body from pathogens and foreign substances. It distinguishes between self and non-self to mount specific responses, acting like the body's security force with different units and weapons to identify and neutralise threats.
antigen — A substance that is foreign to the body and stimulates an immune response (e.g. any large molecule such as a protein).
Antigens are typically large molecules found on the surface of pathogens or released by them. The immune system recognises these as non-self and produces antibodies or activates T cells against them, much like a unique 'ID badge' on a foreign invader that immune cells recognise as a threat.
Students often think antigens are always pathogens, but actually antigens are specific molecules (e.g., proteins, polysaccharides) that can be part of a pathogen or even a non-pathogenic foreign substance.
self — Refers to substances produced by the body that the immune system does not recognise as foreign, so they do not stimulate an immune response.
The immune system undergoes a maturation process where lymphocytes that react to self-antigens are typically destroyed or inactivated. This prevents the immune system from attacking the body's own cells and tissues. Self-antigens are like the 'uniform' worn by your body's own cells, which the immune system recognises as belonging and therefore doesn't attack.
When discussing self/non-self, link it to the concept of immune tolerance and the prevention of autoimmune diseases.
non-self — Refers to any substance or cell that is recognised by the immune system as being foreign and will stimulate an immune response.
These substances, often antigens, trigger the activation of lymphocytes and the production of antibodies or killer cells. This recognition is crucial for defence against pathogens and foreign tissues. Non-self antigens are like 'enemy uniforms' that the immune system identifies as threats, prompting an attack.
immune response — The complex series of responses of the body to the entry of a foreign antigen; it involves the activity of lymphocytes and phagocytes.
This involves the coordinated action of B-lymphocytes (producing antibodies) and T-lymphocytes (killing infected cells or coordinating the response), often with the help of phagocytes. It leads to the elimination of the antigen and the development of immunological memory, much like a coordinated military operation.
The immune system employs various cellular defences. Phagocytes, including neutrophils and macrophages, are crucial for non-specific immunity. They engulf and destroy pathogens through a process called phagocytosis. Macrophages, after engulfing pathogens, also play a vital role in antigen presentation, displaying pathogen antigens on their surface to activate T-lymphocytes.
Be precise when describing phagocytosis: use key terms like phagosome, lysosome, phagolysosome, and hydrolytic enzymes in a clear sequence.
Lymphocytes are key players in specific immunity. B-lymphocytes are responsible for humoral immunity, primarily producing antibodies. T-lymphocytes mediate cell-mediated immunity and include T-helper cells and T-killer cells. The immune response involves the activity of both lymphocytes and phagocytes working in a coordinated manner.
clonal selection — Individual lymphocytes have cell surface receptors specific to one antigen; this specificity is determined as lymphocytes mature and before any antigens enter the body (during an immune response the only lymphocytes to respond are those with receptors specific to antigens on the surface of the invading pathogen).
When an antigen enters the body, it 'selects' the specific B or T lymphocytes that have complementary receptors. This ensures that only the relevant lymphocytes are activated to proliferate, much like finding a specific key that fits only one particular lock (antigen).
Emphasise that specificity is pre-determined before antigen exposure, and the antigen merely 'selects' the appropriate clone.
clonal expansion — The increase in number of specific clones of lymphocytes by mitosis during an immune response.
Once a specific lymphocyte is selected by an antigen, it undergoes rapid mitotic division to produce a large number of identical cells (a clone). This ensures a sufficient number of effector cells to combat the infection, similar to rapidly making many copies of a key to open all the locks quickly.
Students often confuse clonal selection and clonal expansion, but actually selection is the initial recognition, and expansion is the subsequent proliferation.
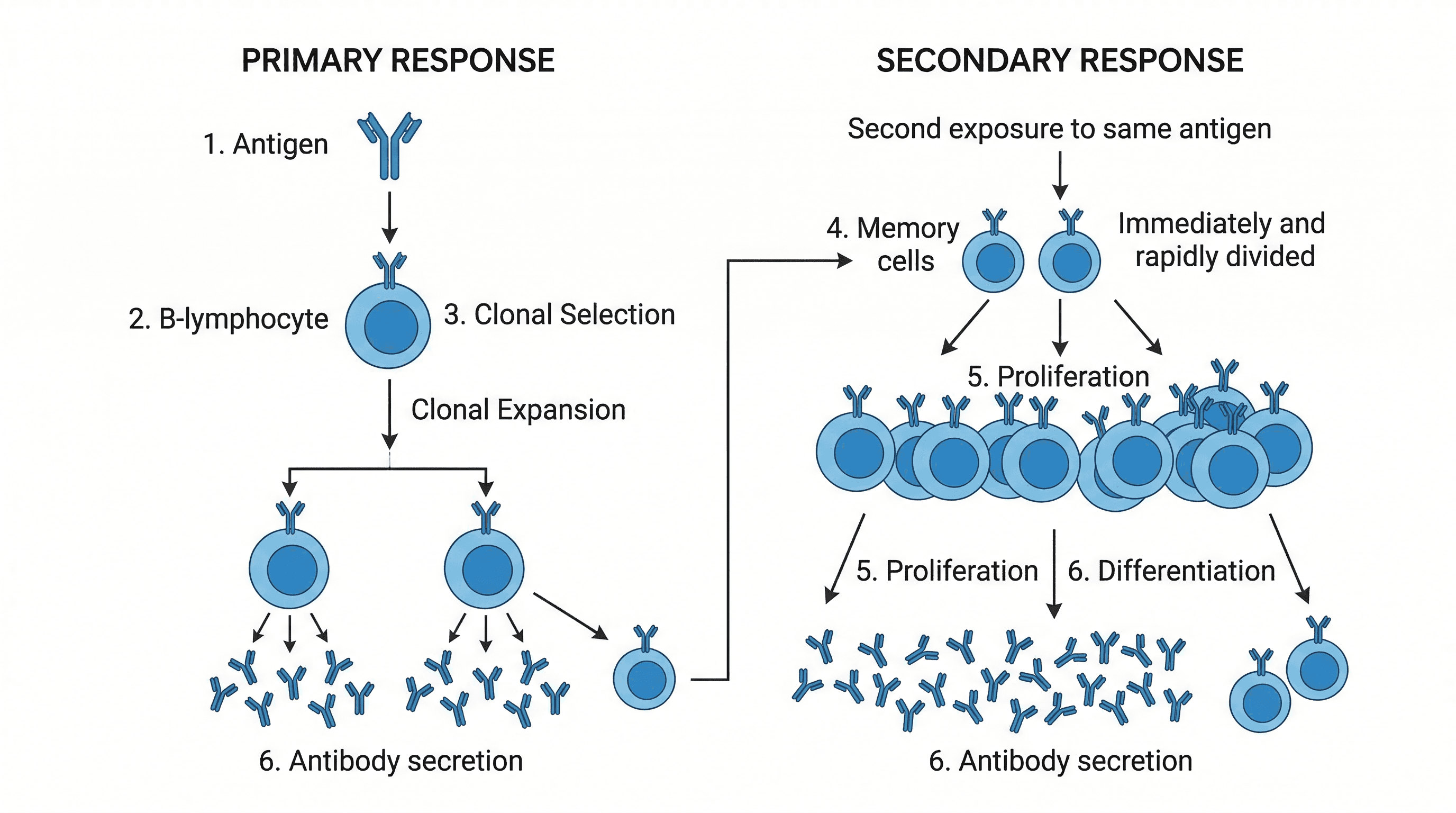
plasma cell — Short-lived, activated B-lymphocyte produced during clonal expansion; plasma cells produce and release antibody molecules.
These cells are highly specialised for antibody production, possessing extensive rough endoplasmic reticulum and mitochondria. They secrete large quantities of specific antibodies into the blood and lymph to combat infection, acting like antibody 'factories'.
Students often think B cells directly produce antibodies, but actually activated B cells differentiate into plasma cells which then secrete antibodies.
memory B cell — Long-lived, activated B-lymphocyte that is specific to one antigen; memory cells are activated to differentiate (develop) into plasma cells during secondary immune responses to the specific antigen.
These cells persist in the body for a long time after the primary infection. Upon re-exposure to the same antigen, they rapidly divide and differentiate into plasma cells and more memory cells, leading to a faster and stronger secondary immune response. They are like the 'veterans' of the immune system.
primary immune response — The first immune response to a specific antigen.
This response is relatively slow and produces a lower concentration of antibodies because there are few specific lymphocytes initially. It takes time for clonal selection and expansion to generate enough effector cells, much like the first time your body encounters a new enemy.
secondary immune response — The second and any subsequent immune responses to a specific antigen.
This response is much faster, stronger, and produces a higher concentration of antibodies due to the presence of memory cells. These memory cells rapidly differentiate into plasma cells upon re-exposure to the antigen, acting like a rapid, pre-planned counter-attack.
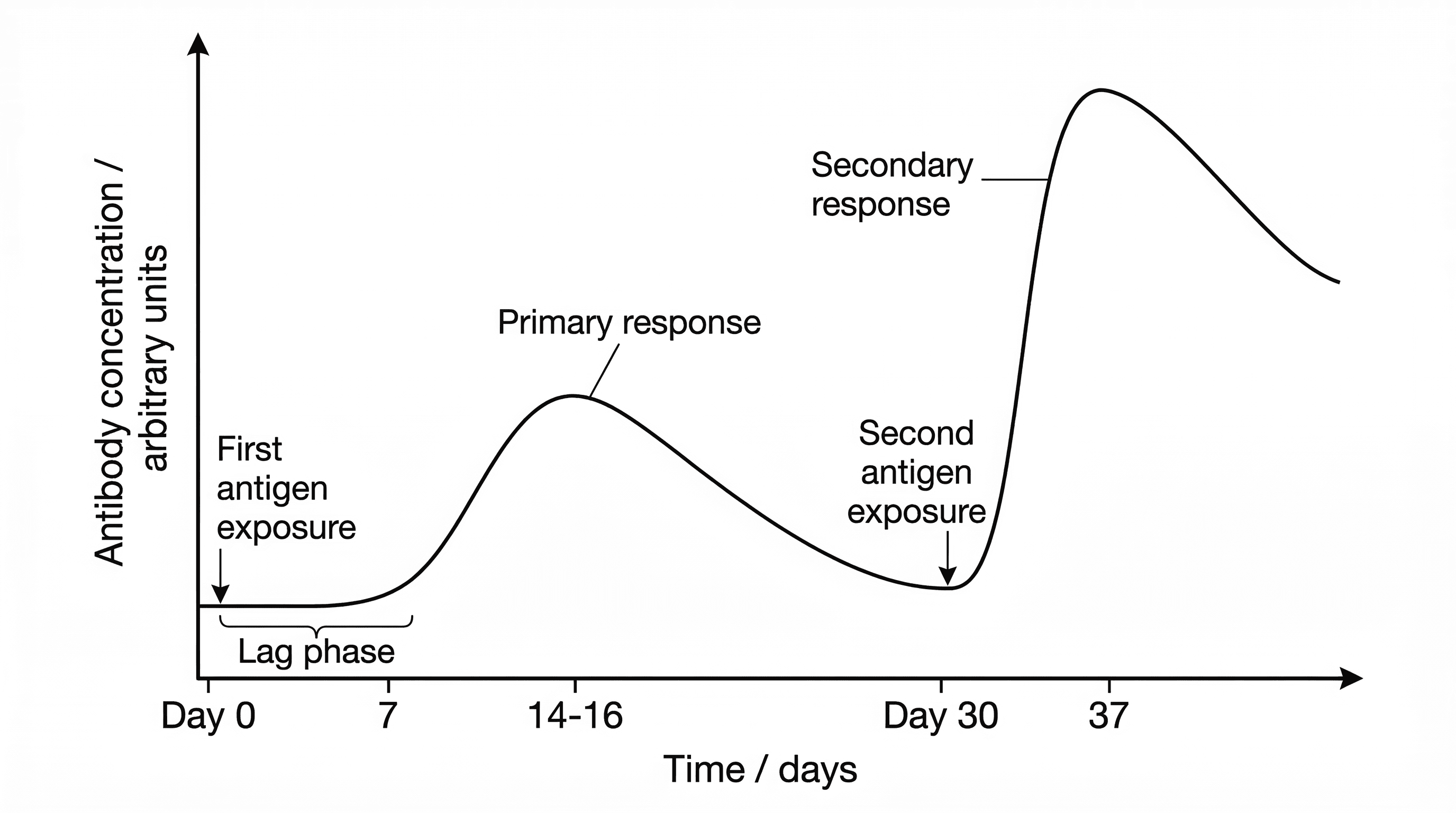
When describing the immune response, always distinguish between the primary and secondary response. Link the secondary response's speed and magnitude to the presence of memory cells.
immunological memory — The ability of the immune system to mount a larger and more rapid response to an antigen that has already been encountered before.
This is the basis of long-term immunity and vaccination. It is mediated by long-lived memory B and T cells that remain in the body after the primary infection, allowing for a swift and effective secondary response, like having a 'most wanted' list for previously encountered criminals.
antibody — A glycoprotein (immunoglobulin) made by specialised lymphocytes in response to the presence of a specific antigen; each type of antibody molecule has a shape that is complementary to its specific antigen.
Antibodies are Y-shaped proteins with two identical antigen-binding sites. They bind to specific antigens, marking pathogens for destruction, neutralising toxins, or preventing viral entry into cells, much like a specific 'key' that fits only one particular 'lock' (antigen).
Students often think antibodies directly kill pathogens, but actually they primarily mark pathogens for destruction by other immune cells or neutralise toxins.
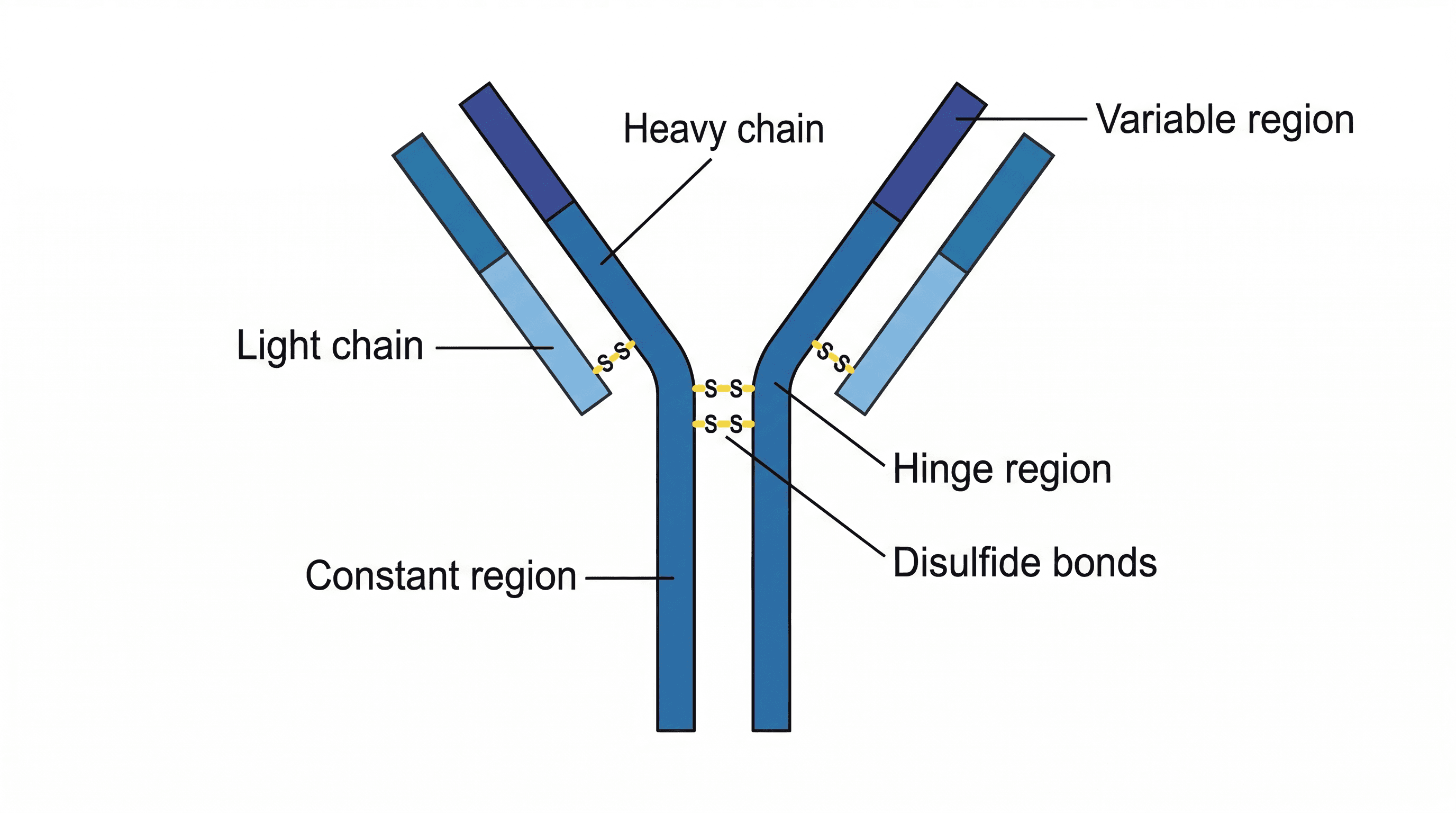
variable region — Region of an antibody molecule composed of parts of the light and heavy polypeptide chains that form the antigen-binding site; the amino acid sequences of the variable site form a specific shape that is complementary to a particular antigen.
This region is highly diverse among different antibody molecules, allowing each antibody to bind specifically to a unique antigen. The precise 3D shape of the variable region determines its antigen specificity, much like the unique 'teeth' of a specific key.
When explaining antibody function, use specific terms like 'agglutination' or 'neutralisation' and state that the variable region determines antigen specificity.
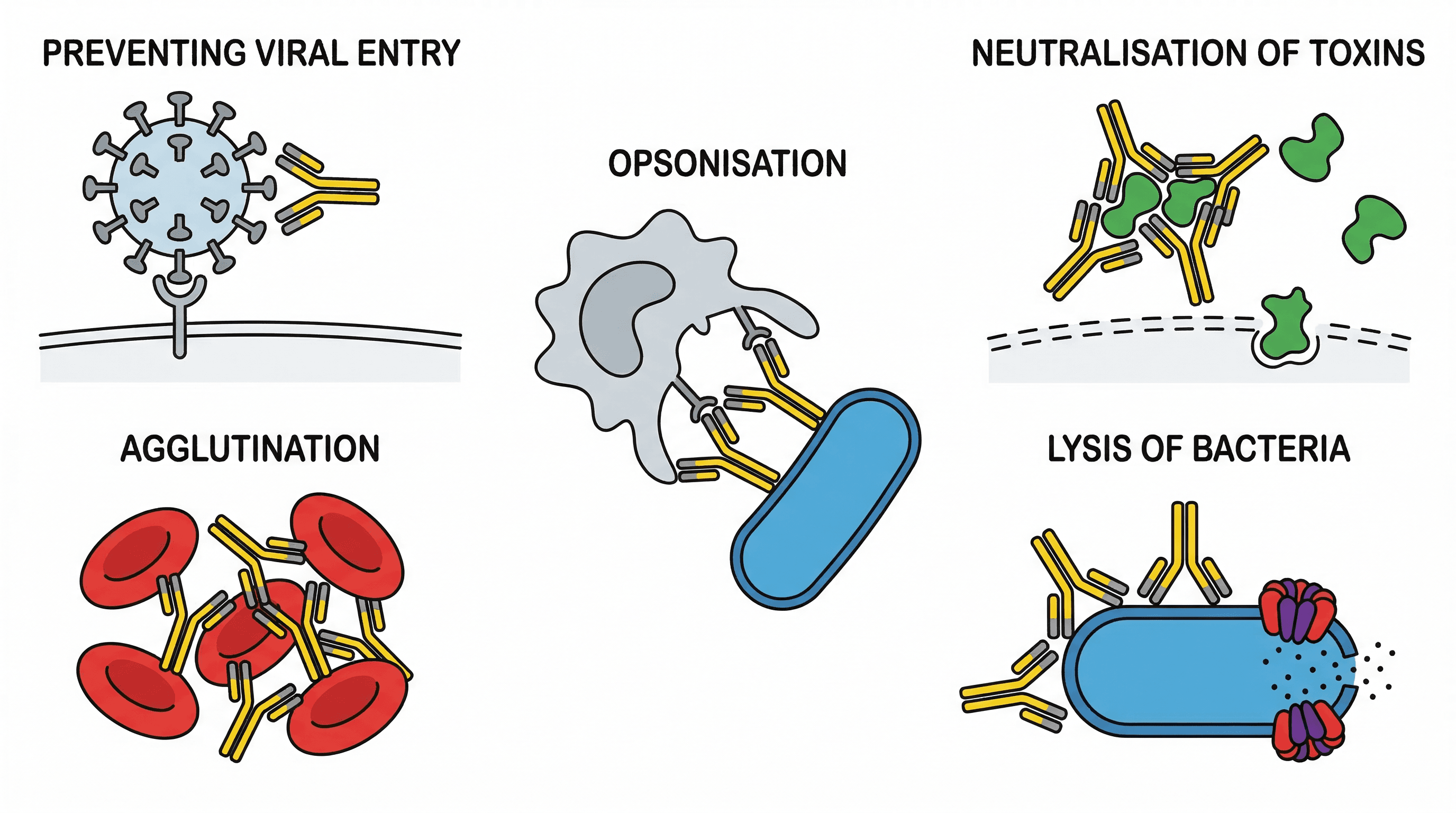
antigen presentation — The process of preparing antigens and exposing them on the surface of host cells (e.g. macrophages) for recognition by T-lymphocytes.
Macrophages, after engulfing pathogens, process their antigens and display them on their cell surface membranes. This allows T-helper cells to recognise the antigen and initiate a coordinated immune response, like a 'wanted poster' displayed by immune cells.
T-helper cell — Type of T-lymphocyte that secretes cytokines to coordinate activity during immune responses.
Upon activation by antigen-presenting cells, T-helper cells release cytokines that stimulate B cells to divide and differentiate into plasma cells, and also enhance the activity of macrophages and T-killer cells. They are like the 'commanders' of the immune system.
T-killer cell — Type of T-lymphocyte that attaches to cells, releasing toxic substances to kill infected cells and cancer cells.
T-killer cells recognise foreign antigens displayed on the surface of infected body cells or cancer cells. They then bind to these cells and release toxic substances, causing the target cells to die, thus eliminating the pathogens within. They are like the 'assassins' of the immune system.
Specify that T-killer cells target and kill *infected host cells* or *cancer cells*, not free pathogens.
cytokine — Any signalling molecule released by cells to influence the growth and/or differentiation of the same or another cell.
Cytokines are a diverse group of small proteins that act as chemical messengers between immune cells. They play crucial roles in regulating the intensity and duration of immune responses, cell proliferation, and differentiation, much like 'walkie-talkies' for immune cells.
Immunity can be broadly categorised into active and passive immunity, and further into natural and artificial types. Active immunity involves the body's own immune system producing antibodies and memory cells, providing long-term protection. Passive immunity involves receiving pre-formed antibodies, offering immediate but temporary protection without memory cell formation.
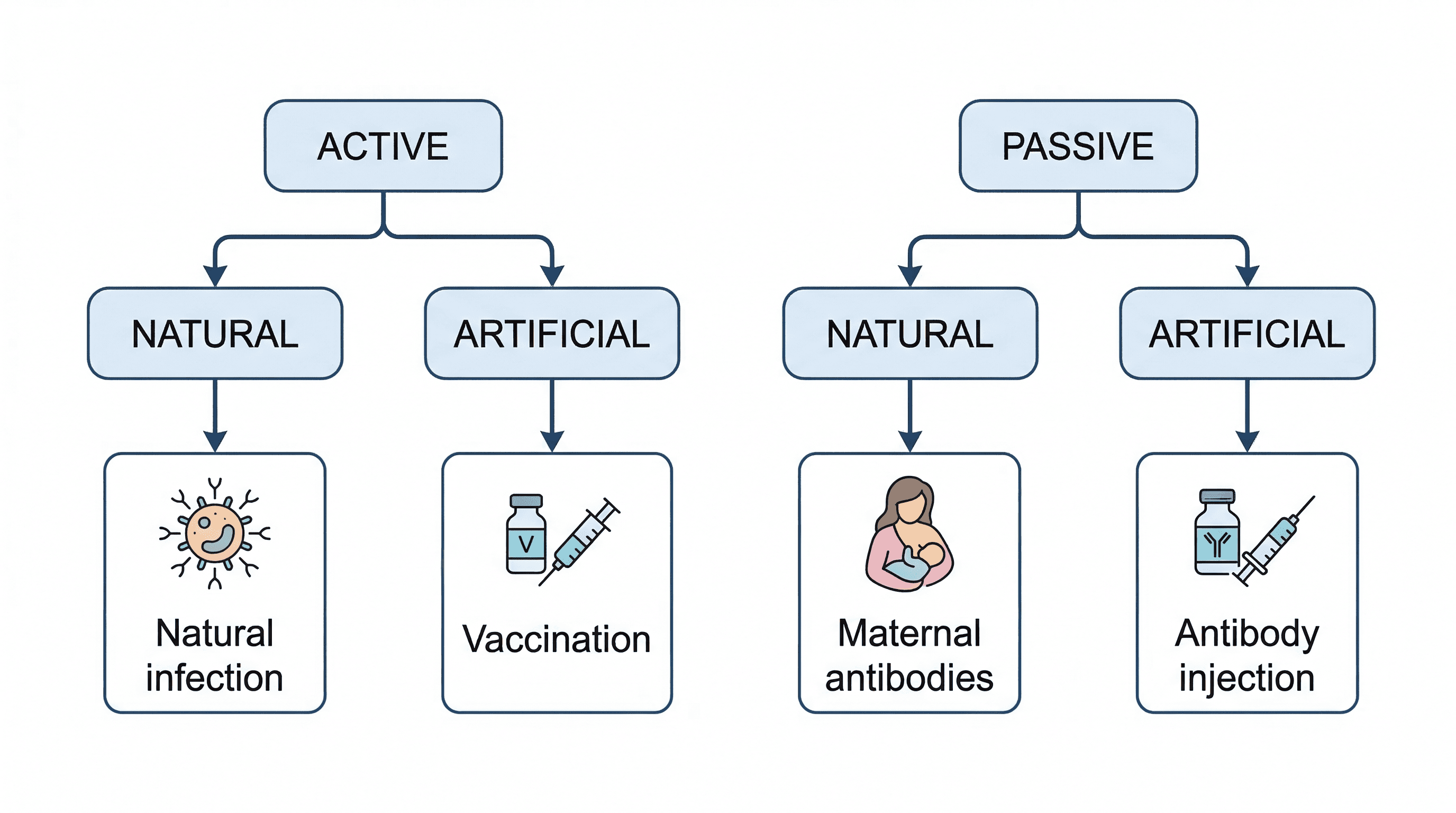
active immunity — Immunity gained when an antigen enters the body, an immune response occurs and antibodies are produced by plasma cells.
This type of immunity involves the body's own immune system actively producing antibodies and memory cells in response to an antigen. It provides long-term protection because of immunological memory, like your body learning to fight an enemy by engaging in battle.
natural active immunity — Immunity gained by being infected by a pathogen.
This occurs when a person naturally encounters a pathogen, gets infected, and their immune system mounts a primary response, leading to the production of memory cells and long-term immunity, like getting sick and then becoming immune.
vaccine — A preparation containing antigens to stimulate active immunity against one or several diseases.
Vaccines introduce antigens (e.g., weakened pathogens, dead pathogens, or pathogen fragments) into the body without causing disease. This stimulates a primary immune response, leading to the formation of memory cells and long-term immunity, acting like a 'training exercise' for your immune system.
Students often think vaccines contain antibodies, but actually they contain antigens to stimulate the body's own antibody production.
artificial active immunity — Immunity gained by putting antigens into the body, either by injection or by mouth.
This is achieved through vaccination, where antigens are deliberately introduced to stimulate the immune system. It results in the production of memory cells and long-term immunity without experiencing the full disease, like getting a 'drill' or 'simulation' of an enemy attack.
vaccination — Giving a vaccine containing antigens for a disease, either by injection or by mouth; vaccination confers artificial active immunity without the development of symptoms of the disease.
This medical procedure aims to prevent infectious diseases by stimulating the immune system to develop immunological memory. It is a key public health strategy for controlling the spread of diseases, like giving your body a 'mugshot' of a criminal.
passive immunity — The temporary immunity gained without there being an immune response.
This type of immunity involves receiving pre-formed antibodies from another source, rather than the body producing its own. It provides immediate but temporary protection because no memory cells are formed, like receiving 'borrowed' weapons.
Students often think passive immunity is long-lasting, but actually it is temporary because the body does not produce its own memory cells.
artificial passive immunity — The immunity gained by injecting antibodies.
This is used for immediate protection against rapidly acting diseases (e.g., tetanus, diphtheria) or toxins. Antibodies are collected from vaccinated donors and injected into the recipient, providing immediate but short-lived protection, like getting an immediate 'reinforcement' of trained soldiers.
natural passive immunity — The immunity gained by a fetus when maternal antibodies cross the placenta or the immunity gained by an infant from breast milk.
This provides temporary protection to newborns and infants against diseases their mother is immune to. Maternal antibodies (IgG across placenta, IgA in colostrum/breast milk) protect the infant until their own immune system matures, like a mother passing on her 'shield' to her baby.
In comparisons, clearly differentiate between active and passive immunity by mentioning the source of antibodies, presence/absence of memory cells, and duration of protection.
Vaccination programmes are crucial for controlling the spread of infectious diseases. By stimulating artificial active immunity, vaccines lead to the formation of memory cells without causing disease. This not only protects vaccinated individuals but also contributes to herd immunity, where a large proportion of the population being immune reduces pathogen transmission, protecting those who cannot be vaccinated.
herd immunity — Vaccinating a large proportion of the population; provides protection for those not immunised as transmission of a pathogen is reduced.
When a high percentage of a population is immune to a disease, it significantly reduces the likelihood of the pathogen spreading, thereby protecting vulnerable individuals who cannot be vaccinated (e.g., infants, immunocompromised). This creates a 'protective bubble' around a community.
For questions on vaccination, explain its role in creating herd immunity, which protects vulnerable individuals in a population by reducing pathogen transmission.
ring immunity — Vaccinating all those people in contact with a person infected with a specific disease to prevent transmission in the immediate area.
This strategy is used to contain outbreaks by creating a 'zone of immunity' around an infected individual. It prevents the pathogen from spreading further into the wider population, like creating a 'firebreak' around a small fire.
Monoclonal antibodies (Mabs) are highly specific antibodies produced in the laboratory. They are generated using the hybridoma method, which involves fusing antibody-producing plasma cells with immortal cancer cells. This creates hybridoma cells that can both secrete specific antibodies and divide indefinitely, allowing for large-scale production.
monoclonal antibody (Mab) — An antibody made by a single clone of hybridoma cells; all the antibody molecules made by the clone have identical variable regions so are specific to one antigen.
Mabs are highly specific antibodies produced in large quantities from a single B cell clone. Their specificity makes them valuable tools in diagnosis (e.g., detecting specific antigens) and treatment (e.g., targeting cancer cells), acting like 'precision-guided missiles'.
hybridoma — A cell formed by the fusion of a plasma cell and a cancer cell; it can both secrete antibodies and divide to form other cells like itself.
Hybridomas combine the antibody-producing ability of plasma cells with the immortality of cancer cells. This allows for the continuous production of large quantities of specific monoclonal antibodies in culture, like a 'super cell' that is an endless factory for specific antibodies.
For monoclonal antibody production, remember the key fusion step: a specific B-lymphocyte (plasma cell) is fused with a myeloma cell to create an immortal, antibody-producing hybridoma cell.
Monoclonal antibodies have diverse medical applications. In diagnosis, their high specificity allows them to detect specific antigens, for example, in pregnancy tests or for identifying disease markers. In treatment, Mabs can be used to target specific cells, such as cancer cells, by binding to unique antigens on their surface, or to deliver drugs directly to diseased tissues, minimising side effects.
When asked to describe the immune system, ensure you mention both cellular and molecular components and their coordinated action.
Definitions Bank
immune system
The body’s internal defence system.
antigen
A substance that is foreign to the body and stimulates an immune response (e.g. any large molecule such as a protein).
self
Refers to substances produced by the body that the immune system does not recognise as foreign, so they do not stimulate an immune response.
non-self
Refers to any substance or cell that is recognised by the immune system as being foreign and will stimulate an immune response.
antibody
A glycoprotein (immunoglobulin) made by specialised lymphocytes in response to the presence of a specific antigen; each type of antibody molecule has a shape that is complementary to its specific antigen.
+25 more definitions
View all →Command Word Guide
| Describe | For 'Describe the mode of action of macrophages and neutrophils', detail the steps of phagocytosis: engulfment, phagosome formation, fusion with lysosome, and enzymatic digestion. For 'Describe what happens during a primary immune response', outline the slower onset, lower antibody production, and generation of memory cells. |
| Explain | For 'Explain what is meant by the term antigen and state the difference between self antigens and non-self antigens', define antigen and then clearly differentiate how the immune system responds to each, linking to immune tolerance. For 'Explain how the molecular structure of antibodies is related to their functions', discuss the Y-shape, variable regions for specificity, and constant regions for effector functions. For 'Explain that vaccines contain antigens that stimulate immune responses to provide long-term immunity', detail how antigens trigger a primary response, leading to memory cell formation and subsequent faster secondary responses. |
| Outline | For 'Outline the hybridoma method for the production of monoclonal antibodies', provide a concise step-by-step account: immunisation, B cell isolation, fusion with myeloma cells, selection of hybridomas, and cloning. For 'Outline the principles of using monoclonal antibodies in the diagnosis and treatment of diseases', briefly state how their specificity allows for detection (diagnosis) or targeted action (treatment). |
| Differentiate | For 'Describe the differences between the different types of immunity: active and passive and natural and artificial', clearly state the key distinctions for each pair, focusing on the source of antibodies, whether memory cells are formed, and the duration of protection. |
Common Mistakes
Confusing antibodies (proteins made by your immune system) with antibiotics (drugs that kill bacteria).
Antibodies are specific proteins produced by lymphocytes, part of the body's natural defence. Antibiotics are medications that target and kill bacteria.
Thinking antigens are always whole pathogens.
Antigens are specific molecules (e.g., proteins, polysaccharides) that can be part of a pathogen, a toxin, or even a non-pathogenic foreign substance, and they trigger an immune response.
Stating that antibodies directly kill pathogens.
Antibodies primarily mark pathogens for destruction by other immune cells (like phagocytes), neutralise toxins, or prevent pathogens from entering cells. They do not directly kill pathogens.
+3 more
View all →This chapter explores the fundamental need for energy in living organisms, focusing on ATP as the universal energy currency. It details the four stages of aerobic respiration and anaerobic respiration pathways, relating these processes to mitochondrial structure and adaptations. The chapter also covers the comparison of energy values, respiratory quotients, and practical investigations using respirometers and redox indicators.
respiration — The enzymatic release of energy from organic compounds in living cells.
Respiration is a catabolic process that breaks down organic molecules, such as glucose, to release chemical potential energy. This energy is then used to synthesise ATP, the cell's energy currency. It can occur aerobically (with oxygen) or anaerobically (without oxygen), much like a power plant burning fuel to generate electricity.
Students often think respiration is just breathing, but actually it's a cellular process of energy release, while breathing is gas exchange.
anabolic — A chemical reaction in which small molecules are built up into larger ones.
Anabolic reactions require energy input, often supplied by ATP, to synthesise complex molecules like proteins or DNA from simpler precursors. These reactions are essential for growth, repair, and storage in living organisms, similar to building a LEGO castle from individual bricks.
When asked to 'explain' the need for energy, link anabolic reactions directly to ATP hydrolysis and the formation of larger molecules.
phosphorylation — The addition of a phosphate group to a molecule.
In glycolysis, phosphorylation of glucose by ATP raises its energy level, making it more reactive and easier to split. This initial energy investment is crucial for the subsequent energy-releasing steps, much like 'priming the pump' to get a bigger process going.
ATP synthase — The enzyme that catalyses the phosphorylation of ADP to produce ATP.
ATP synthase is a large protein complex embedded in the inner mitochondrial membrane. It acts as a channel for protons to flow down their concentration gradient, using this energy to synthesise ATP from ADP and Pi, much like a molecular turbine generating ATP as protons flow through it.
substrate-linked reaction — In the context of ATP formation, the transfer of phosphate from a substrate molecule directly to ADP to produce ATP, using energy provided directly by another chemical reaction.
This is a direct method of ATP synthesis, occurring in glycolysis and the Krebs cycle, where an enzyme transfers a phosphate group from a high-energy intermediate molecule to ADP. It does not involve the electron transport chain, acting like a direct cash transfer rather than a complex investment scheme.
Students often think all ATP is made by chemiosmosis, but actually substrate-linked phosphorylation provides a small but crucial amount of ATP.
chemiosmosis — The synthesis of ATP using energy released by the movement of hydrogen ions down their concentration gradient, across a membrane in a mitochondrion or chloroplast.
In mitochondria, protons are pumped into the intermembrane space, creating a gradient. Their subsequent flow back into the matrix through ATP synthase drives ATP production. This is the primary method of ATP synthesis in aerobic respiration, similar to a dam where water flowing through turbines generates electricity.
Be precise when describing ATP synthesis; specify 'substrate-linked phosphorylation' for direct transfers and 'chemiosmosis' for the proton gradient mechanism.
Living organisms require a constant supply of energy to carry out essential life processes. This energy is primarily used for anabolic reactions, where small molecules are built up into larger, more complex ones, crucial for growth, repair, and storage. Adenosine triphosphate (ATP) serves as the universal energy currency, providing readily available energy for these cellular activities. ATP is synthesised through processes like substrate-linked reactions and chemiosmosis, ensuring a continuous energy supply.
NAD (nicotinamide adenine dinucleotide) — A hydrogen carrier used in respiration.
NAD accepts hydrogen atoms (protons and electrons) during glycolysis, the link reaction, and the Krebs cycle, becoming reduced NAD. It then transports these hydrogens to the electron transport chain for ATP synthesis, much like a taxi picking up passengers (hydrogens) and dropping them off at their destination.
oxidation — The addition of oxygen, or the removal of hydrogen or electrons from a substance.
In respiration, organic molecules are progressively oxidised, releasing energy. The removal of hydrogen atoms, which include electrons, is a key form of oxidation in metabolic pathways, similar to 'stripping' a molecule of its energy-rich components.
Students often think oxidation only involves oxygen, but actually it's a broader concept involving loss of electrons or hydrogen.
reduction — The removal of oxygen, or the addition of hydrogen or electrons to a substance.
In respiration, carrier molecules like NAD and FAD are reduced when they accept hydrogen atoms (protons and electrons). This allows them to transport energy to the electron transport chain, much like a battery getting charged by gaining energy-rich components.
redox reaction — A chemical reaction in which one substance is reduced and another is oxidised.
Redox reactions are fundamental to respiration, as electrons and hydrogen atoms are transferred between molecules. This transfer of energy is central to ATP production, acting like a 'give and take' relationship where one molecule gives up electrons while another takes them.
Always specify 'reduced NAD' when it has accepted hydrogens and 'oxidised NAD' when it has released them, to avoid ambiguity.
Aerobic respiration is the complete enzymatic release of energy from organic compounds in the presence of oxygen. It is a highly efficient process that occurs in four main stages: glycolysis, the link reaction, the Krebs cycle, and oxidative phosphorylation. This complex series of redox reactions progressively breaks down glucose, releasing chemical potential energy to synthesise a large amount of ATP.
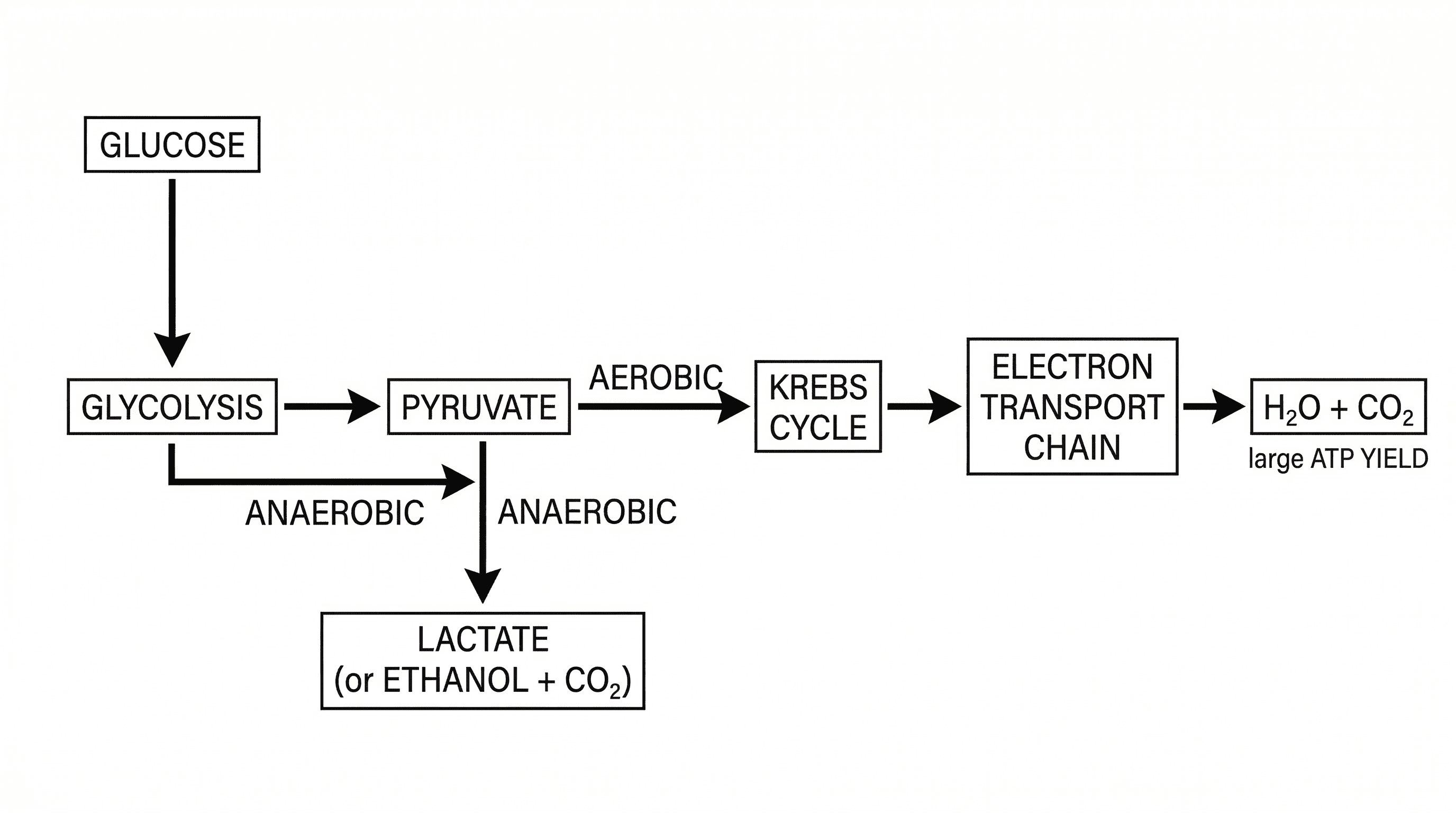
glycolysis — The splitting (lysis) of glucose; the first stage in aerobic respiration.
Glycolysis occurs in the cytoplasm and breaks down one molecule of glucose (6C) into two molecules of pyruvate (3C). This process produces a net gain of 2 ATP via substrate-linked phosphorylation and 2 reduced NAD. It can proceed in both aerobic and anaerobic conditions, much like breaking a large log into two smaller pieces for easier handling.
Students often think glycolysis requires oxygen, but actually it is an anaerobic process that can occur with or without oxygen.
link reaction — Decarboxylation and dehydrogenation of pyruvate, resulting in the formation of acetyl coenzyme A, linking glycolysis with the Krebs cycle.
This reaction occurs in the mitochondrial matrix. Pyruvate loses a carbon dioxide molecule (decarboxylation) and hydrogen atoms (dehydrogenation), which are picked up by NAD, forming acetyl CoA. It acts as the 'gateway' process preparing glycolysis products for the Krebs cycle.
decarboxylation — The removal of carbon dioxide from a substance.
Decarboxylation occurs in the link reaction and the Krebs cycle, where carbon atoms are removed from organic molecules and released as CO2. This is a key step in breaking down glucose completely, similar to 'venting' excess carbon from a molecule.
dehydrogenation — The removal of hydrogen from a substance.
Dehydrogenation reactions are crucial in glycolysis, the link reaction, and the Krebs cycle, as they provide hydrogen atoms (protons and electrons) to carrier molecules like NAD and FAD. These reduced carriers then fuel ATP synthesis in oxidative phosphorylation, much like 'harvesting' energy-rich hydrogen atoms from a fuel molecule.
coenzyme A (CoA) — A molecule that supplies acetyl groups required for the link reaction.
CoA is a complex coenzyme that combines with the 2-carbon acetyl group produced from pyruvate in the link reaction, forming acetyl coenzyme A. This molecule then delivers the acetyl group to the Krebs cycle, acting like a shuttle bus for the acetyl group.
acetyl coenzyme A — A molecule made up of CoA and a 2C acetyl group, important in the link reaction.
Formed from pyruvate and CoA in the link reaction, acetyl CoA is the entry molecule for the Krebs cycle. It delivers the 2-carbon acetyl group to oxaloacetate to form citrate, much like a delivery truck carrying a specific package to the next processing station.
Krebs cycle — A cycle of reactions in aerobic respiration in the matrix of a mitochondrion in which hydrogens pass to hydrogen carriers for subsequent ATP synthesis and some ATP is synthesised directly; also known as the citric acid cycle.
The Krebs cycle is a central metabolic pathway where acetyl CoA is completely oxidised, producing CO2, ATP (via substrate-linked phosphorylation), and a large amount of reduced NAD and FAD. These reduced carriers are essential for oxidative phosphorylation, much like a circular conveyor belt processing fuel and loading energy carriers.
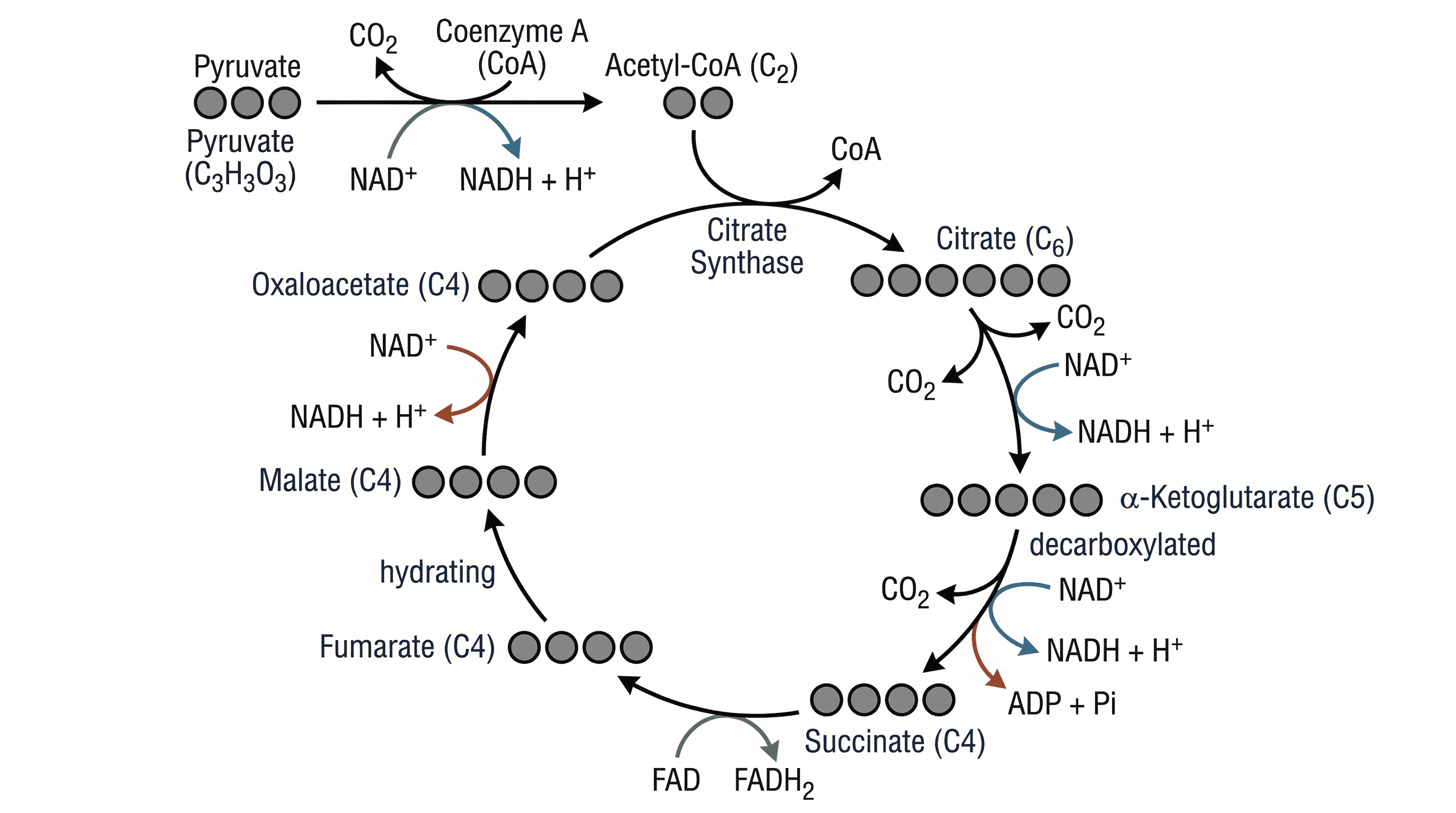
Students often think the Krebs cycle directly produces a lot of ATP, but actually its main output is reduced NAD and FAD for oxidative phosphorylation.
When describing the Krebs cycle, mention its cyclical nature, the regeneration of oxaloacetate, and the production of CO2, ATP, reduced NAD, and reduced FAD.
oxidative phosphorylation — The synthesis of ATP from ADP and Pi using energy from oxidation reactions in aerobic respiration.
This is the final and most productive stage of aerobic respiration, occurring on the inner mitochondrial membrane. It involves the electron transport chain and chemiosmosis, where oxygen acts as the final electron acceptor, much like the 'grand finale' of energy production.
electron transport chain — A chain of adjacently arranged carrier molecules in the inner mitochondrial membrane, along which electrons pass in redox reactions.
Electrons, derived from reduced NAD and FAD, move along this chain, releasing energy. This energy is used to pump protons into the intermembrane space, establishing the gradient for chemiosmosis, similar to a series of cascading waterfalls releasing energy.
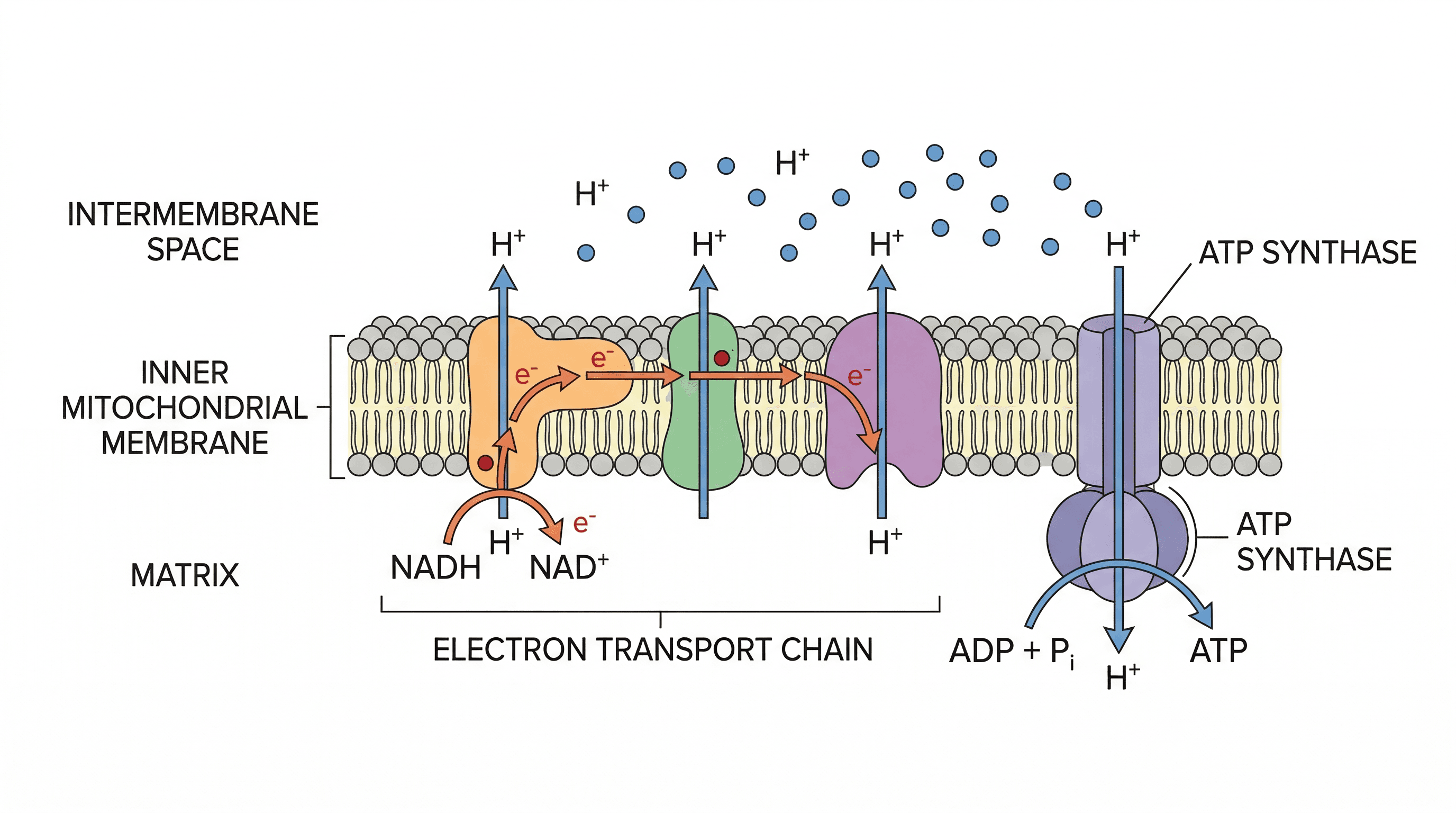
Clearly link oxidative phosphorylation to the inner mitochondrial membrane, the electron transport chain, proton gradient, ATP synthase, and the role of oxygen.
Mitochondria are often called the 'powerhouses' of the cell due to their central role in aerobic respiration. Their structure is highly adapted for this function. The inner mitochondrial membrane is extensively folded into cristae, which significantly increases the surface area for the electron transport chain and ATP synthase enzymes. The mitochondrial matrix, enclosed by the inner membrane, contains the enzymes for the link reaction and the Krebs cycle, facilitating these crucial stages of respiration.
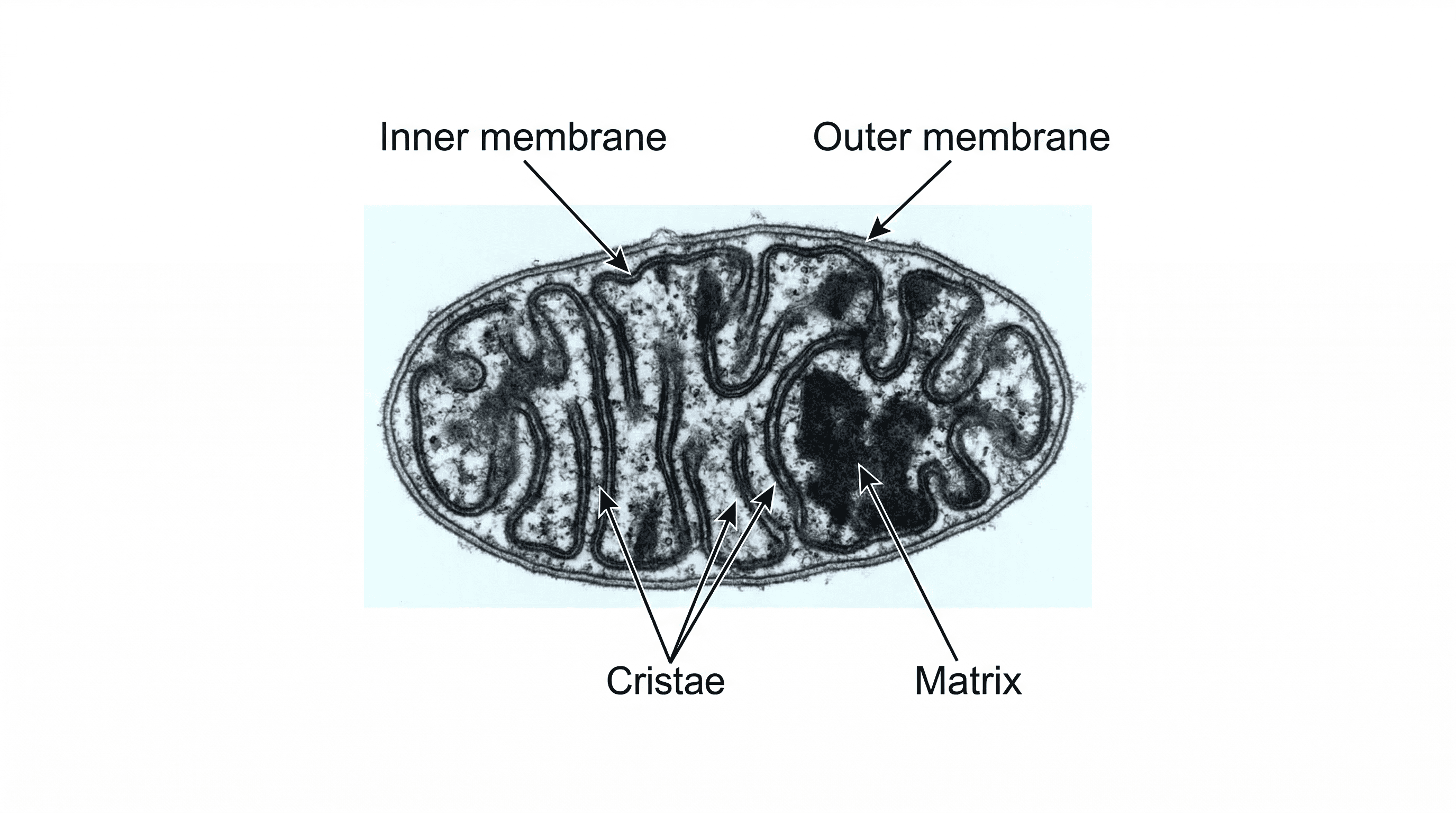
Explicitly link mitochondrial structure to function: cristae provide a large surface area for the electron transport chain and ATP synthase.
anaerobic — Without oxygen.
Anaerobic conditions mean that oxygen is not available as the final electron acceptor in respiration. This leads to alternative pathways like fermentation, which produce much less ATP than aerobic respiration, much like trying to run a car without enough air, resulting in inefficient operation.
ethanol fermentation — Anaerobic respiration in which pyruvate is converted to ethanol.
This pathway occurs in yeast and some plant tissues. Pyruvate is first decarboxylated to ethanal, then reduced by reduced NAD to ethanol, regenerating oxidised NAD for glycolysis to continue. It's like a temporary workaround when the main power grid (aerobic respiration) is down, allowing minimal energy production.
lactate fermentation — Anaerobic respiration in which pyruvate is converted to lactate.
This pathway occurs in mammalian muscles during intense exercise when oxygen supply is limited. Pyruvate is directly reduced by reduced NAD to lactate, regenerating oxidised NAD for glycolysis to continue. It acts as a short-term emergency power supply for muscles.
Students often think lactate is a waste product that cannot be used, but actually it can be converted back to pyruvate or glycogen in the liver.
aerenchyma — Plant tissue containing air spaces.
Aerenchyma tissue, found in plants like rice, provides a pathway for gases, including oxygen, to diffuse from aerial parts to submerged roots. This helps the roots respire aerobically even in flooded conditions, acting like a natural ventilation system within the plant.
While most organisms rely on aerobic respiration, some can adapt to or perform respiration in the absence of oxygen. For instance, rice plants, which often grow with their roots submerged in water, have evolved aerenchyma tissue. This specialised tissue contains air spaces that allow oxygen to diffuse from the aerial parts of the plant down to the submerged roots, enabling partial aerobic respiration even in flooded environments. This adaptation is crucial for their survival in such conditions.
respiratory quotient (RQ) — The ratio of the volume of carbon dioxide produced to the volume of oxygen used.
The RQ value indicates the type of respiratory substrate being used (carbohydrate ~1.0, lipid ~0.7, protein ~0.9) and can also signal anaerobic respiration (RQ > 1 or infinity). It's like a 'fuel gauge' for the cell, telling you what kind of fuel it's currently burning based on the gas exchange ratio.
Respiratory Quotient (RQ)
Used to determine the type of respiratory substrate or if anaerobic respiration is occurring.
Respiratory Quotient (RQ) from moles/molecules
Used when a balanced chemical equation for respiration is available.
For RQ calculations, always write the formula (RQ = CO2/O2), show your substitution, and then state the final calculated value.
respirometer — A piece of apparatus that can be used to measure the rate of oxygen uptake by respiring organisms.
A respirometer typically consists of sealed tubes containing organisms and a CO2 absorbent, connected to a manometer. Changes in manometer fluid level indicate oxygen consumption, allowing calculation of respiration rate and RQ, much like a miniature sealed environment for observing gas exchange.
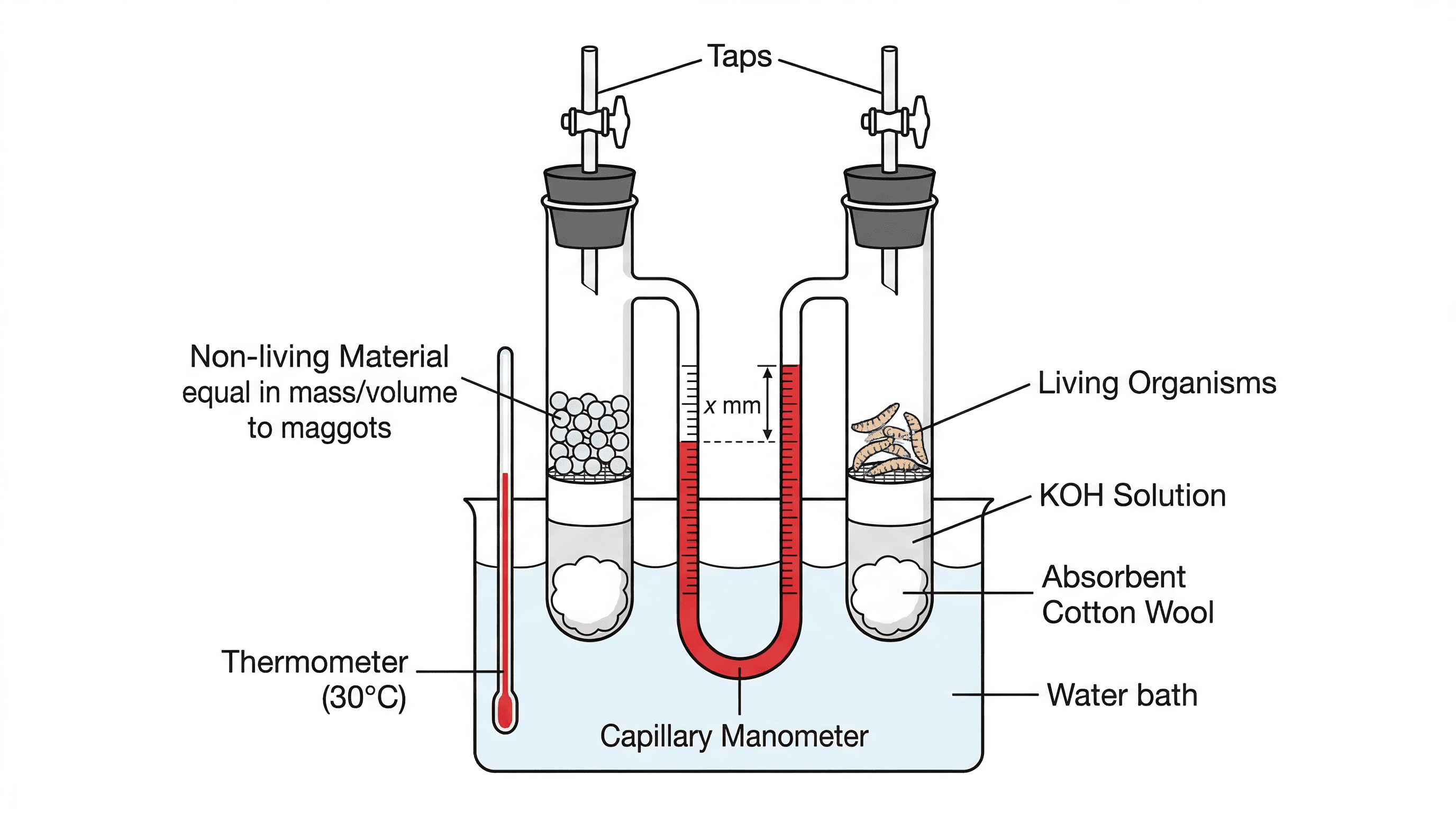
redox indicator — A substance that changes colour when it is oxidised or reduced.
Redox indicators like DCPIP or methylene blue can accept hydrogens from respiratory substrates, becoming reduced and changing colour (e.g., blue to colourless). The rate of colour change indicates the rate of respiration, acting like a chemical 'dipstick' for reaction speed.
The rate of respiration can be investigated using various methods. Respirometers are commonly used to measure the rate of oxygen uptake by respiring organisms. These setups typically include a CO2 absorbent to isolate oxygen consumption, and a manometer to quantify gas volume changes. Alternatively, redox indicators like DCPIP or methylene blue can be employed. These indicators change colour as they are reduced by hydrogen atoms released during respiration, providing a visual measure of the reaction rate.
Always state the precise location for each stage of respiration (e.g., 'mitochondrial matrix for the Krebs cycle').
Use specific terminology: 'dehydrogenation' for hydrogen removal, 'decarboxylation' for CO2 removal, and 'phosphorylation' for adding phosphate.
When describing oxidative phosphorylation, clearly state that oxygen is the 'final electron acceptor'.
Definitions Bank
anabolic
A chemical reaction in which small molecules are built up into larger ones.
respiration
The enzymatic release of energy from organic compounds in living cells.
substrate-linked reaction
In the context of ATP formation, the transfer of phosphate from a substrate molecule directly to ADP to produce ATP, using energy provided directly by another chemical reaction.
chemiosmosis
The synthesis of ATP using energy released by the movement of hydrogen ions down their concentration gradient, across a membrane in a mitochondrion or chloroplast.
glycolysis
The splitting (lysis) of glucose; the first stage in aerobic respiration.
+21 more definitions
View all →Command Word Guide
| Outline | Provide a brief summary of the main points without extensive detail. For 'Outline the need for energy', focus on anabolic reactions and ATP. |
| Explain | Give reasons or mechanisms for a phenomenon. For 'Explain how ATP is suited...', describe its properties like small, soluble, immediate energy release. For 'Explain how rice is adapted...', detail aerenchyma and its function. |
| Describe | Give a detailed account of a process or structure. For 'Describe the stages in aerobic respiration', list each stage, its location, key inputs/outputs, and the roles of specific molecules. |
| Compare | Identify similarities and differences between two or more things. For 'Compare energy values of different respiratory substrates', state typical RQ values and their implications. |
+1 more
View all →Common Mistakes
Confusing cellular respiration (energy release in cells) with breathing (gas exchange).
Respiration is a cellular process of energy release from organic compounds, while breathing is the physical process of gas exchange (taking in oxygen, releasing carbon dioxide).
Thinking glycolysis requires oxygen.
Glycolysis is an anaerobic process that occurs in the cytoplasm and does not require oxygen. It is the first stage of both aerobic and anaerobic respiration.
Believing all ATP is made by chemiosmosis.
While chemiosmosis produces the majority of ATP in aerobic respiration, a small but crucial amount of ATP is also made directly by substrate-linked phosphorylation in glycolysis and the Krebs cycle.
+3 more
View all →Photosynthesis is a vital energy transfer process where light energy is converted into chemical potential energy stored in carbohydrates. This chapter explores the intricate structure of chloroplasts, the roles of various photosynthetic pigments, and the two main stages: the light-dependent reactions and the light-independent Calvin cycle. Finally, it examines the environmental factors that limit the rate of photosynthesis and methods for its investigation.
photosynthetic pigments — Coloured substances that absorb light of particular wavelengths, supplying energy to drive the reactions in the light-dependent stage of photosynthesis.
These pigments, including chlorophyll a, chlorophyll b, carotene, and xanthophyll, are embedded in the thylakoid membranes. They capture light energy and funnel it to reaction centres, initiating the electron transport chain. Photosynthetic pigments are like an array of different-coloured 'antennae' that can each pick up specific 'radio frequencies' (wavelengths of light) to gather as much energy as possible.
chlorophyll — A green pigment that absorbs energy from light, used in photosynthesis.
Chlorophyll is the primary photosynthetic pigment found in chloroplasts, responsible for capturing light energy. It exists in two main forms, chlorophyll a and chlorophyll b, which absorb slightly different wavelengths of light, primarily in the red and blue regions of the spectrum, reflecting green light. Think of chlorophyll like a solar panel on a house; it's designed to capture energy from sunlight and convert it into a usable form for the plant.
Students often think chlorophyll absorbs all light, but actually it reflects green light, which is why plants appear green.
When asked to describe the role of chlorophyll, ensure you mention its ability to absorb light energy and channel it to reaction centres, not just 'it's green'.
absorption spectrum — A graph showing the absorbance of different wavelengths of light by a photosynthetic pigment.
An absorption spectrum illustrates which wavelengths of light a particular pigment absorbs most effectively. For example, chlorophylls absorb strongly in the blue and red regions, while reflecting green light. An absorption spectrum is like a 'light preference chart' for a pigment, showing which colours of light it 'likes' to absorb the most.
action spectrum — A graph showing the effect of different wavelengths of light on a process, for example the rate of photosynthesis.
An action spectrum plots the rate of a biological process (like photosynthesis) against the wavelength of light. It typically mirrors the combined absorption spectra of the pigments involved, showing peaks where light is most effectively used. An action spectrum is like a 'performance report' for photosynthesis, showing how well it 'works' at different colours of light.
Students often confuse absorption spectra with action spectra, but actually absorption spectra show what light is absorbed, while action spectra show how effective different wavelengths are at driving photosynthesis.
Be able to compare and contrast absorption and action spectra, explaining why they are similar (pigments absorb light used for photosynthesis) but also why they might not be identical (e.g., energy transfer efficiency, accessory pigments).
chromatography — A technique that can separate substances in a mixture according to their solubility in a solvent.
In paper chromatography for pigments, a solvent moves up the paper by capillary action, carrying dissolved pigments with it. Pigments separate based on their differential solubility in the solvent and their adsorption to the paper, resulting in distinct spots. Chromatography is like a 'race' where different pigments run at different speeds depending on how well they dissolve in the solvent and how much they stick to the paper, causing them to separate.
Rf value — A number that indicates how far a substance travels during chromatography, calculated by dividing the distance travelled by the substance by the distance travelled by the solvent; Rf values can be used to identify the substance.
The Rf value is a ratio, always between 0 and 1. A higher Rf value indicates greater solubility in the solvent and less adsorption to the stationary phase. It is a characteristic value for a given substance under specific chromatographic conditions. The Rf value is like a 'speed score' for each pigment in the chromatography race. A higher score means it travelled further relative to the solvent.
Rf value calculation
Used in chromatography to identify substances; values are always between 0 and 1.
Remember the formula: Rf = distance travelled by pigment spot / distance travelled by solvent. Know the relative Rf values for common chloroplast pigments (carotenoids highest, then chlorophyll a, then chlorophyll b).
Students often forget to measure the solvent front, but actually it's crucial for calculating the Rf value accurately.
Photosynthesis is an essential energy transfer process that converts light energy into chemical potential energy, stored within carbohydrate molecules. The overall equation for this process is 6CO₂ + 6H₂O → C₆H₁₂O₆ + 6O₂. This equation represents the net input of carbon dioxide and water, and the output of glucose and oxygen, driven by light energy in the presence of chlorophyll.
Overall equation for photosynthesis
Represents the overall input and output of photosynthesis, not the detailed steps.
stroma — The background material in a chloroplast in which the light-independent stage of photosynthesis takes place.
The stroma is the fluid-filled space within the inner membrane of a chloroplast, analogous to the cytoplasm of a cell. It contains enzymes, ribosomes, DNA, and starch grains, providing the necessary environment for the Calvin cycle reactions. The stroma is like the 'factory floor' of the chloroplast, where all the machinery (enzymes) for building carbohydrates from carbon dioxide is located.
lamellae — Membranes found within a chloroplast.
Lamellae are internal membranes within the chloroplast that connect the grana. They are part of the thylakoid membrane system and contain photosynthetic pigments and electron transport chain components. If grana are stacks of pancakes, lamellae are the single pancakes connecting different stacks, allowing communication and transport between them.
thylakoid membranes — The membranes inside a chloroplast that enclose fluid-filled sacs; the light-dependent stage of photosynthesis takes place in these membranes.
These membranes are highly folded and form flattened sacs called thylakoids, which are often stacked into grana. They embed photosynthetic pigments, electron carriers, and ATP synthase, providing the site for light absorption, electron transport, and ATP synthesis. These membranes are like the 'solar panels' and 'power lines' of the chloroplast, where light energy is captured and converted into chemical energy (ATP and reduced NADP).
thylakoid spaces — Fluid-filled sacs enclosed by the thylakoid membranes.
The thylakoid spaces (or lumen) are the internal compartments within the thylakoids. Protons are pumped into these spaces during the light-dependent stage, creating a proton gradient that drives ATP synthesis. These spaces are like a 'reservoir' for protons. As protons build up inside, they create a pressure that drives the 'water wheel' (ATP synthase) to make energy.
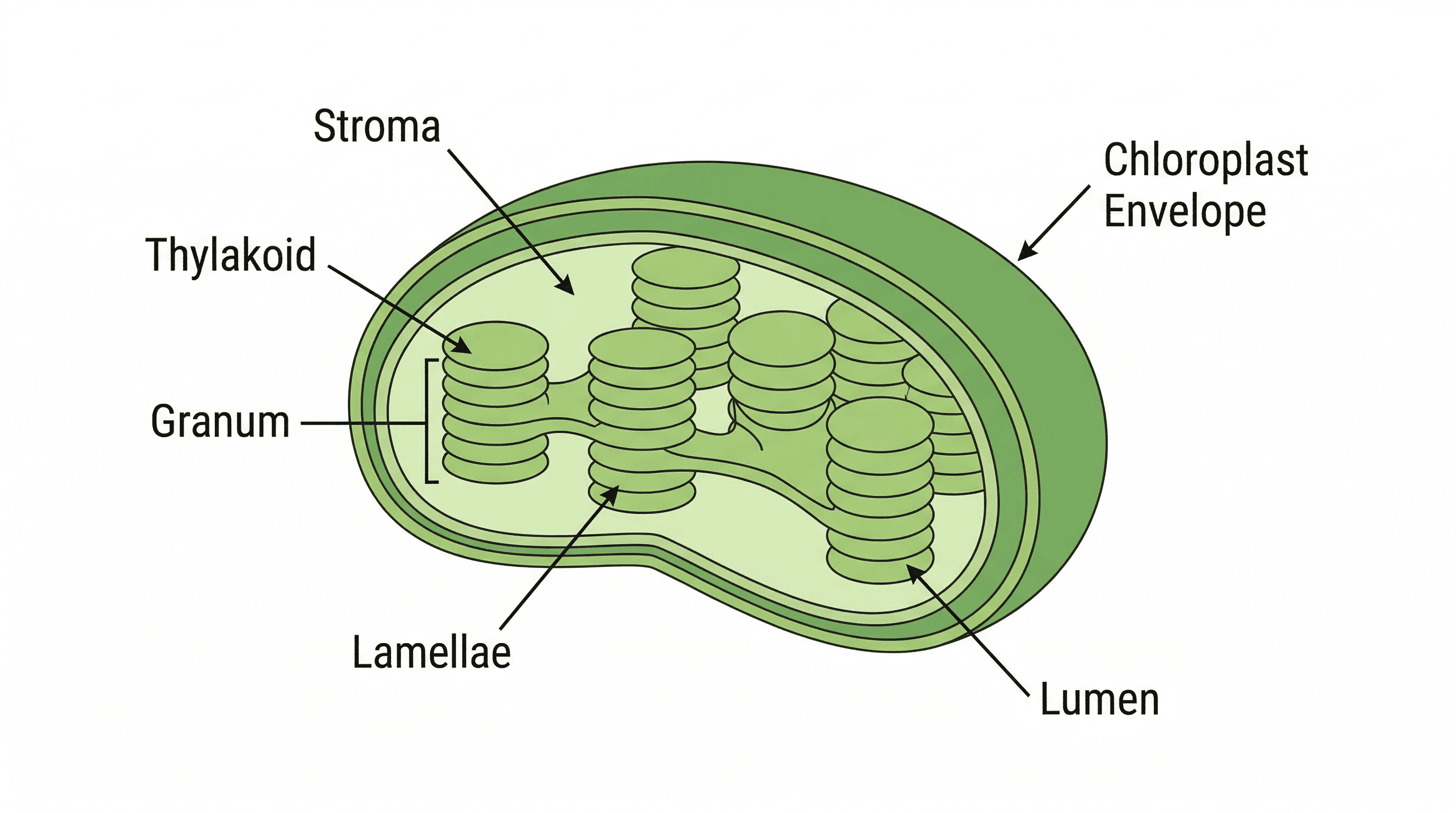
Clearly state that the light-independent stage (Calvin cycle) occurs in the stroma and mention the presence of enzymes, ribosomes, and DNA. Highlight the large surface area provided by thylakoid membranes for efficient light absorption and electron transport, linking structure to function.
light-dependent stage — The first series of reactions that take place in photosynthesis; it requires energy absorbed from light.
This stage occurs in the thylakoid membranes of chloroplasts. It involves the absorption of light energy by pigments, leading to the splitting of water (photolysis), the production of ATP (photophosphorylation), and the reduction of NADP to reduced NADP. Oxygen is released as a byproduct. The light-dependent stage is the initial power generation and raw material processing unit, creating the energy (ATP) and reducing power (reduced NADP) needed for the next steps.
Be precise about the products of the light-dependent stage: ATP, reduced NADP, and oxygen. State where it occurs (thylakoid membranes) and its energy requirement (light).
Students often think this stage directly produces glucose, but actually it produces ATP and reduced NADP, which are then used in the light-independent stage.
photosystem — A cluster of light-harvesting pigments surrounding a reaction centre.
Photosystems are functional units located in the thylakoid membranes. Each photosystem contains many pigment molecules (chlorophylls and accessory pigments) that absorb light energy and transfer it to a central reaction centre, which contains a pair of chlorophyll a molecules. A photosystem is like a 'satellite dish' with many small antennae (pigments) collecting signals (light energy) and focusing them onto a central receiver (reaction centre).
reaction centre — The part of a photosystem towards which energy from light is funnelled; it contains a pair of chlorophyll a molecules, which absorb the energy and emit electrons.
Located at the core of a photosystem, the reaction centre receives light energy from surrounding accessory pigments. This energy excites electrons in its chlorophyll a molecules, causing them to be emitted and enter the electron transport chain. The reaction centre is the 'engine' of the photosystem. All the light energy collected by the surrounding pigments is directed here to power the initial step of electron emission.
photoactivation — The emission of an electron from a molecule as a result of the absorption of energy from light.
When a chlorophyll a molecule in a reaction centre absorbs light energy, its electrons become excited to a higher energy level. If this energy is sufficiently high, the electron is emitted from the molecule and captured by an electron acceptor. Photoactivation is like a 'kick' from light energy that makes an electron jump out of its atom, ready to start a chain reaction.
photophosphorylation — Producing ATP using energy that originated as light.
This process occurs in the thylakoid membranes during the light-dependent stage. Light energy excites electrons, which then pass along an electron transport chain, releasing energy to pump protons and create a gradient. This proton gradient drives ATP synthase to produce ATP from ADP and Pi. It's like a hydroelectric dam, but instead of water, it uses the flow of electrons (energised by light) to generate a 'current' (proton gradient) that powers a turbine (ATP synthase) to make energy (ATP).
Students often confuse photophosphorylation with oxidative phosphorylation, but actually photophosphorylation uses light energy, while oxidative phosphorylation uses energy from chemical reactions (respiration).
cyclic photophosphorylation — The production of ATP using energy from light, involving only photosystem I.
In cyclic photophosphorylation, excited electrons from photosystem I are passed along an electron transport chain and then return to photosystem I. This electron flow generates a proton gradient, leading to ATP synthesis, but does not produce reduced NADP or oxygen. This is like a closed-loop power generator. Electrons are energised by light, generate ATP as they cycle through carriers, and then return to their starting point to be re-energised.
State that only Photosystem I is involved in cyclic photophosphorylation, only ATP is produced, and no water is split or oxygen released.
Students often think all photophosphorylation produces reduced NADP, but actually cyclic photophosphorylation only produces ATP.
non-cyclic photophosphorylation — The production of ATP using energy from light, involving both photosystem I and photosystem II; this process also produces reduced NADP.
This process, also known as the Z-scheme, involves both photosystem I and photosystem II. Electrons flow from photosystem II to photosystem I, generating ATP, and then from photosystem I to NADP, reducing it to reduced NADP. Water is split to replace electrons in photosystem II, releasing oxygen. This is like an open-ended power generator. Electrons move in one direction, generating ATP along the way, and then are used to create reduced NADP, requiring a constant supply of new electrons from water.
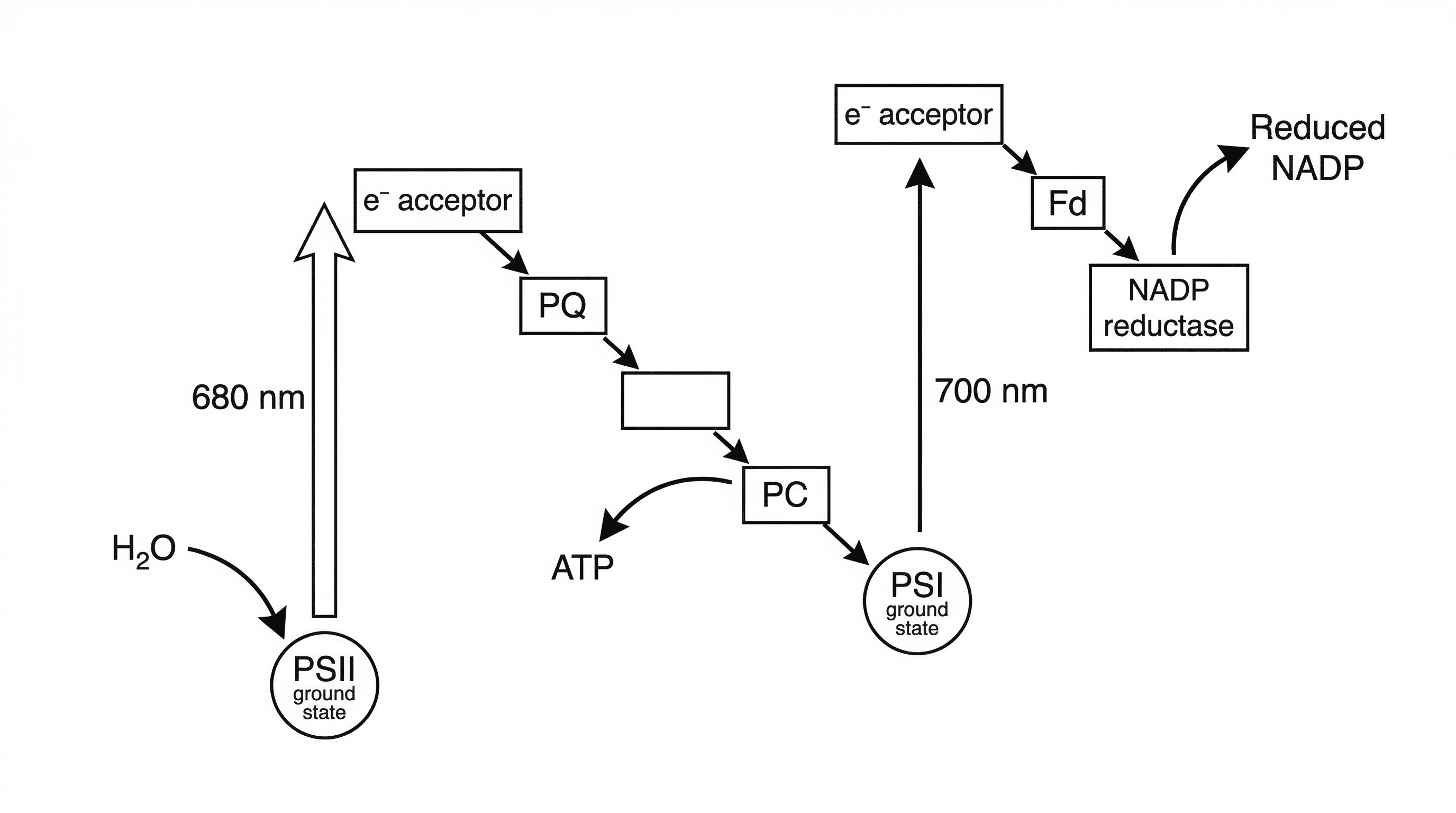
Emphasise that both ATP and reduced NADP are products of non-cyclic photophosphorylation, both photosystems are involved, and photolysis of water occurs, releasing oxygen.
photolysis — Splitting a water molecule, using energy from light.
Photolysis occurs in photosystem II during the light-dependent stage. Water molecules are split into hydrogen ions (protons), electrons, and oxygen. The electrons replace those lost from chlorophyll a, the protons contribute to the proton gradient for ATP synthesis, and oxygen is released. Think of photolysis as a water-splitting machine powered by sunlight, breaking water into its components to provide essential parts for the photosynthetic process.
Photolysis of water
Occurs in photosystem II during the light-dependent stage.
Students often think oxygen comes from carbon dioxide, but actually it comes from the splitting of water during photolysis.
Clearly state the products of water splitting (2H+, 2e-, ½O2) and its role in replacing electrons in photosystem II.
oxygen-evolving complex — An enzyme found in photosystem II that catalyses the breakdown of water, using energy from light.
This enzyme, also known as the water-splitting complex, is associated with photosystem II. It uses light energy to split water molecules (photolysis) into protons, electrons, and oxygen, providing electrons to replace those lost by chlorophyll a in photosystem II. This complex is like a 'water cracker' that breaks water molecules apart to supply the necessary electrons and protons for photosynthesis.
NADP — A coenzyme that transfers hydrogen from one substance to another, in the reactions of photosynthesis.
NADP acts as an electron and proton carrier in photosynthesis. In the light-dependent stage, it accepts electrons and hydrogen ions (protons) to become reduced NADP, which then carries this reducing power to the light-independent stage to reduce carbon dioxide. NADP is like a rechargeable battery or a shuttle bus for hydrogen and electrons. It picks them up in the light-dependent stage and delivers them where needed in the light-independent stage.
Reduction of NADP
Occurs at the end of the electron transport chain in non-cyclic photophosphorylation.
Students often confuse NADP with NAD (used in respiration), but actually NADP is specific to photosynthesis, while NAD is specific to respiration.
Always refer to it as 'reduced NADP' when it has accepted hydrogen and electrons, and 'oxidised NADP' when it has released them.
light-independent stage — The final series of reactions that take place in photosynthesis; it does not require light but does need substances that are produced in the light-dependent stage.
Also known as the Calvin cycle, this stage occurs in the stroma of the chloroplast. It uses the ATP and reduced NADP from the light-dependent stage to fix carbon dioxide and reduce it to carbohydrates like triose phosphate, which can then be converted into glucose and other organic molecules. The light-independent stage is the main manufacturing unit, taking the energy and raw materials from the first stage to build the final product (carbohydrates).
Students often think 'light-independent' means it can happen in complete darkness indefinitely, but actually it relies on the products of the light-dependent stage, which will run out in prolonged darkness.
Emphasise that while the light-independent stage doesn't directly use light, it is dependent on the products (ATP and reduced NADP) of the light-dependent stage. Mention its location (stroma) and key inputs (CO2, ATP, reduced NADP).
Calvin cycle — A cycle of reactions in the light-independent stage of photosynthesis in which carbon dioxide is reduced to form carbohydrate.
The Calvin cycle takes place in the stroma of chloroplasts. It involves three main phases: carbon fixation (CO2 combines with RuBP), reduction (GP is reduced to TP using ATP and reduced NADP), and regeneration (RuBP is regenerated from TP using ATP). The Calvin cycle is like a biochemical 'recycling plant' that takes in carbon dioxide, processes it using energy and reducing power, produces a small amount of carbohydrate, and then regenerates its starting material to keep the process going.
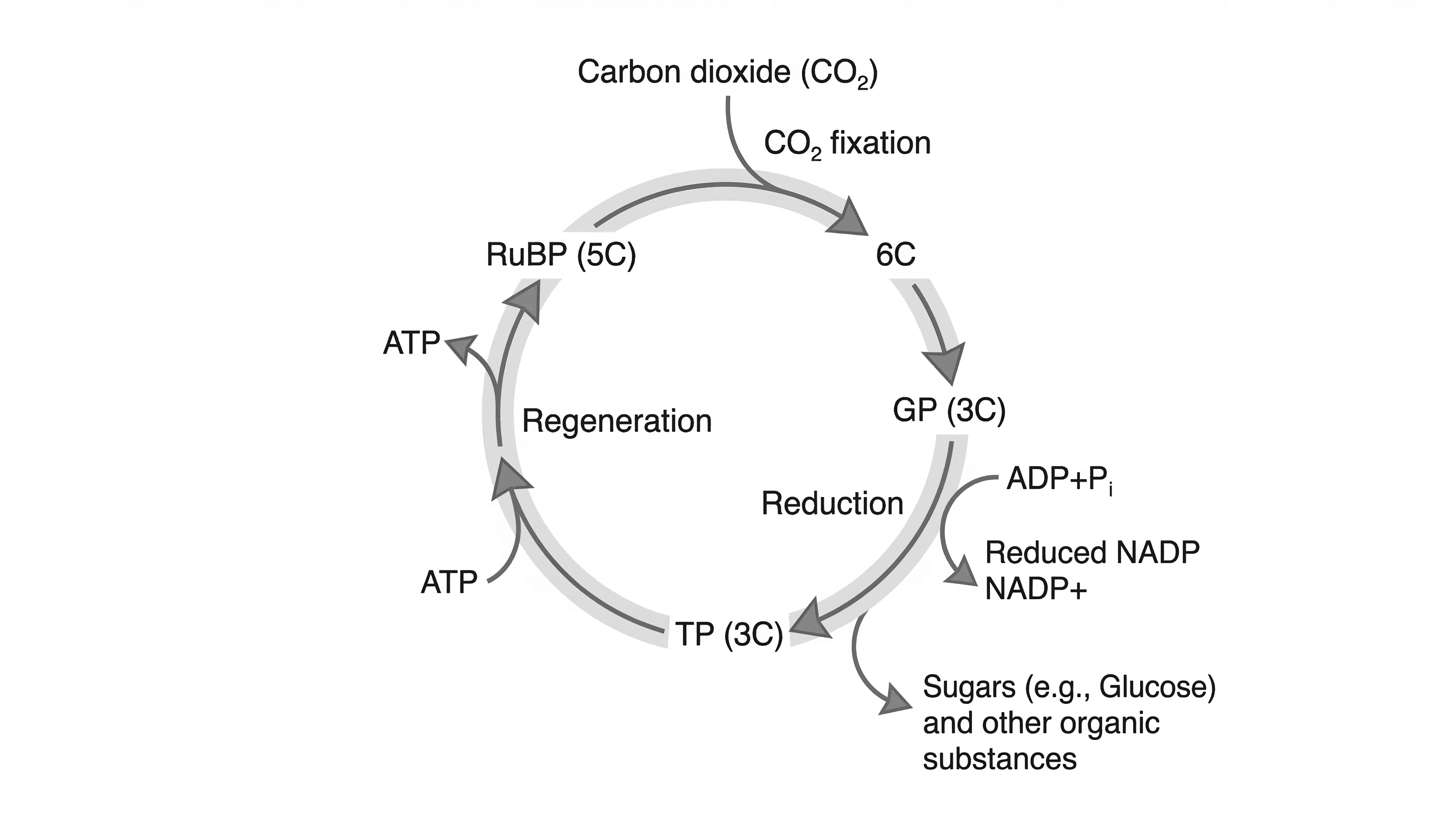
ribulose bisphosphate (RuBP) — A five-carbon phosphorylated sugar which is the first compound to combine with carbon dioxide during the light-independent stage of photosynthesis.
RuBP is a key molecule in the Calvin cycle. It acts as the carbon dioxide acceptor, combining with CO2 in a reaction catalysed by rubisco to form an unstable six-carbon intermediate, which then splits into two molecules of GP. RuBP is like the 'empty seat' waiting for carbon dioxide to join the Calvin cycle. Once CO2 sits down, the cycle can begin.
rubisco — The enzyme that catalyses the combination of RuBP with carbon dioxide.
Rubisco (ribulose bisphosphate carboxylase-oxygenase) is arguably the most abundant enzyme on Earth. It catalyses the crucial carbon fixation step in the Calvin cycle, where CO2 is incorporated into an organic molecule (RuBP). Rubisco is the 'gatekeeper' enzyme that allows carbon dioxide to enter the photosynthetic pathway and become part of organic molecules.
glycerate-3-phosphate (GP) — A three-carbon compound which is formed when RuBP combines with carbon dioxide.
After RuBP combines with CO2 and the unstable 6C intermediate splits, two molecules of GP are formed. GP is then reduced to triose phosphate (TP) using ATP and reduced NADP from the light-dependent stage. GP is like the 'intermediate product' in the carbohydrate factory. It's not the final product, but it's the next step after the raw material (CO2) has been incorporated.
triose phosphate (TP) — A three-carbon phosphorylated sugar, the first carbohydrate to be formed during the light-independent stage of photosynthesis.
TP is formed from the reduction of GP using ATP and reduced NADP. It is a crucial molecule because some TP is used to regenerate RuBP, while the rest is used to synthesise other organic molecules like glucose, starch, sucrose, lipids, and amino acids. TP is the 'versatile building block' produced by the Calvin cycle. It can either be recycled to keep the cycle going or used to build all the other essential organic compounds for the plant.
Students often think the Calvin cycle directly produces glucose, but actually it produces triose phosphate (TP), which is then used to synthesise glucose and other organic molecules.
Be able to describe the key compounds (RuBP, GP, TP) and the role of rubisco, ATP, and reduced NADP in each step of the Calvin cycle.
limiting factor — The requirement for a process to take place that is in the shortest supply; an increase in this factor will allow the process to take place more rapidly.
In photosynthesis, common limiting factors include light intensity, carbon dioxide concentration, and temperature. The rate of photosynthesis is determined by the factor that is furthest from its optimum level, even if other factors are abundant. Think of a car assembly line. If you have plenty of parts and workers but only one wrench, the wrench is the limiting factor for how fast you can build cars. Increasing the number of wrenches will speed up production until something else becomes limiting.
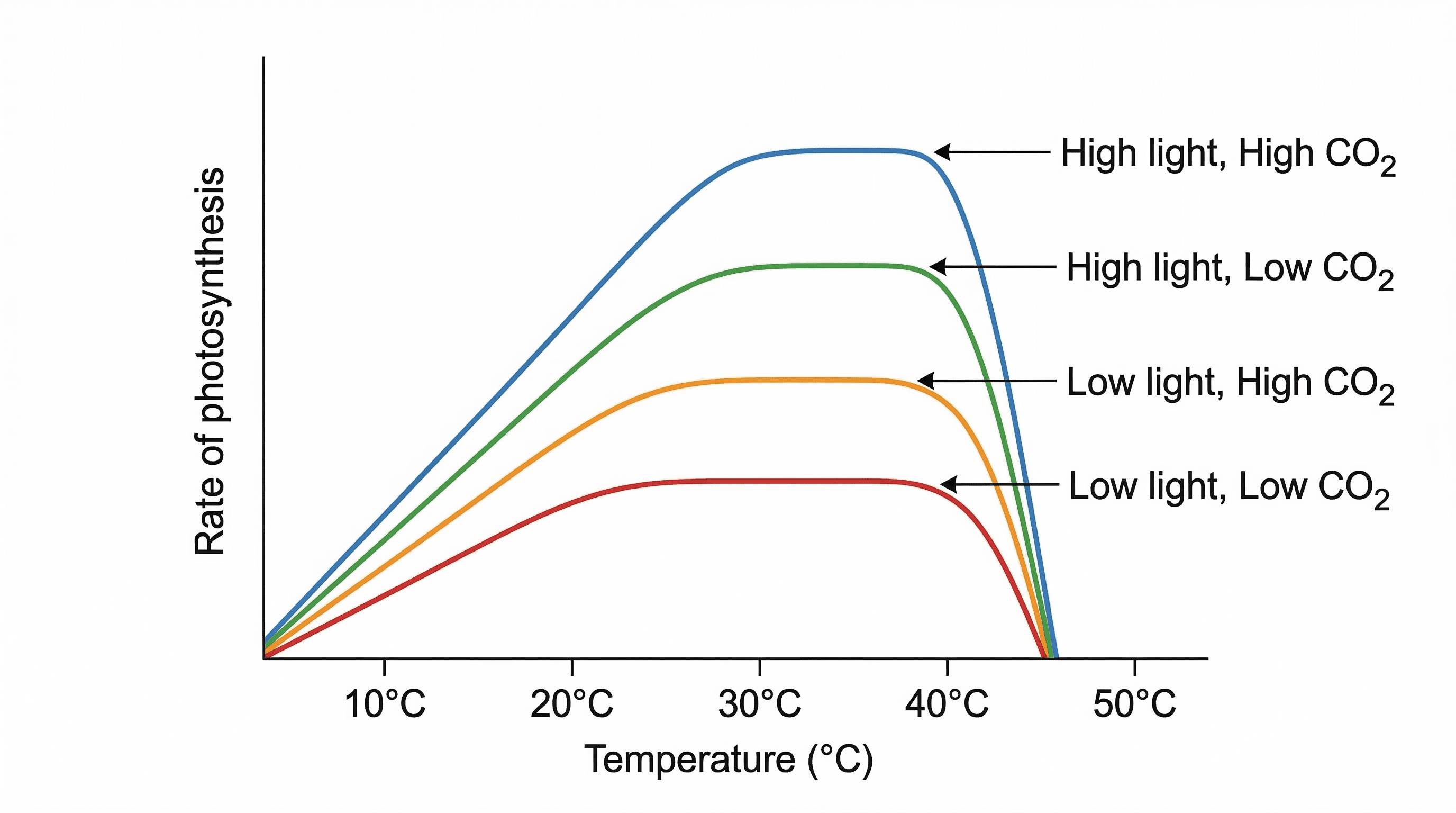
Students often think that increasing any factor will always increase the rate, but actually only increasing the *limiting* factor will increase the rate.
When interpreting graphs, identify the region where the rate is increasing in response to a change in a factor, indicating that factor is limiting. Where the rate plateaus, another factor has become limiting.
The rate of photosynthesis can be investigated by measuring the uptake of carbon dioxide or the production of oxygen. For aquatic plants, the rate of oxygen bubble production can be counted or the change in pH (due to CO2 uptake) can be monitored. Using a redox indicator with a chloroplast suspension allows for the investigation of the effect of light intensity and wavelengths on the light-dependent stage, as the indicator changes colour when reduced by electrons from the electron transport chain.
When asked to investigate photosynthesis (e.g., using an aquatic plant), state the independent and dependent variables and describe how you would control all other variables to ensure a valid result.
Definitions Bank
chlorophyll
A green pigment that absorbs energy from light, used in photosynthesis.
light-dependent stage
The first series of reactions that take place in photosynthesis; it requires energy absorbed from light.
light-independent stage
The final series of reactions that take place in photosynthesis; it does not require light but does need substances that are produced in the light-dependent stage.
photolysis
Splitting a water molecule, using energy from light.
photophosphorylation
Producing ATP using energy that originated as light.
+22 more definitions
View all →Command Word Guide
| Describe | For chloroplast structure, describe the thylakoid membranes, grana, stroma, and their relative positions. For stages, describe the sequence of events, inputs, and outputs. For chromatography, describe the practical steps. |
| Explain | For energy transfer, explain how light energy is converted to chemical energy. For structure-function, explain how specific features of the chloroplast (e.g., large surface area of thylakoids) relate to their roles. For limiting factors, explain why a factor limits the rate and how increasing it affects the process. |
| Interpret | For absorption and action spectra, interpret the peaks and troughs in relation to pigment absorption and photosynthetic efficiency. For limiting factor graphs, interpret the different regions of the curve to identify the limiting factor. |
| Calculate | For chromatography, calculate Rf values using the provided formula and measurements. |
Common Mistakes
Thinking oxygen produced in photosynthesis comes from carbon dioxide.
Oxygen actually comes from the splitting of water (photolysis) during the light-dependent stage.
Confusing the light-dependent and light-independent stages, or thinking the latter can occur indefinitely in darkness.
The light-independent stage relies on ATP and reduced NADP from the light-dependent stage, so it will cease when these products run out in prolonged darkness.
Confusing absorption spectra (what light is absorbed) with action spectra (how effective different wavelengths are at driving photosynthesis).
Absorption spectra show the wavelengths a pigment absorbs, while action spectra show the rate of photosynthesis at different wavelengths.
+3 more
View all →This chapter explores homeostasis, the vital process of maintaining a stable internal environment in mammals, primarily through negative feedback mechanisms coordinated by the nervous and endocrine systems. It details kidney function in urine formation and osmoregulation, blood glucose control by insulin and glucagon, and plant homeostasis focusing on stomatal regulation.
homeostasis — The maintenance of a relatively constant internal environment for the cells within the body.
This ensures that cells can function efficiently by keeping physiological factors like temperature, pH, and solute concentrations within optimum ranges, preventing damage to enzymes and metabolic processes. Like a thermostat in a house, homeostasis keeps the body's internal conditions stable despite external changes, ensuring all 'rooms' (cells) are comfortable for work.
Students often think homeostasis means conditions are absolutely constant, but actually, they fluctuate slightly around a set point.
set point — The ideal value of a physiological factor that the body controls in homeostasis.
The set point is the target value around which a physiological factor is maintained. Homeostatic mechanisms work to keep the actual value fluctuating within a narrow range around this ideal. The set temperature on a thermostat is the set point for the room's temperature.
stimulus — A change in the external or internal environment that is detected by a receptor and which may cause a response.
Stimuli are the triggers that initiate homeostatic responses. They can be internal, such as a change in blood glucose, or external, such as a change in ambient temperature. A sudden loud noise is a stimulus that might cause you to jump (a response).
receptor — A cell or tissue that is sensitive to a specific stimulus and communicates with a control centre by generating nerve impulses or sending a chemical messenger.
Receptors detect changes in internal or external conditions, initiating a response to maintain homeostasis. They are the 'sensors' of the body's control systems. Like a smoke detector in a building, a receptor detects a specific change (smoke/stimulus) and sends a signal to a central alarm system (control centre).
effector — A tissue or organ that carries out an action in response to a stimulus; muscles and glands are effectors.
Effectors are the 'responders' in a homeostatic loop, performing corrective actions to restore the physiological factor to its set point. Their actions are coordinated by the nervous or endocrine system. If a thermostat detects the room is too cold, the effector (furnace) turns on to heat the room, correcting the temperature.
corrective action — A response or series of responses that return a physiological factor to the set point so maintaining a constant environment for the cells within the body.
These actions are the output of the homeostatic control system, designed to reverse the initial deviation from the set point and restore balance. If your body temperature rises, sweating and vasodilation are corrective actions to cool you down.
negative feedback — A process in which a change in some parameter (e.g. blood glucose concentration) brings about processes which return it towards normal.
This mechanism minimises the difference between the actual value of a factor and its ideal set point, ensuring stability. If a factor increases, negative feedback causes it to decrease, and vice versa. Imagine a car's cruise control: if the car speeds up (change), the system reduces engine power (response) to bring it back to the set speed (normal).
Students often think negative feedback means a bad response, but actually, it refers to a response that reverses the initial change, maintaining stability.
When describing negative feedback, clearly identify the stimulus, receptor, control centre, effector, and the corrective action that reverses the initial change.
positive feedback — A process in which a change in some parameter such as a physiological factor brings about processes that move its level further in the direction of the initial change.
Unlike negative feedback, positive feedback amplifies the initial change, moving the system further away from the set point. It is rare in homeostatic control but important in processes like childbirth. A microphone feedback loop where the sound gets progressively louder and louder, amplifying the initial noise.
hormone — A substance secreted by an endocrine gland that is carried in blood plasma to another part of the body where it has an effect.
Hormones are chemical messengers of the endocrine system, enabling long-distance cell signalling to coordinate physiological responses, often slower but longer-lasting than nervous responses. Like a message in a bottle sent across an ocean, a hormone travels through the bloodstream to deliver its specific message to distant target cells.
Homeostasis is crucial for mammals to maintain a relatively constant internal environment, which is essential for efficient cell function. By keeping physiological factors like temperature, pH, and solute concentrations within optimum ranges, it prevents damage to enzymes and metabolic processes, ensuring coordinated function of the body's systems.
excretion — The removal of toxic or waste products of metabolism from the body.
Excretion is vital for maintaining a healthy internal environment by eliminating substances that could be harmful if allowed to accumulate, such as carbon dioxide and urea. Like taking out the trash from your house, excretion removes unwanted metabolic byproducts from the body.
Students often think excretion is the same as egestion, but actually, egestion is the removal of undigested food, while excretion is the removal of metabolic wastes.
deamination — The breakdown of excess amino acids in the liver, by the removal of the amine group; ammonia and, eventually, urea are formed from the amine group.
This process allows the body to utilise the energy content of excess amino acids by converting the remaining keto acid into glucose or fat, while safely disposing of the nitrogenous waste. Like disassembling a toy to reuse its parts: the amine group is removed for disposal, and the remaining carbon skeleton is repurposed for energy or storage.
urea — A nitrogenous excretory product produced in the liver from the deamination of amino acids.
Urea is less toxic than ammonia, which is initially formed during deamination, making it a safer form for transport in the blood to the kidneys for excretion. Like converting highly toxic industrial waste into a less harmful, transportable form before disposal, the body converts ammonia to urea.
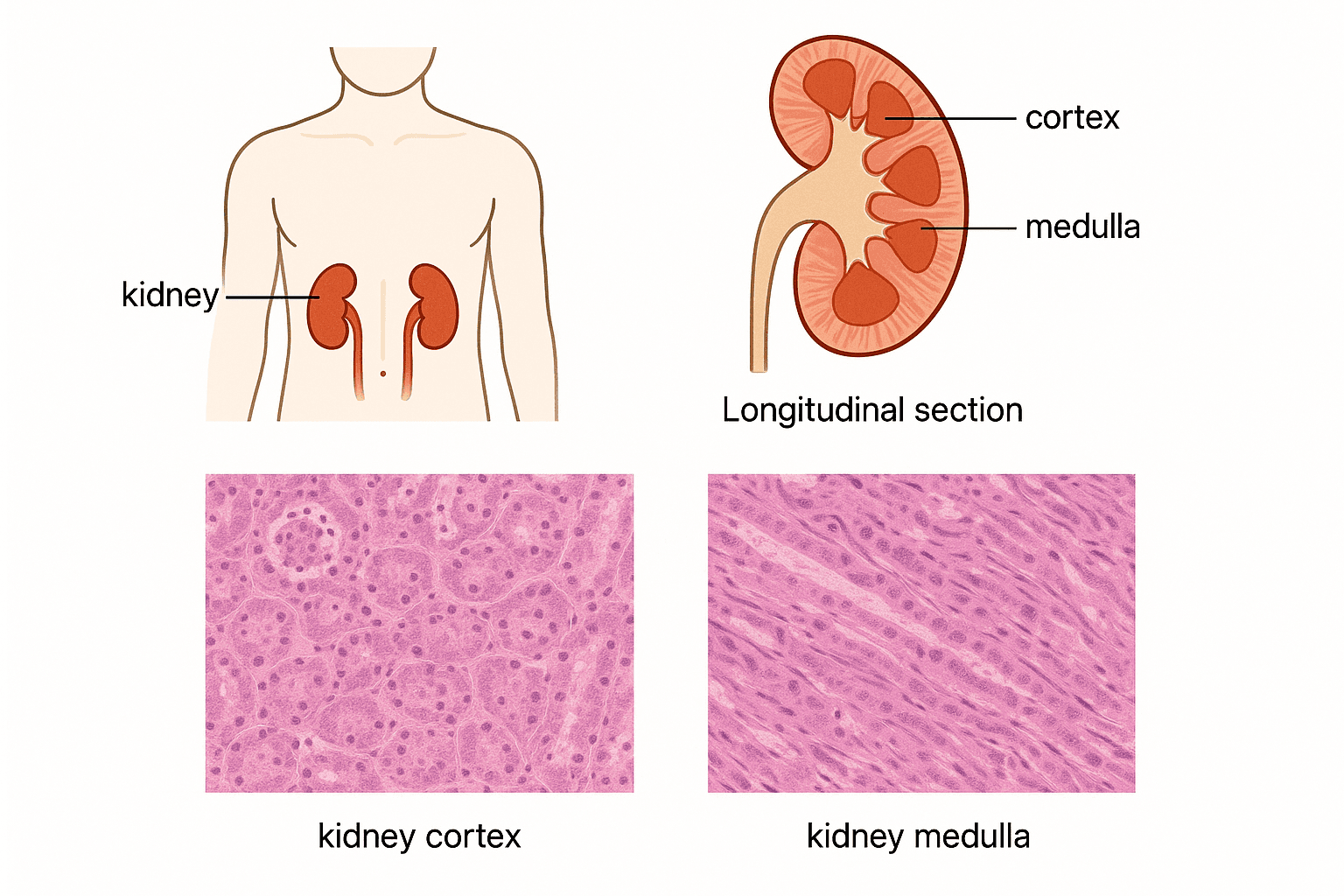
nephron — The structural and functional unit of the kidney composed of Bowman’s capsule and a tubule divided into three regions: proximal convoluted tubule, loop of Henle and distal convoluted tubule.
Each kidney contains thousands of nephrons, which are responsible for filtering blood, reabsorbing useful substances, and forming urine to regulate body fluid composition. Like a miniature water treatment plant, each nephron filters, purifies, and concentrates waste from the blood.
afferent arteriole — Arteriole leading to glomerular capillaries.
It has a wider diameter than the efferent arteriole, contributing to the high hydrostatic pressure within the glomerulus, which is essential for ultrafiltration. Like a wide pipe feeding water into a narrower hose, it creates pressure in the glomerulus.
efferent arteriole — Arteriole leading away from glomerular capillaries.
Its narrower diameter, compared to the afferent arteriole, helps maintain the high pressure in the glomerulus, facilitating ultrafiltration. Like a narrower pipe restricting water flow out of a system, it helps build up pressure upstream.
glomerulus — A group of capillaries within the ‘cup’ of a Bowman’s capsule in the cortex of the kidney.
The high pressure within these capillaries, due to the afferent arteriole being wider than the efferent, drives the process of ultrafiltration, forcing fluid and small solutes into Bowman's capsule. Like a high-pressure sprinkler head, the glomerulus forces water (blood plasma) through tiny holes (capillary pores) into the surrounding collection basin (Bowman's capsule).
Bowman’s capsule — The cup-shaped part of a nephron that surrounds a glomerulus and collects filtrate from the blood.
This is where ultrafiltration occurs, as blood plasma (minus large proteins and cells) is forced out of the glomerulus and into the capsule to form glomerular filtrate. Like a coffee filter and funnel, the Bowman's capsule collects the liquid (filtrate) that passes through the filter (glomerulus).
podocyte — One of the cells that makes up the lining of Bowman’s capsule surrounding the glomerular capillaries.
These cells have finger-like projections with gaps (slit pores) that form part of the filtration barrier, allowing filtrate to pass through while preventing the passage of larger molecules. Like a hand with fingers spread out, the podocyte's projections create spaces for fluid to pass through, but still act as a barrier.
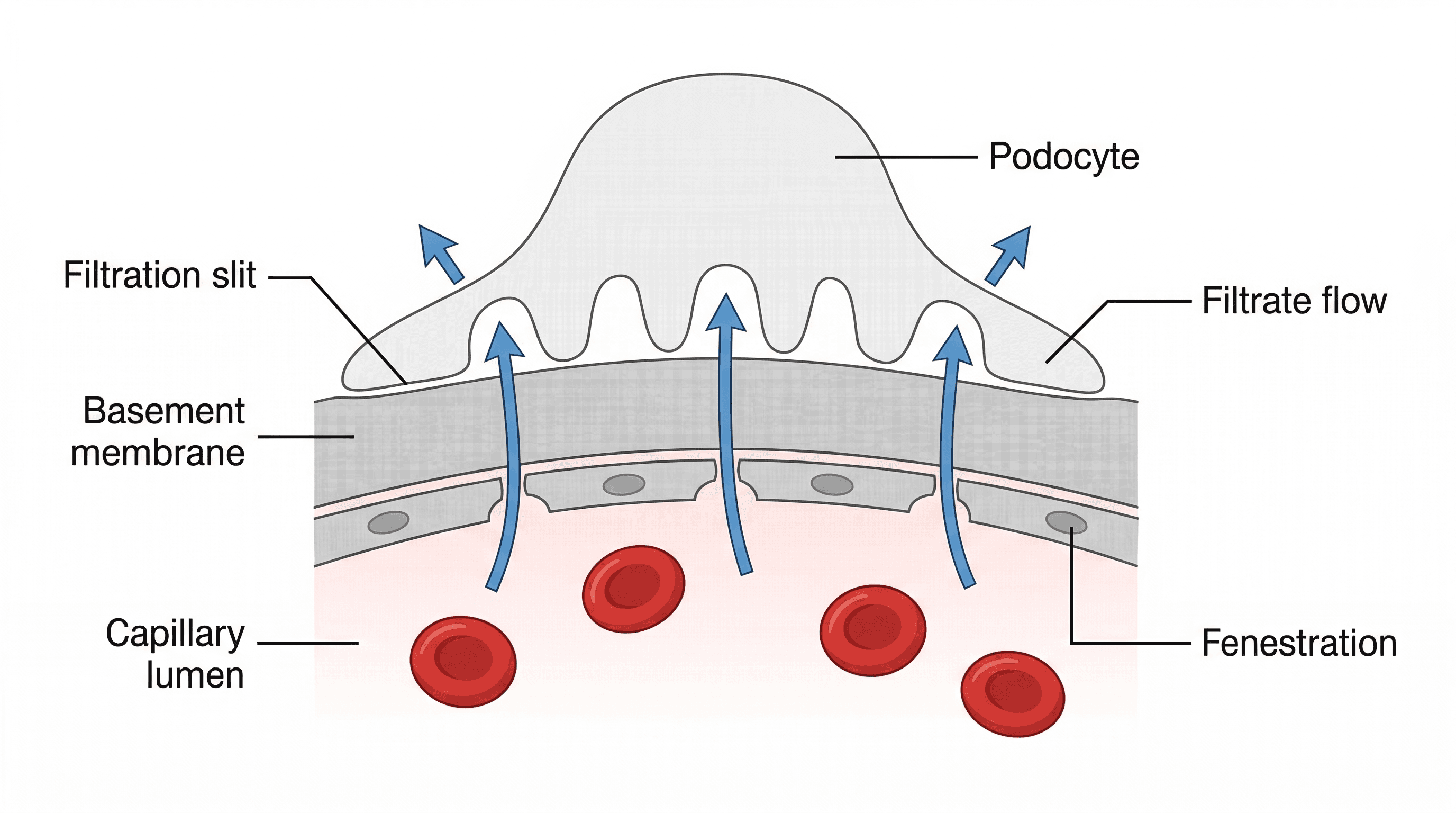
ultrafiltration — Filtration on a molecular scale separating small molecules from larger molecules, such as proteins (e.g. the filtration that occurs as blood flows through capillaries, especially those in glomeruli in the kidney).
This process occurs in the Bowman's capsule, driven by high blood pressure, forcing water and small solutes out of the blood while retaining large proteins and blood cells. Like a very fine sieve, ultrafiltration allows small particles (water, ions, glucose) to pass through but blocks larger ones (proteins, blood cells).
For ultrafiltration, always mention the high hydrostatic pressure in the glomerulus forcing small molecules from the blood into the Bowman's capsule.
selective reabsorption — Movement of certain substances from the filtrate in nephrons back into the blood.
This process ensures that useful substances like glucose, amino acids, and most water are reclaimed from the filtrate and returned to the bloodstream, preventing their loss in urine. Like a quality control check, the nephron selectively reclaims valuable items from the filtered fluid, sending them back to circulation.
proximal convoluted tubule — Part of the nephron that leads from Bowman’s capsule to the loop of Henle.
This region is responsible for the majority of selective reabsorption, taking back essential substances like glucose, amino acids, and most water from the filtrate into the blood. Like a diligent recycling plant, the proximal convoluted tubule reclaims almost all the valuable materials from the initial waste stream.
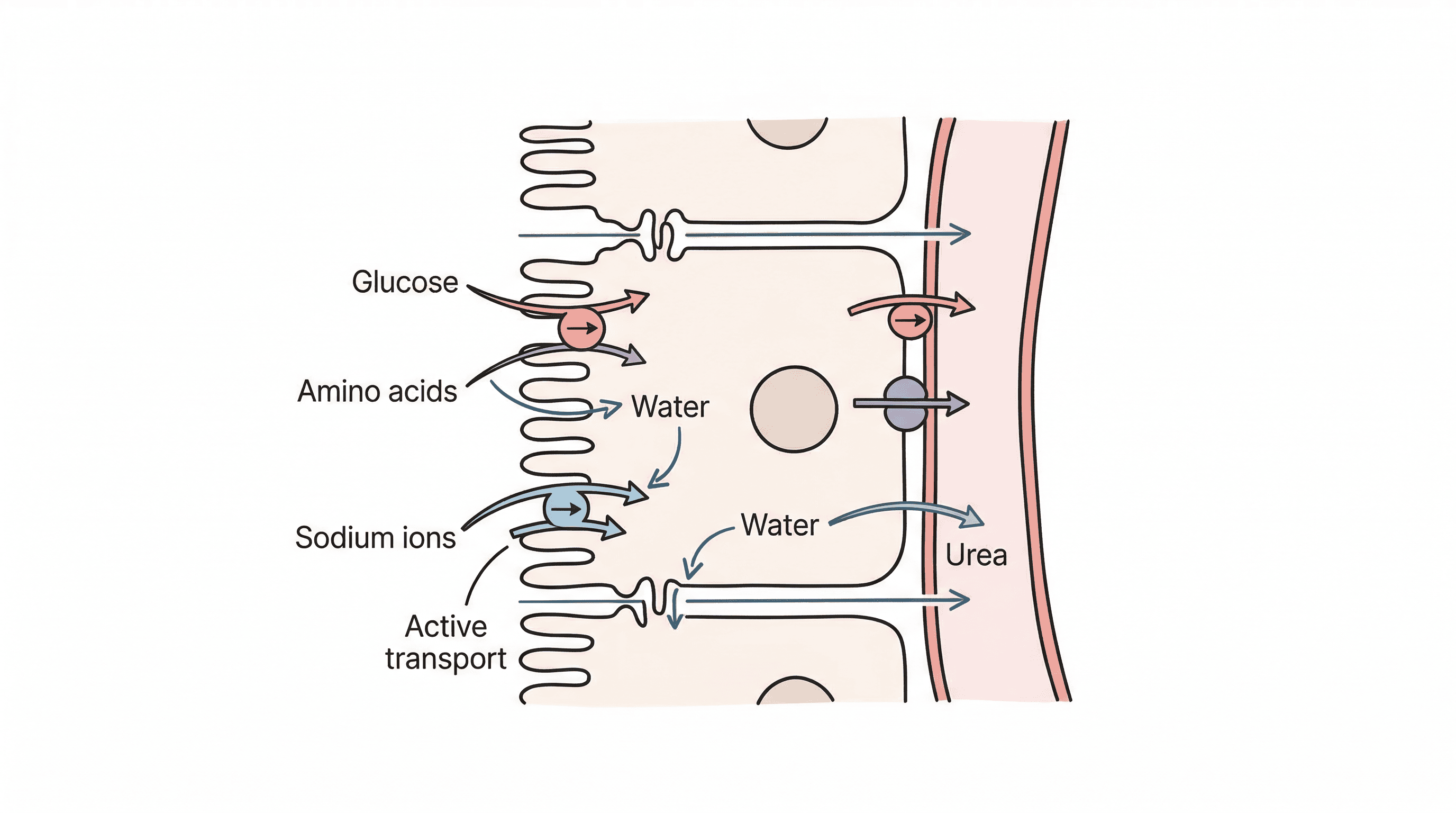
Link the structure of the proximal convoluted tubule (PCT) to its function: microvilli for large surface area and many mitochondria for active transport in selective reabsorption.
loop of Henle — The part of the nephron between the proximal and distal convoluted tubules.
Its primary function is to create a high concentration of sodium and chloride ions in the tissue fluid of the medulla, establishing a water potential gradient crucial for water reabsorption from the collecting duct. Like a counter-current heat exchanger, the loop of Henle uses opposing flows to build up a concentration gradient in the surrounding tissue.
distal convoluted tubule — Part of the nephron that leads from the loop of Henle to the collecting duct.
This region allows for fine-tuning of ion and water reabsorption, with its permeability to water being regulated by ADH, contributing to the final concentration of urine. Like the final adjustment knob on a machine, the distal convoluted tubule makes precise changes to the filtrate composition before it becomes urine.
collecting duct — Tube in the medulla of the kidney that carries urine from the distal convoluted tubules of many nephrons to the renal pelvis.
The collecting duct's permeability to water is regulated by ADH, allowing variable amounts of water to be reabsorbed by osmosis into the highly concentrated medullary tissue fluid, thus controlling urine concentration. Like a water tap that can be opened or closed, the collecting duct's permeability is controlled by ADH to adjust how much water is retained or lost.
Urine is formed in the nephrons through two main processes: ultrafiltration and selective reabsorption. Ultrafiltration occurs in the Bowman's capsule, where high hydrostatic pressure forces blood plasma (excluding large proteins and cells) into the capsule. The resulting filtrate then undergoes selective reabsorption in the proximal convoluted tubule, loop of Henle, distal convoluted tubule, and collecting duct, where useful substances are returned to the blood, and waste products are concentrated into urine.
osmoregulation — The control of the water potential of blood and tissue fluid by controlling the water content and/or the concentration of ions, particularly sodium ions.
This homeostatic process maintains the appropriate water balance in the body, preventing cells from swelling or shrinking due to osmotic imbalances, which would disrupt metabolic functions. Like a gardener carefully watering plants to keep the soil moisture just right, osmoregulation maintains the perfect water balance in the body's internal environment.
osmoreceptor — Type of receptor that detects changes in the water potential of blood.
Located in the hypothalamus, these specialised sensory neurones monitor blood water potential and initiate the release of ADH when a decrease is detected, triggering water reabsorption. Like a humidity sensor, an osmoreceptor detects changes in the 'wetness' (water potential) of the blood and signals for adjustment.
antidiuretic hormone (ADH) — Hormone secreted from the posterior pituitary gland that increases water reabsorption in the kidneys and therefore reduces water loss in urine.
ADH acts on the collecting ducts and distal convoluted tubules, increasing their permeability to water by inserting aquaporins, allowing more water to be reabsorbed into the blood when the body is dehydrated. Like a 'water-saving' signal, ADH tells the kidneys to open more water channels, allowing the body to conserve water.
Students often think ADH directly causes water to move, but actually, it increases the *permeability* of collecting ducts to water, allowing osmosis to occur down an existing water potential gradient.
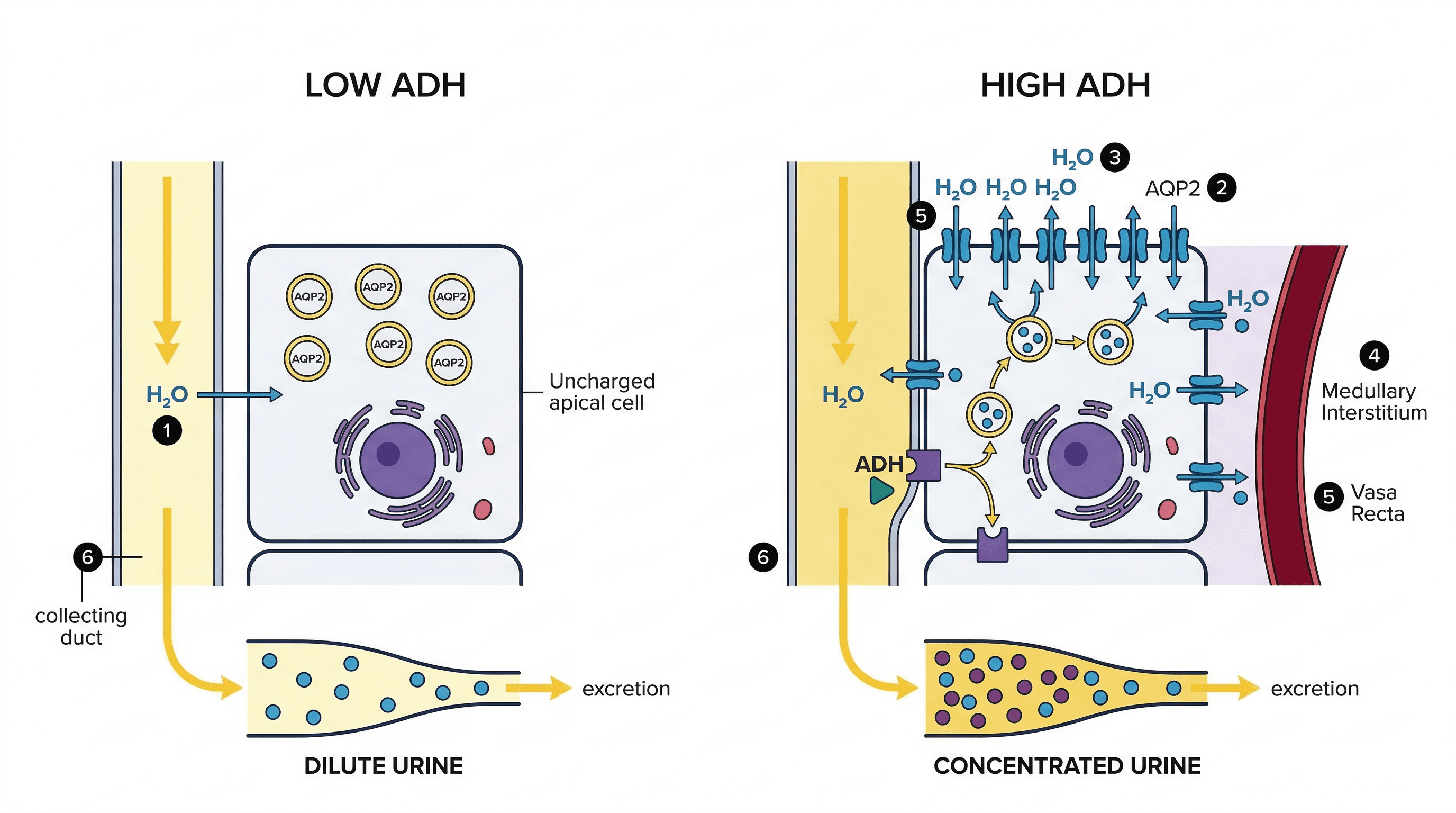
Be precise about ADH's mechanism: it binds to receptors, causing aquaporins to be inserted into the cell surface membranes of the collecting duct and DCT.
The kidneys control blood water potential through osmoregulation, a process coordinated by osmoreceptors in the hypothalamus and the hormone ADH. When blood water potential decreases, osmoreceptors stimulate the posterior pituitary gland to release ADH. ADH increases the permeability of the distal convoluted tubule and collecting duct to water, leading to increased water reabsorption and the production of more concentrated urine.
islet of Langerhans — A group of cells in the pancreas which secrete glucagon and insulin.
These endocrine clusters within the pancreas contain alpha cells (secreting glucagon) and beta cells (secreting insulin), which are crucial for the homeostatic control of blood glucose concentration. Like tiny islands of specialized factories within a larger organ, the islets produce the specific hormones needed to regulate blood sugar.
insulin — A small peptide hormone secreted by the β cells in the islets of Langerhans in the pancreas to bring about a decrease in the concentration of glucose in the blood.
Insulin stimulates liver, muscle, and adipose tissue cells to increase glucose uptake from the blood, convert glucose to glycogen (glycogenesis), and increase glucose use in respiration, thereby lowering blood glucose. Like a 'key' that unlocks cells to let glucose in, insulin helps remove excess glucose from the bloodstream.
glycogenesis — Synthesis of glycogen by addition of glucose monomers.
This process is stimulated by insulin when blood glucose levels are high, converting excess glucose into glycogen for storage primarily in the liver and muscle cells. It is a key mechanism for lowering blood glucose concentration.
glucagon — A small peptide hormone secreted by the α cells in the islets of Langerhans in the pancreas to bring about an increase in the concentration of glucose in the blood.
Glucagon primarily acts on liver cells, stimulating glycogenolysis (breakdown of glycogen) and gluconeogenesis (formation of new glucose) to release glucose into the bloodstream when blood glucose levels are low. Like an 'emergency release' button for stored energy, glucagon signals the liver to release glucose when blood sugar drops.
glycogenolysis — The breakdown of glycogen by removal of glucose monomers.
This process is stimulated by glucagon when blood glucose levels are low, releasing stored glucose from glycogen in the liver into the bloodstream. It is a crucial mechanism for increasing blood glucose concentration.
gluconeogenesis — The formation of glucose in the liver from non-carbohydrate sources such as amino acids, pyruvate, lactate, fatty acids and glycerol.
This process is stimulated by glucagon when blood glucose levels are low and glycogen stores are depleted, providing an alternative source of glucose to maintain blood sugar. It is vital for long-term glucose regulation.
adenylyl cyclase — Enzyme that catalyses formation of the second messenger cyclic AMP.
Adenylyl cyclase is a key enzyme in cell signalling pathways, particularly in the response to hormones like glucagon. Its activation leads to the production of cAMP, which then amplifies the hormonal signal within the cell.
cyclic AMP (c-AMP) — A second messenger in cell-signalling pathways.
Cyclic AMP acts as an intracellular signal, relaying and amplifying the message from a hormone (first messenger) that binds to a receptor on the cell surface. It triggers a cascade of events inside the cell, leading to the final physiological response.
protein kinase A — Enzyme that is activated by c-AMP and once activated adds phosphate groups to other proteins, including phosphorylase kinase, to activate them.
Protein kinase A is a crucial enzyme in many cell signalling pathways. Its activation by cAMP initiates a phosphorylation cascade, where it modifies other proteins by adding phosphate groups, thereby altering their activity and leading to a cellular response.
phosphorylase kinase — An enzyme that is part of the enzyme cascade that acts in response to glucagon; the enzyme activates glycogen phosphorylase by adding a phosphate group.
Phosphorylase kinase plays a specific role in the glucagon signalling pathway. Upon activation by protein kinase A, it phosphorylates and activates glycogen phosphorylase, which is the enzyme directly responsible for breaking down glycogen into glucose.
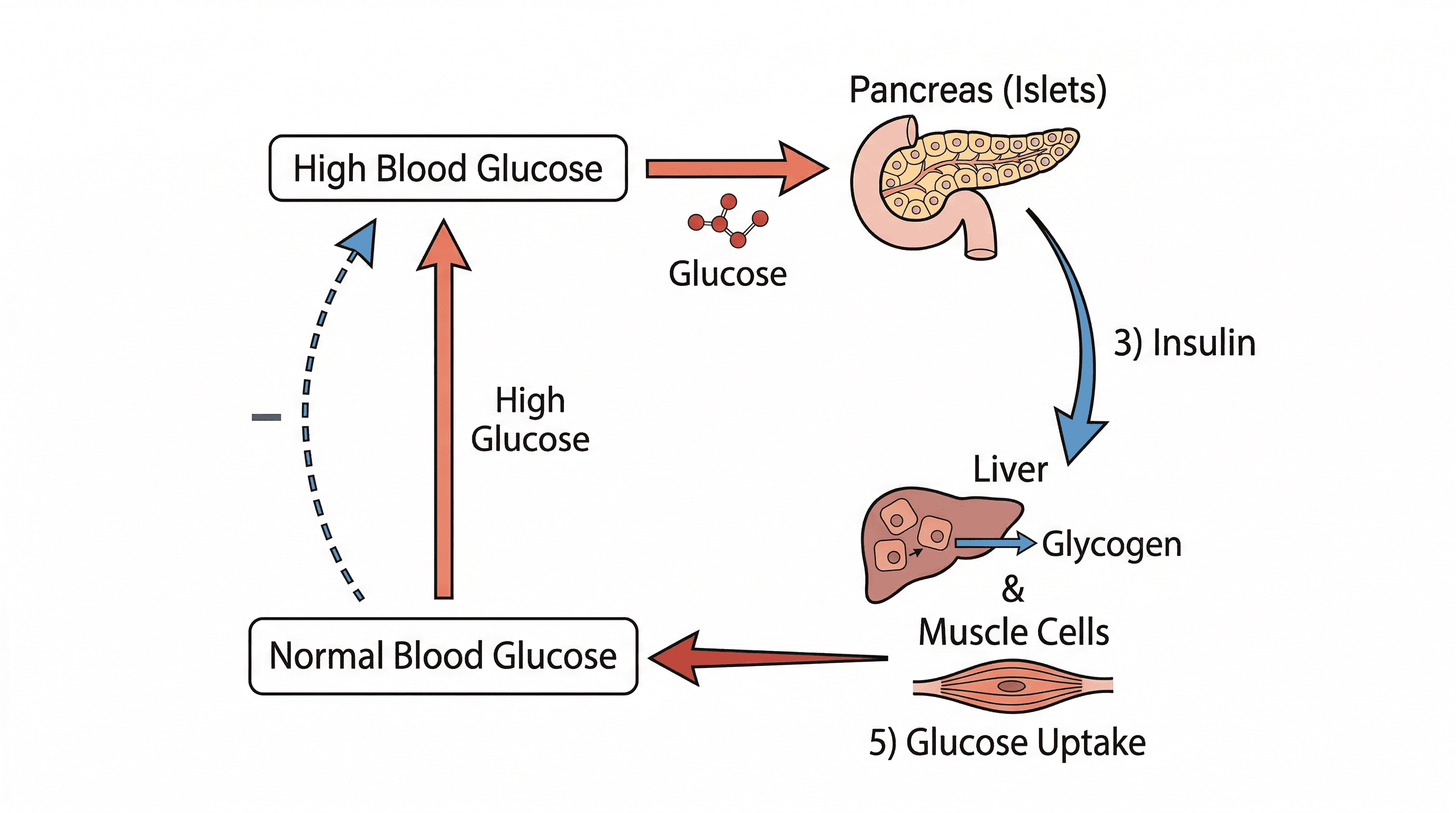
Blood glucose concentration is tightly controlled by the hormones insulin and glucagon, secreted by the islets of Langerhans in the pancreas. When blood glucose is high, beta cells release insulin, which promotes glucose uptake by liver, muscle, and adipose cells, and stimulates glycogenesis. When blood glucose is low, alpha cells release glucagon, which acts on the liver to stimulate glycogenolysis and gluconeogenesis, releasing glucose into the blood.
Students often think glucagon acts on muscle cells, but actually, muscle cells do not have glucagon receptors; its primary target for glucose release is the liver.
Clearly distinguish between insulin and glucagon effects. Use key terms: insulin promotes glycogenesis (glucose to glycogen), while glucagon promotes glycogenolysis (glycogen to glucose).
biosensor — A device that uses a biological material such as an enzyme to measure the concentration of a chemical compound.
Biosensors are used for measuring blood and urine glucose, often employing enzymes like glucose oxidase and peroxidase. They provide a rapid and accurate way to monitor physiological parameters. Their core function relies on a biological component, typically an enzyme, to detect the target substance.
Students often think biosensors are purely electronic, but actually, their core function relies on a biological component, typically an enzyme.
Test strips and biosensors are commonly used to measure glucose concentration in blood and urine. Biosensors utilise biological materials, typically enzymes like glucose oxidase and peroxidase, to detect and quantify glucose. This allows for rapid and accurate monitoring, crucial for managing conditions like diabetes.
guard cell — A kidney-shaped epidermal cell found with another, in a pair surrounding a stoma and controlling its opening or closure.
Guard cells regulate the stomatal aperture, balancing carbon dioxide entry for photosynthesis with water loss by transpiration. Their turgor changes, influenced by ion movement and water potential, dictate whether the stoma is open or closed.
electrochemical gradient — A gradient across a cell surface membrane that involves both a difference in concentrations of ions and a potential difference.
This gradient is crucial for the movement of ions, such as potassium ions, into and out of guard cells, which in turn affects their turgor and the opening or closing of stomata. It represents the combined force driving ion movement across a membrane.
abscisic acid (ABA) — An inhibitory plant growth regulator that causes closure of stomata in dry conditions.
Abscisic acid plays a vital role in plant water conservation. During water shortage, ABA signals guard cells to close stomata, reducing transpiration and preventing excessive water loss, thus helping the plant cope with drought stress.
Plants also exhibit homeostatic mechanisms, particularly in regulating stomatal aperture to balance carbon dioxide entry for photosynthesis with water loss by transpiration. Guard cells, surrounding the stomata, control their opening and closing. Stomata have daily rhythms and respond to environmental changes, such as light and water availability. During water shortage, the plant hormone abscisic acid (ABA) triggers stomatal closure to conserve water.
Use specific terminology accurately, such as 'water potential gradient', 'selective reabsorption', and 'deamination' to maximise marks.
Definitions Bank
homeostasis
The maintenance of a relatively constant internal environment for the cells within the body.
negative feedback
A process in which a change in some parameter (e.g. blood glucose concentration) brings about processes which return it towards normal.
receptor
A cell or tissue that is sensitive to a specific stimulus and communicates with a control centre by generating nerve impulses or sending a chemical messenger.
effector
A tissue or organ that carries out an action in response to a stimulus; muscles and glands are effectors.
stimulus
A change in the external or internal environment that is detected by a receptor and which may cause a response.
+36 more definitions
View all →Command Word Guide
| Explain | When explaining the importance of homeostasis, link it directly to enzyme activity and metabolic efficiency, mentioning denaturation at extremes. For ADH action, explain the mechanism: binding to receptors, cAMP production, vesicle fusion, aquaporin insertion, and increased water permeability. |
| Describe | When describing negative feedback, clearly identify the stimulus, receptor, control centre, effector, and the corrective action that reverses the initial change. For deamination, specify that it occurs in the liver, involves the removal of the amine group, and produces ammonia (which is then converted to urea) and a keto acid. |
| State | State that urea is produced by deamination of excess amino acids in the liver. State the location (hypothalamus) and specific function (detecting blood water potential) of osmoreceptors. |
| Identify | When identifying effectors, name the specific organ or tissue (e.g., liver cells, muscle cells, collecting ducts) and the action they perform. Be able to label and describe the function of each part of the nephron and its associated blood vessels. |
Common Mistakes
Thinking homeostasis means conditions are absolutely constant.
Homeostasis maintains conditions that fluctuate slightly around a set point, not absolute constancy.
Confusing negative feedback with a 'bad' response.
Negative feedback refers to a response that reverses the initial change, maintaining stability, which is beneficial.
Confusing excretion with egestion.
Excretion is the removal of metabolic waste products (e.g., urea), while egestion is the removal of undigested food.
+3 more
View all →This chapter explores the intricate mechanisms of control and coordination in living organisms, focusing on the endocrine and nervous systems in mammals, and electrical and chemical communication in plants. It details the structure and function of neurones, the transmission of nerve impulses via action potentials and synapses, and the ultrastructure and contraction mechanism of striated muscle. Additionally, it covers rapid plant responses and the roles of plant growth regulators such as auxins and gibberellins.
endocrine system — Consists of all the endocrine glands in the body together with the hormones that they secrete.
This system is one of the body's main communication and coordination systems, working alongside the nervous system to regulate various physiological processes, often over longer durations. It is like the body's slow-acting, widespread postal service, delivering chemical messages (hormones) to many different locations.
endocrine gland — An organ that secretes hormones directly into the blood; endocrine glands are also known as ductless glands.
These glands are part of the endocrine system and release their chemical messengers (hormones) into the bloodstream, allowing them to travel to distant target cells throughout the body. This contrasts with exocrine glands, which secrete substances through ducts. Think of an endocrine gland as a radio station broadcasting a signal (hormone) to anyone with a receiver (receptor) tuned to that frequency, rather than sending a letter (exocrine) to a specific address.
When asked to describe endocrine glands, always mention 'ductless' and 'secrete hormones directly into the blood' for full marks.
Students often think endocrine glands have ducts, but actually they are ductless and secrete directly into the blood.
neurone — A nerve cell; a cell which is specialised for the conduction of nerve impulses.
Neurones are the basic functional units of the nervous system, designed to transmit electrical signals rapidly over long distances to coordinate body activities. A neurone is like a telegraph wire, specifically designed to carry electrical signals (impulses) from one point to another very quickly.
nerve impulse — (usually shortened to impulse) a wave of electrical depolarisation that is transmitted along neurones.
This is the fundamental unit of information transfer in the nervous system, involving rapid changes in the electrical potential across the neurone's cell surface membrane due to ion movement. A nerve impulse is like a ripple effect in a pond, where the disturbance (depolarisation) travels along the surface (neurone membrane) without the water itself moving far.
Students often think a nerve impulse is a flow of electrons like an electric current, but actually it's a wave of electrical depolarisation caused by ion movement.
Avoid using 'electrical current' when describing nerve impulses; instead, use 'wave of electrical depolarisation' or 'action potential' and mention ion movement.
The endocrine and nervous systems are the body's main communication and coordination systems. The endocrine system uses chemical messengers (hormones) transported by the blood, resulting in slower, widespread, and often longer-lasting responses. In contrast, the nervous system uses electrical impulses transmitted along neurones, leading to rapid, targeted, and short-lived responses.
When comparing the endocrine and nervous systems, focus on the speed and duration of response, and the mode of transport (blood vs. electrical impulses).
sensory neurone — A neurone that transmits nerve impulses from a receptor to the central nervous system (CNS).
These neurones are responsible for conveying information about internal and external stimuli from sensory receptors to the brain and spinal cord for processing. A sensory neurone is like a security camera cable, sending signals from a sensor (receptor) to the central monitoring station (CNS).
motor neurone — A neurone whose cell body is in the brain, spinal cord or a ganglion (a swelling on a nerve), and that transmits nerve impulses to an effector such as a muscle or gland.
Motor neurones are crucial for initiating responses by carrying commands from the CNS to muscles or glands, causing them to contract or secrete. A motor neurone is like a control wire from a central command center (CNS) to a robot's arm (effector), telling it to move.
intermediate neurone — A neurone that transmits nerve impulses between other neurones, typically within the central nervous system (CNS).
Intermediate neurones, also known as relay neurones, connect sensory and motor neurones, allowing for complex processing and integration of signals within the CNS. They are essential for reflex arcs and higher-order brain functions.
Remember that sensory neurones carry information 'to' the CNS, while motor neurones carry information 'from' the CNS.
Students often think all neurones are the same, but actually there are different types (sensory, motor, intermediate) with distinct structures and functions.
myelin — Insulating material that surrounds the axons of many neurones; myelin is made of layers of cell surface membranes formed by Schwann cells so that they are very rich in phospholipids and therefore impermeable to water and ions in tissue fluid.
Myelin acts as an electrical insulator, significantly increasing the speed of nerve impulse conduction by allowing action potentials to 'jump' between nodes of Ranvier. Myelin is like the plastic insulation around an electrical wire, preventing signal leakage and allowing the electrical impulse to travel much faster.
node of Ranvier — A very short gap between Schwann cells where myelinated axons are not covered in myelin so are exposed to tissue fluid.
These gaps are critical for saltatory conduction, as action potentials are regenerated only at these points, allowing the impulse to 'jump' from node to node. Nodes of Ranvier are like stepping stones across a river; the impulse jumps from one stone to the next, rather than having to wade through the entire river.
Explain that myelin is rich in phospholipids, making it impermeable to ions, and that it enables saltatory conduction for faster impulse transmission.
Students often think myelin is a continuous sheath, but actually it has small gaps called nodes of Ranvier where depolarisation occurs.
The resting potential is the electrical potential difference across the cell surface membrane of a neurone when it is not transmitting an action potential, typically around -70 mV inside. This negative charge is established and maintained by the sodium-potassium pump, which actively transports three sodium ions out for every two potassium ions in, along with the differential permeability of the membrane to ions and the presence of large organic anions inside the cell. This state prepares the neurone to fire an action potential.
potential difference — The difference in electrical potential between two points; in the nervous system, between the inside and the outside of a cell surface membrane such as the membrane that encloses an axon.
This electrical gradient is crucial for nerve impulse transmission, as changes in potential difference across the membrane drive the opening and closing of voltage-gated ion channels. Potential difference is like the height difference between two points on a hill; it creates a 'drive' for things (like ions) to move from higher to lower potential.
resting potential — The difference in electrical potential that is maintained across the cell surface membrane of a neurone when it is not transmitting an action potential; it is normally about –70 mV inside and is partly maintained by sodium–potassium pumps.
This negative charge inside the neurone is established and maintained by the sodium-potassium pump, differential ion permeability, and the presence of large organic anions, preparing the neurone to fire an action potential. The resting potential is like a loaded spring, storing potential energy (electrical charge) that can be released quickly when triggered (by a stimulus).
Students often think the resting potential is solely due to the sodium-potassium pump, but actually the differential permeability of the membrane to ions and large intracellular anions also contribute significantly.
When explaining resting potential, mention the roles of the sodium-potassium pump, the impermeability of the membrane to sodium ions, and the presence of organic anions.
A nerve impulse is transmitted as an action potential, a rapid, brief change in the potential difference across the neurone membrane. This 'all-or-none' event is triggered when a stimulus causes the membrane potential to reach a critical threshold potential. Once initiated, the action potential propagates along the neurone, involving sequential depolarisation and repolarisation.
action potential — A brief change in the potential difference from –70 mV to +30 mV across the cell surface membranes of neurones and muscle cells caused by the inward movement of sodium ions.
This is the electrical signal transmitted along a neurone, involving a rapid depolarisation followed by repolarisation, driven by the sequential opening and closing of voltage-gated ion channels. An action potential is like flipping a light switch on and off very quickly; the voltage rapidly changes from negative to positive and back again.
voltage-gated channel protein — A channel protein through a cell membrane that opens or closes in response to changes in electrical potential across the membrane.
These proteins are fundamental to action potential generation and propagation, as their opening and closing in response to depolarisation allows the rapid influx and efflux of ions. A voltage-gated channel protein is like an automatic door that only opens when a specific electrical 'key' (voltage change) is detected.
depolarisation — The reversal of the resting potential across the cell surface membrane of a neurone or muscle cell, so that the inside becomes positively charged compared with the outside.
This is the initial phase of an action potential, caused by the rapid influx of sodium ions through voltage-gated channels, making the inside of the membrane less negative and then positive. Depolarisation is like a sudden surge of water into a dry well, quickly filling it up and even overflowing, changing its 'potential' from empty to full.
threshold potential — The critical potential difference across the cell surface membrane of a sensory receptor or neurone which must be reached before an action potential is initiated.
This 'all-or-none' principle means that if the stimulus is too weak to reach this threshold, no action potential will be generated, ensuring that only significant stimuli trigger responses. The threshold potential is like the minimum pressure needed to push a button; if you don't push hard enough, nothing happens, but once you reach that pressure, the button activates fully.
repolarisation — Returning the potential difference across the cell surface membrane of a neurone or muscle cell to normal following the depolarisation of an action potential.
This phase involves the efflux of potassium ions through voltage-gated channels, restoring the negative charge inside the membrane and preparing the neurone for another action potential. Repolarisation is like draining the overflowing well back to its original dry state, restoring its 'potential' to be filled again.
refractory period — A period of time during which a neurone is recovering from an action potential, and during which another action potential cannot be generated.
This period ensures that action potentials are discrete events, travel in one direction only, and limits the maximum frequency at which impulses can be transmitted. The refractory period is like the cool-down time for a camera flash; you can't take another picture immediately after one flash, you have to wait for it to recharge.
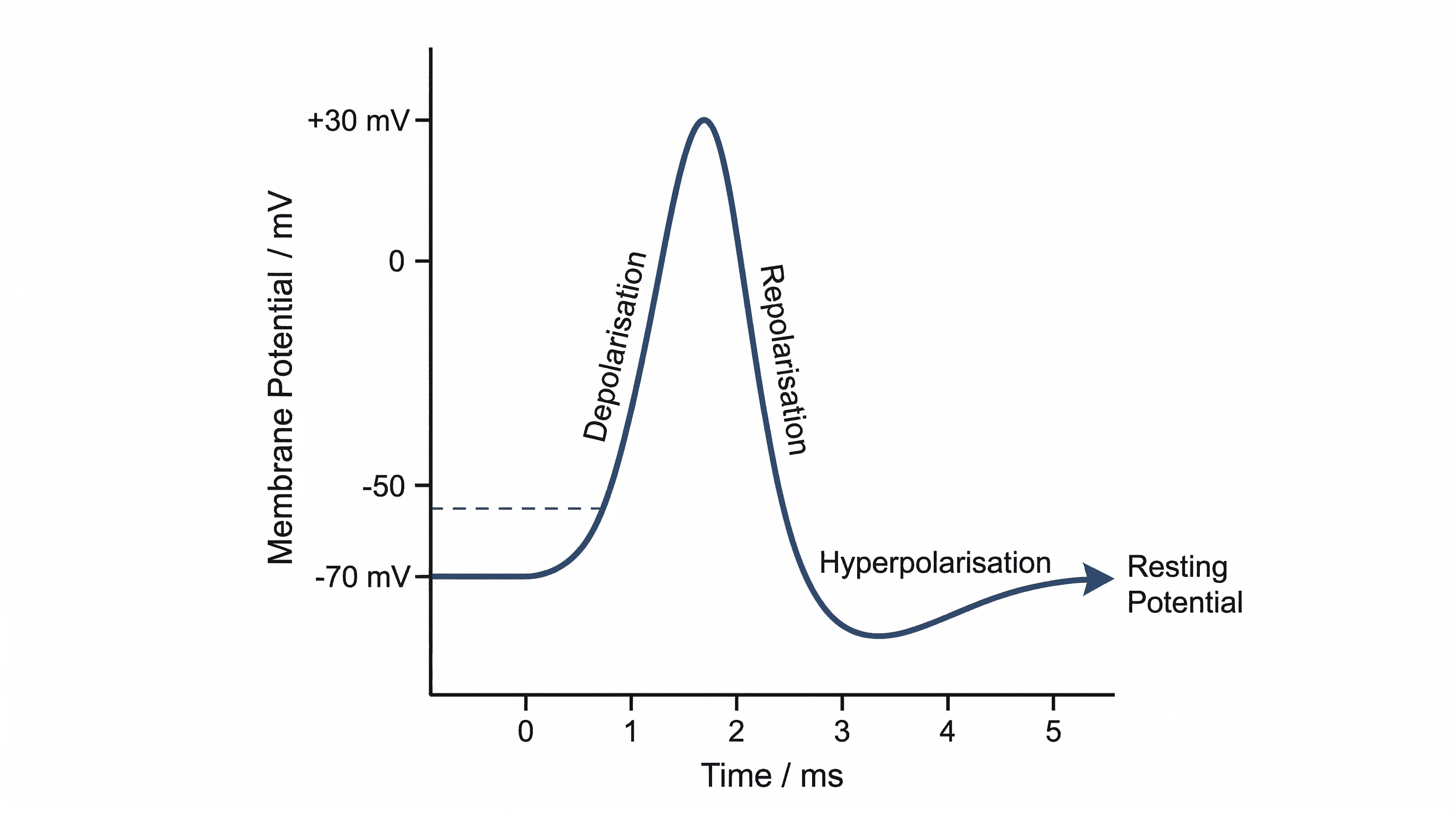
Clearly state the voltage changes (e.g., -70mV to +30mV) and the primary ion responsible for depolarisation (sodium ions) when describing an action potential.
Students often think action potentials vary in size with stimulus strength, but actually they are 'all-or-none' events with a constant amplitude; only their frequency changes.
all-or-none law — Neurones and muscle cells only transmit impulses if the initial stimulus is sufficient to increase the membrane potential above a threshold potential.
This principle dictates that once the threshold is reached, an action potential of a fixed amplitude is generated, regardless of further increases in stimulus strength; information about stimulus intensity is encoded by frequency. The all-or-none law is like flushing a toilet; once you push the handle past a certain point, the flush happens completely, regardless of how much harder you push.
Emphasise the 'all-or-none' nature of action potentials in relation to the threshold potential; a sub-threshold stimulus produces no action potential.
The speed of nerve impulse conduction is significantly increased in myelinated neurones through a process called saltatory conduction. Myelin acts as an electrical insulator, preventing ion flow across the membrane except at the nodes of Ranvier. This allows the action potential to 'jump' from one node to the next, rather than propagating continuously along the entire axon.
saltatory conduction — Movement of an action potential along a myelinated axon, in which the action potential ‘jumps’ from one node of Ranvier to the next.
This mechanism significantly increases the speed of impulse transmission in myelinated neurones compared to unmyelinated ones, as depolarisation only occurs at the nodes. Saltatory conduction is like skipping steps on a staircase instead of taking each one; it's a much faster way to get from top to bottom.
Explain that myelin insulates the axon, preventing ion flow, and that voltage-gated channels are concentrated at the nodes, allowing the 'jumping' effect.
Students often think saltatory conduction means the impulse literally jumps through the air, but actually it means the action potential is regenerated only at the nodes of Ranvier.
chemoreceptor — A receptor cell that responds to chemical stimuli; chemoreceptors are found in taste buds on the tongue, in the nose and in blood vessels where they detect changes in oxygen and carbon dioxide concentrations.
These specialised cells convert chemical signals into electrical impulses, enabling senses like taste and smell, and playing vital roles in homeostatic regulation by monitoring blood chemistry. A chemoreceptor is like a chemical sensor that detects specific molecules in its environment and then sends an alarm signal (electrical impulse) when it finds them.
receptor potential — A change in the normal resting potential across the membrane of a receptor cell, caused by a stimulus.
This is a graded potential, meaning its magnitude depends on the strength of the stimulus; if it reaches a threshold, it can trigger an action potential in an associated sensory neurone. A receptor potential is like the initial push on a swing; a small push causes a small swing, but a strong push can make it swing high enough to trigger a larger event (like an action potential).
Students often think receptor potentials are always action potentials, but actually receptor potentials are graded and only trigger action potentials if they reach a threshold.
Synapses are crucial junctions where nerve impulses are transmitted from one neurone to another, or from a neurone to an effector. Unlike direct electrical connections, synapses involve a small gap called the synaptic cleft, across which chemical messengers called neurotransmitters diffuse. This chemical transmission allows for signal integration, modulation, and ensures unidirectional flow of information.
synapse — A point at which two neurones meet but do not touch; the synapse is made up of the end of the presynaptic neurone, the synaptic cleft and the end of the postsynaptic neurone.
Synapses are crucial for information processing in the nervous system, allowing for one-way transmission, integration of multiple signals, and modulation of nerve impulses. A synapse is like a junction box in an electrical circuit, where signals can be routed, amplified, or inhibited before being passed on.
synaptic cleft — A very small gap between two neurones at a synapse; nerve impulses are transmitted across synaptic clefts by neurotransmitters.
This physical separation prevents direct electrical transmission, necessitating chemical transmission via neurotransmitters, which allows for signal integration and modulation. The synaptic cleft is like a small river separating two banks; the message (neurotransmitter) has to be ferried across to reach the other side.
Students often think neurones touch at a synapse, but actually there is a distinct gap, the synaptic cleft, which neurotransmitters must cross.
neurotransmitter — A chemical released at synapses to transmit impulses between neurones or between a motor neurone and a muscle fibre.
These chemical messengers diffuse across the synaptic cleft and bind to receptors on the postsynaptic membrane, causing a change in its potential and potentially generating a new action potential. A neurotransmitter is like a chemical key that unlocks a specific door (receptor) on the next cell, allowing a message to pass through.
presynaptic neurone — A neurone ending at a synapse from which neurotransmitter is released when an action potential arrives.
This neurone transmits the signal towards the synapse, releasing neurotransmitters into the synaptic cleft upon the arrival of an action potential. The presynaptic neurone is like the sender of a text message, initiating the communication by releasing the message (neurotransmitter).
postsynaptic neurone — The neurone on the opposite side of a synapse to the neurone in which the action potential arrives.
This neurone receives the neurotransmitter signal, which can cause depolarisation and potentially trigger its own action potential, propagating the impulse. The postsynaptic neurone is like the receiver of a text message, interpreting the message (neurotransmitter binding) and deciding how to respond.
voltage-gated calcium ion channel protein — A channel protein in presynaptic membranes that responds to depolarisation by opening to allow diffusion of calcium ions down their electrochemical gradient.
The influx of calcium ions through these channels into the presynaptic terminal is the critical trigger for the exocytosis of neurotransmitter vesicles. This channel is like a gate that only opens when an electrical signal arrives, letting in calcium ions which then act as a signal to release neurotransmitters.
receptor protein — A protein on a postsynaptic membrane that is a ligand-gated channel protein opening in response to binding of a neurotransmitter.
These proteins specifically bind neurotransmitters, leading to a conformational change that opens an ion channel, allowing ions to flow across the postsynaptic membrane and alter its potential. A receptor protein is like a specific lock on the postsynaptic membrane that only opens when the correct key (neurotransmitter) fits into it.
acetylcholine (ACh) — A type of neurotransmitter released by cholinergic synapses.
ACh is a common neurotransmitter involved in muscle contraction (at neuromuscular junctions), learning, and memory, and its action is rapidly terminated by acetylcholinesterase. ACh is like a specific key for muscle contraction; when it binds to the lock (receptor), the muscle is told to contract.
cholinergic synapse — A synapse at which the transmitter substance is ACh.
These synapses are prevalent in the peripheral nervous system, particularly at neuromuscular junctions, and are critical for voluntary muscle control. A cholinergic synapse is like a specific type of communication channel that only uses ACh as its language.
acetylcholinesterase — An enzyme in the synaptic cleft and on the postsynaptic membrane that hydrolyses ACh to acetate and choline.
This enzyme rapidly breaks down acetylcholine, ensuring that the neurotransmitter's effect is brief and allowing the postsynaptic membrane to repolarise and be ready for subsequent signals. Its action is crucial for precise control of muscle contraction and nerve impulse transmission.
noradrenaline — A type of neurotransmitter, which is also released by cells in the adrenal glands as a hormone.
Noradrenaline acts as both a neurotransmitter in the nervous system and a hormone in the endocrine system, playing roles in the 'fight or flight' response, alertness, and mood. Noradrenaline is like a versatile messenger that can be delivered quickly and locally (neurotransmitter) or broadcast widely through the bloodstream (hormone).
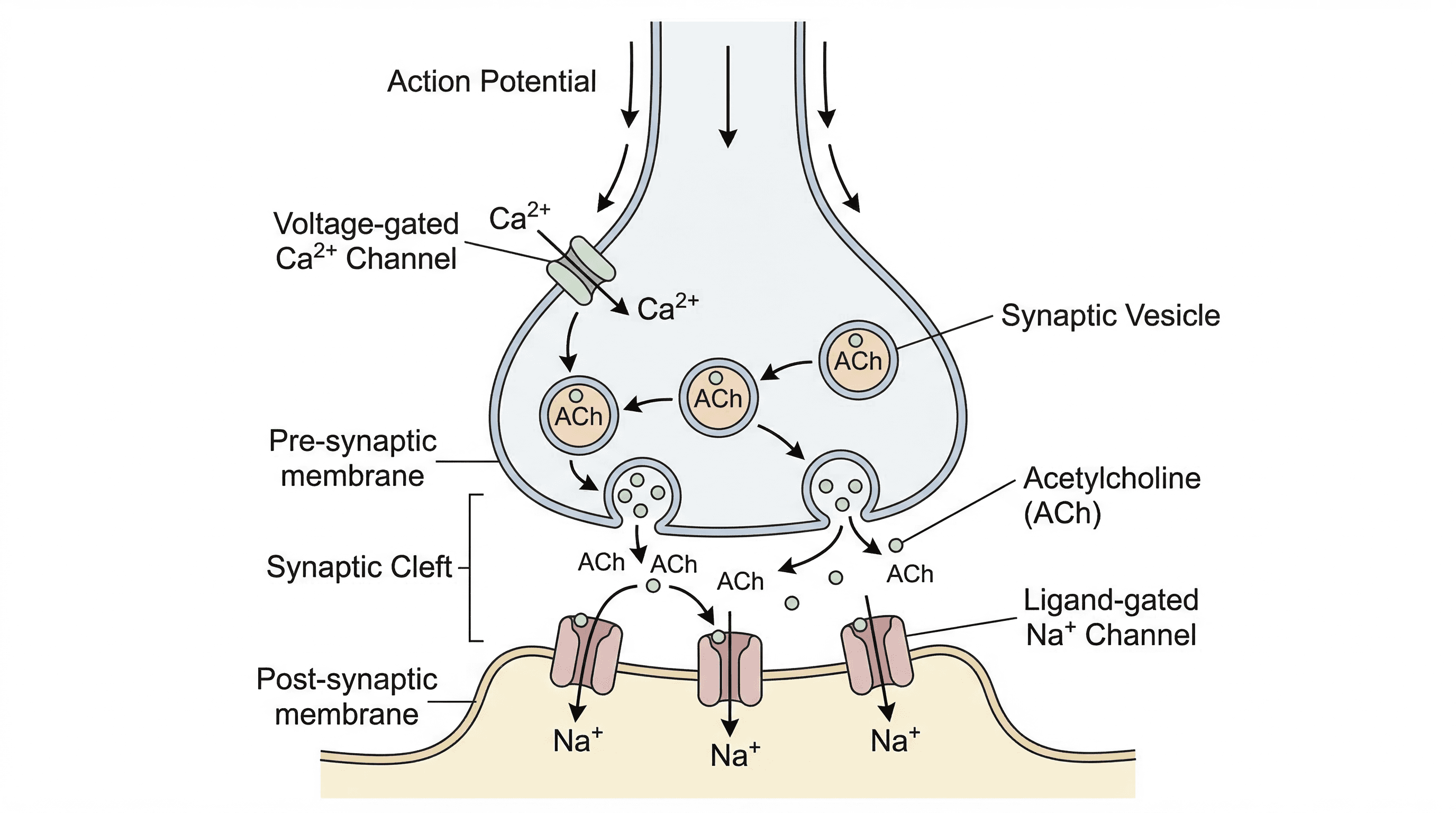
Always mention acetylcholinesterase when discussing ACh, as its role in breaking down ACh is crucial for proper synaptic function.
Muscle contraction is a complex process initiated by nerve impulses from motor neurones. Striated muscle, found in skeletal muscles, exhibits a characteristic banded appearance due to the organised arrangement of contractile proteins. The contraction itself occurs via the sliding filament model, where thick and thin filaments slide past each other, shortening the sarcomere.
neuromuscular junction — A synapse between a motor neurone and a muscle.
This specialised synapse is where the motor neurone transmits its electrical signal to the muscle fibre, initiating muscle contraction. It is a type of cholinergic synapse, using acetylcholine as its neurotransmitter.
striated muscle — Type of muscle tissue in skeletal muscles; the muscle fibres have regular striations that can be seen under the light microscope.
Striated muscle is responsible for voluntary movements and is characterised by its organised structure, which gives it a striped appearance under a microscope. This organisation is key to its efficient contractile function.
sarcolemma — The cell surface membrane of a muscle fibre.
The sarcolemma is the plasma membrane of a muscle cell, which plays a crucial role in transmitting the electrical impulse from the neuromuscular junction throughout the muscle fibre via T-tubules, initiating contraction.
sarcoplasm — The cytoplasm of muscle cells.
The sarcoplasm is the specialised cytoplasm of muscle fibres, containing numerous mitochondria for ATP production, glycogen stores, and a high concentration of calcium ions, which are essential for muscle contraction.
sarcoplasmic reticulum (SR) — The endoplasmic reticulum of a muscle fibre.
The sarcoplasmic reticulum is a specialised network of membranes within muscle cells that stores and releases calcium ions, which are critical for triggering muscle contraction. It plays a central role in regulating the availability of calcium for the contractile proteins.
transverse system tubule (or T-system tubule or T-tubule) — Infolding of the sarcolemma that go deep into a muscle fibre and conducts impulses to the SR.
T-tubules are invaginations of the sarcolemma that extend deep into the muscle fibre, allowing the action potential to rapidly reach all parts of the muscle cell, including the sarcoplasmic reticulum, ensuring coordinated contraction.
myofibril — One of many cylindrical bundles of thick (myosin) and thin (actin) filaments inside a muscle fibre.
Myofibrils are the contractile units within muscle fibres, composed of repeating units called sarcomeres. Their organised arrangement of actin and myosin filaments is responsible for muscle contraction.
myosin — The protein that makes up the thick filaments in striated muscle; the globular heads of each molecule break down ATP (they act as an ATP-ase).
Myosin is a motor protein that forms the thick filaments in muscle. Its globular heads bind to actin and use ATP hydrolysis to generate the force for muscle contraction, pulling the thin filaments.
actin — The protein that makes up the thin filaments in striated muscle.
Actin is a globular protein that polymerises to form the thin filaments in muscle. It provides the binding sites for myosin heads during muscle contraction.
sarcomere — The part of a myofibril between two Z discs.
The sarcomere is the fundamental contractile unit of striated muscle, extending from one Z-disc to the next. It contains the organised arrangement of actin and myosin filaments that slide past each other during contraction.
tropomyosin — A fibrous protein that is part of the thin filaments in myofibrils in striated muscle; tropomyosin blocks the attachment site on the thin filament for myosin heads so preventing the formation of cross-bridges.
Tropomyosin is a regulatory protein that, in resting muscle, covers the myosin-binding sites on actin, preventing contraction. Its movement, triggered by calcium, exposes these sites, allowing myosin to bind.
troponin — A calcium-binding protein that is part of the thin filaments in myofibrils in striated muscle.
Troponin is a complex of three proteins that binds to calcium ions. This binding causes a conformational change in troponin, which in turn moves tropomyosin away from the myosin-binding sites on actin, initiating muscle contraction.
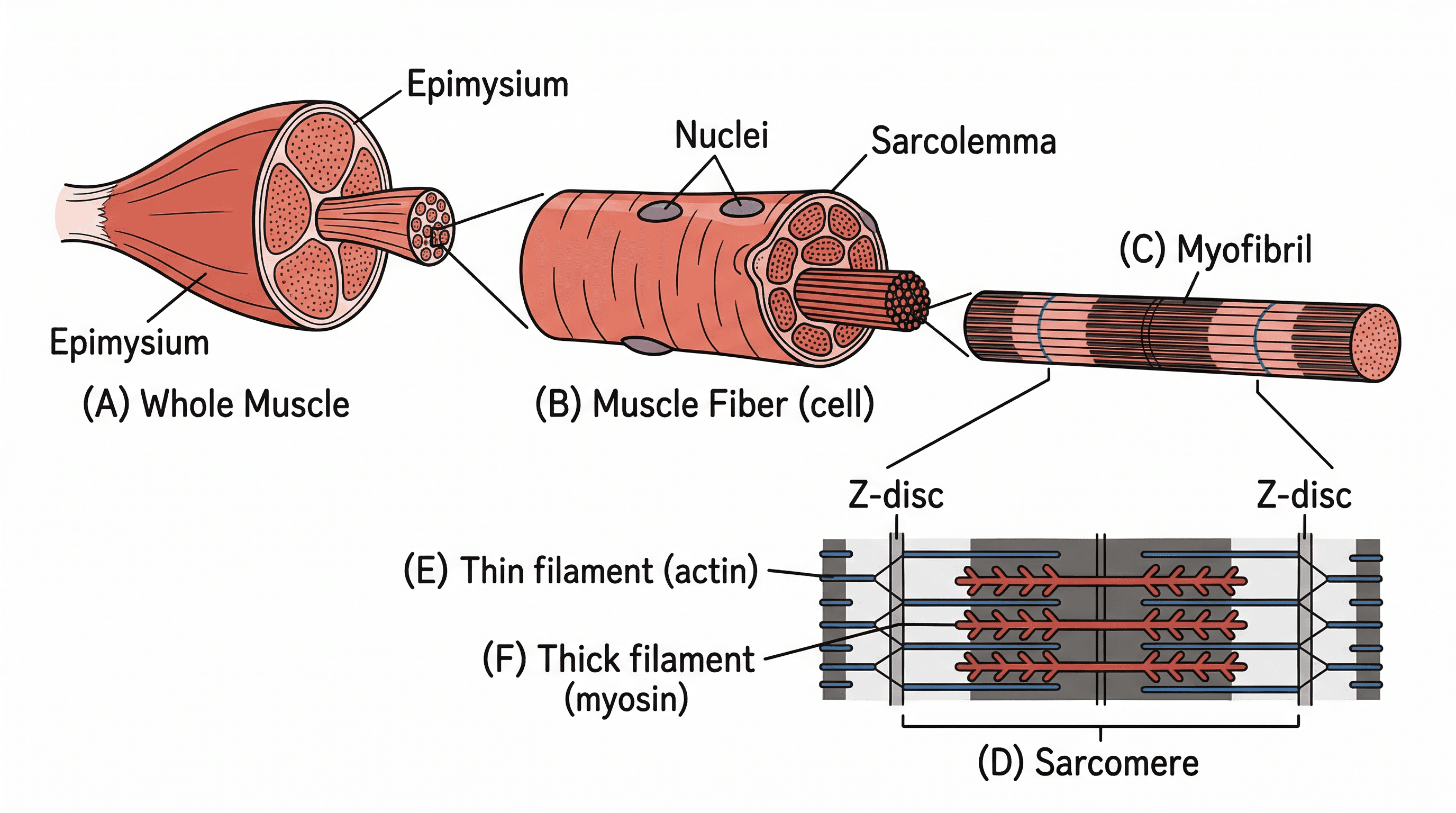
sliding filament model — The mechanism of muscle contraction; within each sarcomere the movement of thin filaments closer together by the action of myosin heads in the thick filaments shortens the overall length of each muscle fibre.
This model explains how muscle contraction occurs: myosin heads bind to actin filaments, form cross-bridges, and then pivot, pulling the actin filaments towards the centre of the sarcomere. This process shortens the sarcomere without the individual filaments themselves changing length.
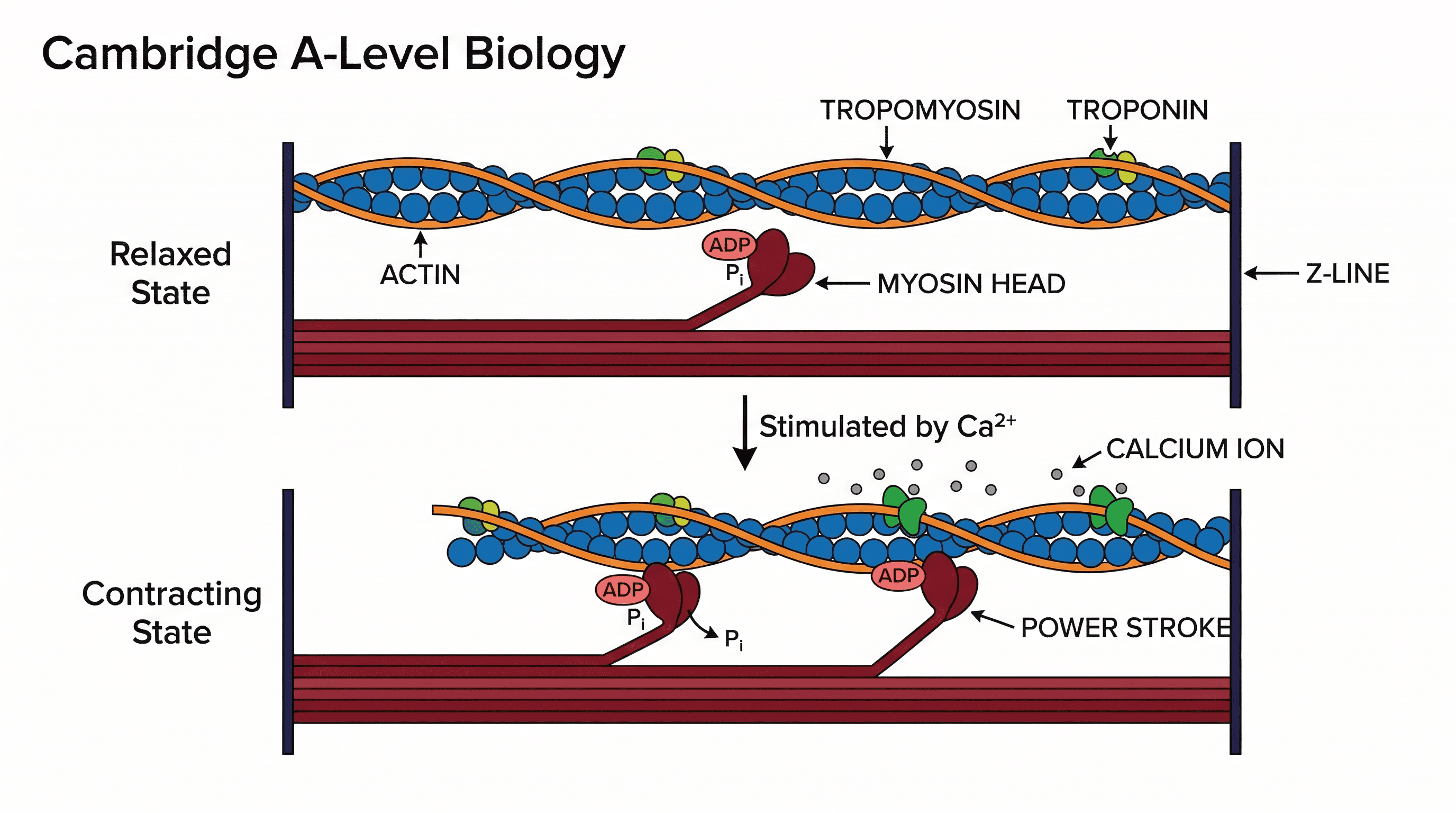
Students often think the actin and myosin filaments shorten during contraction, but actually they maintain their length and slide past each other.
For muscle contraction, be able to label a sarcomere diagram (Z-line, I-band, A-band, H-zone) and clearly state which bands shorten (I-band, H-zone) and which stay the same length (A-band).
Plants also exhibit control and coordination, though their mechanisms differ from animals. They respond to stimuli using both electrical and chemical communication. Rapid responses, such as the Venus fly trap closing, involve electrical signals. Chemical communication relies on plant growth regulators, often called plant hormones, which influence growth and development.
Students often think plant hormones are produced in specialised glands like animal hormones, but actually they are produced in various tissues throughout the plant.
plant growth regulator — (plant hormone) any chemical produced in plants that influences their growth and development (e.g. auxins, gibberellins, cytokinins and ABA).
These chemical messengers are produced in various plant tissues and regulate a wide range of physiological processes, including cell division, elongation, differentiation, and responses to environmental cues. They are crucial for coordinating plant growth and development.
auxin — A plant growth regulator (plant hormone) that stimulates cell elongation.
Auxins are primarily involved in cell elongation, particularly in shoots, and play roles in phototropism, gravitropism, and apical dominance. They promote the loosening of cell walls, allowing cells to expand.
expansins — Proteins in the cell walls of plants that loosen the attachment of microfibrils of cellulose during elongation growth.
Expansins are cell wall proteins that facilitate cell expansion by loosening the cellulose microfibril network, allowing the cell wall to stretch under turgor pressure. Auxins promote the activity of expansins, contributing to cell elongation.
gibberellin — A plant growth regulator (plant hormone) that stimulates seed germination and regulates plant height (stem growth); a lack of gibberellin causes dwarfness.
Gibberellins are a class of plant hormones involved in various developmental processes, including stem elongation, fruit development, and particularly, the breaking of seed dormancy and promotion of germination. They are crucial for normal plant growth.
endosperm — A tissue in some seeds, such as barley, that is a store of starch and other nutrients.
The endosperm serves as a primary food reserve for the developing embryo in many seeds, providing energy and building blocks until the seedling can photosynthesise independently. In barley, it is rich in starch.
aleurone layer — A layer of tissue around the endosperm that synthesises amylase during germination.
The aleurone layer is a specialised tissue in cereal grains that responds to gibberellins during germination by synthesising and secreting alpha-amylase, an enzyme that breaks down the starch in the endosperm into usable sugars for the embryo.
During barley seed germination, the embryo releases gibberellins. These gibberellins diffuse to the aleurone layer, stimulating it to synthesise and secrete alpha-amylase. Alpha-amylase then hydrolyses the starch stored in the endosperm into maltose, which is further broken down into glucose to provide energy for the growing embryo.
Always link structure to function. For example, the myelin sheath acts as an electrical insulator, and the many mitochondria in a presynaptic knob provide ATP for neurotransmitter synthesis.
Use precise terminology. Distinguish between 'neurone' (a cell) and 'nerve' (a bundle of axons). Use terms like 'presynaptic terminal', 'synaptic cleft', and 'postsynaptic membrane' correctly.
Be prepared for comparison questions. Create a table comparing nervous vs. endocrine control based on speed, duration, transmission method (neurones vs. blood), and nature of the message (electrical vs. chemical).
Definitions Bank
endocrine gland
An organ that secretes hormones directly into the blood; endocrine glands are also known as ductless glands.
endocrine system
Consists of all the endocrine glands in the body together with the hormones that they secrete.
nerve impulse
(usually shortened to impulse) a wave of electrical depolarisation that is transmitted along neurones.
neurone
A nerve cell; a cell which is specialised for the conduction of nerve impulses.
sensory neurone
A neurone that transmits nerve impulses from a receptor to the central nervous system (CNS).
+46 more definitions
View all →Command Word Guide
| Describe | Provide a detailed account of the features or process, e.g., 'Describe the features of the endocrine system' requires mentioning ductless glands, hormone secretion into blood, and widespread action. |
| Explain | Give reasons for a phenomenon or process, e.g., 'Explain the transmission of nerve impulses' requires detailing ion movements, voltage changes, and the role of channels. |
| Compare | Identify similarities and differences between two or more concepts, e.g., 'Compare the endocrine and nervous systems' requires discussing speed, duration, and mode of transmission for both. |
| Outline | Give the main features or general principles of something, e.g., 'Outline the roles of sensory receptor cells' requires a brief summary of their function in detecting stimuli and generating receptor potentials. |
Common Mistakes
Thinking a nerve impulse is a flow of electrons like an electric current.
A nerve impulse is a wave of electrical depolarisation caused by ion movement across the neurone membrane.
Believing action potentials vary in size with stimulus strength.
Action potentials are 'all-or-none' events with a constant amplitude; only their frequency changes with stimulus strength.
Assuming neurones touch at a synapse.
There is a distinct gap, the synaptic cleft, which neurotransmitters must cross to transmit the impulse.
+3 more
View all →This chapter explores the fundamental principles of inheritance, detailing how genetic information is passed from parents to offspring through meiosis and sexual reproduction. It covers Mendelian genetics, including various inheritance patterns and the use of genetic diagrams and the chi-squared test, before examining the molecular basis of gene expression and its control in both prokaryotes and eukaryotes.
sexual reproduction — Reproduction involving the fusion of gametes (fertilisation) to produce a zygote.
This process combines genetic material from two parents, leading to offspring with unique combinations of alleles. It is a key source of genetic variation in populations, much like mixing two decks of cards to create a new, unique deck.
gamete — A sex cell; during sexual reproduction, two gametes fuse together to form a zygote; gametes are usually haploid.
Gametes carry half the genetic information of a somatic cell, ensuring that when two fuse during fertilisation, the resulting zygote has the correct diploid chromosome number. Examples include sperm and egg cells, acting like half-keys that combine to unlock full genetic potential.
Students often think gametes are diploid, but actually they are haploid, containing only one set of chromosomes.
Always specify that gametes are haploid (n) and contain one allele for each gene when discussing their role in inheritance.
fertilisation — The fusing of the nuclei of two gametes, to form a zygote.
This crucial event restores the diploid chromosome number and combines genetic material from both parents, initiating the development of a new individual. It is a random process, contributing to genetic variation, much like two puzzle pieces fitting together perfectly to complete a picture.
When defining fertilisation, explicitly mention the fusion of nuclei and the formation of a zygote to gain full marks.
zygote — A cell formed by the fusion of the nuclei of two gametes; most zygotes are diploid.
The zygote is the first diploid cell of a new organism, containing a complete set of chromosomes from both parents. It undergoes repeated mitotic divisions to develop into a multicellular organism, acting like the blueprint for a new building.
diploid — Containing two complete sets of chromosomes; can be signified by the symbol 2n.
Most somatic cells in sexually reproducing organisms are diploid, meaning they have two copies of each chromosome, one inherited from each parent. This provides a backup copy of genes and allows for greater genetic diversity, like having two identical instruction manuals.
haploid — Containing one complete set of chromosomes; can be signified by the symbol n.
Gametes are haploid cells, meaning they contain only one chromosome from each homologous pair. This ensures that upon fertilisation, the diploid number is restored in the zygote. If a diploid cell is a full deck of cards, a haploid cell is half a deck.
homologous chromosomes — Two chromosomes that carry the same genes in the same positions.
These pairs of chromosomes are similar in size and shape and carry alleles for the same traits at corresponding loci. One homologous chromosome is inherited from the mother and the other from the father, like a pair of matching shoes.
Students often confuse homologous chromosomes with sister chromatids; homologous chromosomes are a pair (one from each parent) carrying the same genes, while sister chromatids are identical copies of a single chromosome joined at the centromere.
Emphasise that homologous chromosomes carry the 'same genes in the same positions' but not necessarily the 'same alleles' when distinguishing them from sister chromatids.
meiosis — Nuclear division that results in the production of four daughter cells with half the chromosome number of the parent cell and with reshuffled alleles; in animals and plants it results in the formation of gametes.
Meiosis involves two rounds of division (Meiosis I and Meiosis II) and is essential for sexual reproduction, reducing the chromosome number by half and introducing genetic variation through crossing over and independent assortment. It's like a genetic lottery, shuffling and halving material.
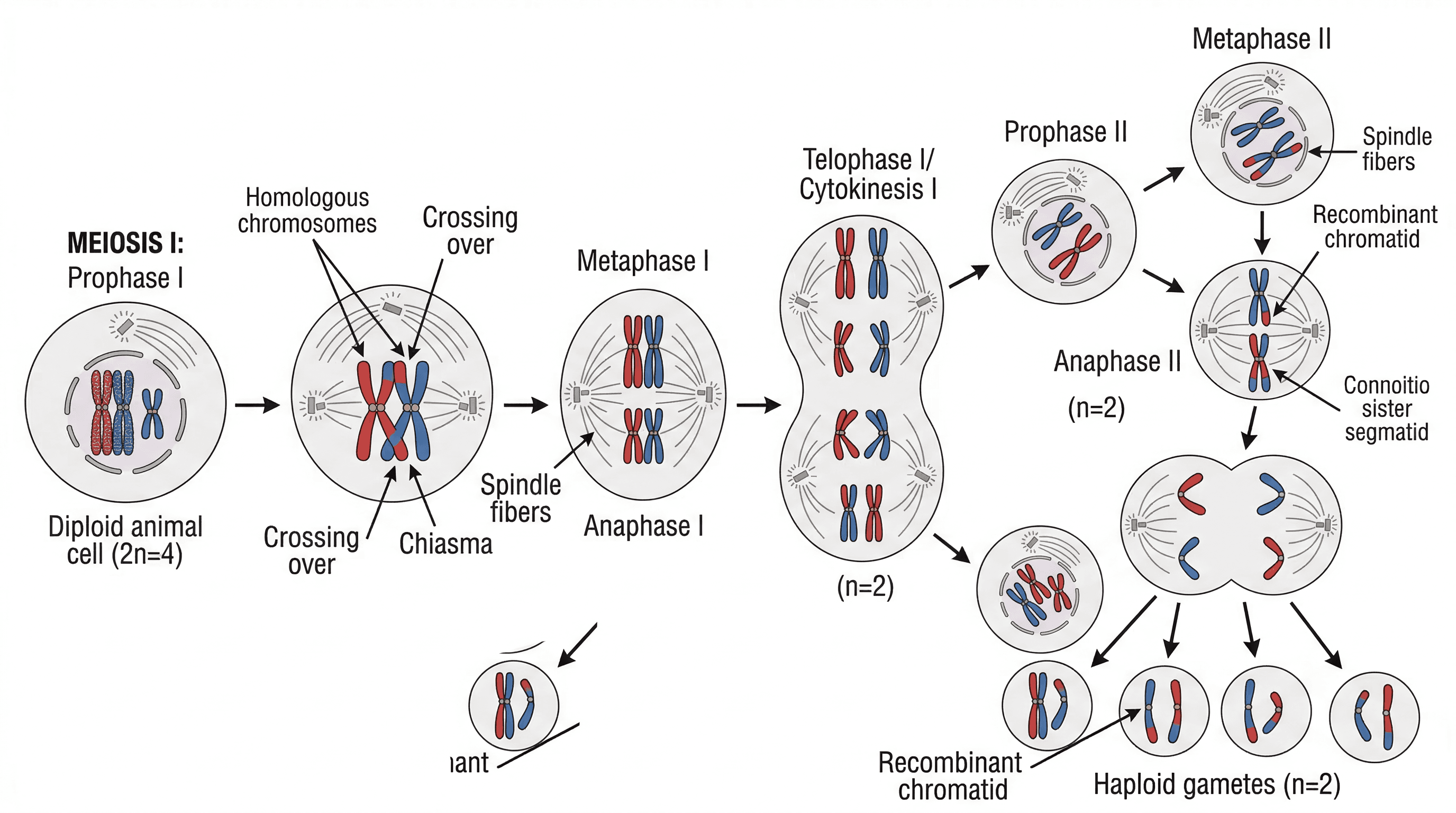
When describing meiosis, ensure you mention both the reduction in chromosome number and the generation of genetic variation, detailing the mechanisms (crossing over, independent assortment).
reduction division — Nuclear division that results in a reduction in chromosome number; the first division of meiosis is a reduction division.
Meiosis I is termed a reduction division because it separates homologous chromosomes, halving the chromosome number from diploid (2n) to haploid (n) in the daughter cells. This is crucial for maintaining a constant chromosome number across generations, like dividing a full set of tools into two half-sets.
Meiosis is fundamental to sexual reproduction, not only by producing haploid gametes but also by generating significant genetic variation among offspring. This variation is crucial for adaptation and evolution. Three primary mechanisms contribute to this genetic diversity: crossing over, independent assortment, and random fertilisation.
bivalent — Two homologous chromosomes lying alongside each other during meiosis I.
During prophase I of meiosis, homologous chromosomes pair up to form a bivalent, allowing for crossing over to occur between non-sister chromatids. This pairing is crucial for accurate segregation, like two dance partners holding hands.
chiasma (plural: chiasmata) — A position at which non-sister chromatids of homologous chromosomes cross over each other.
Chiasmata are visible manifestations of crossing over, where genetic material is exchanged between homologous chromosomes. They help hold homologous chromosomes together until anaphase I, like a knot tied between two ropes.
crossing over — The exchange of alleles between non-sister chromatids of homologous chromosomes during meiosis I.
This process shuffles alleles between homologous chromosomes, creating new combinations of alleles on the chromatids. It is a major source of genetic variation in gametes, like swapping pages between two identical books to create new versions.
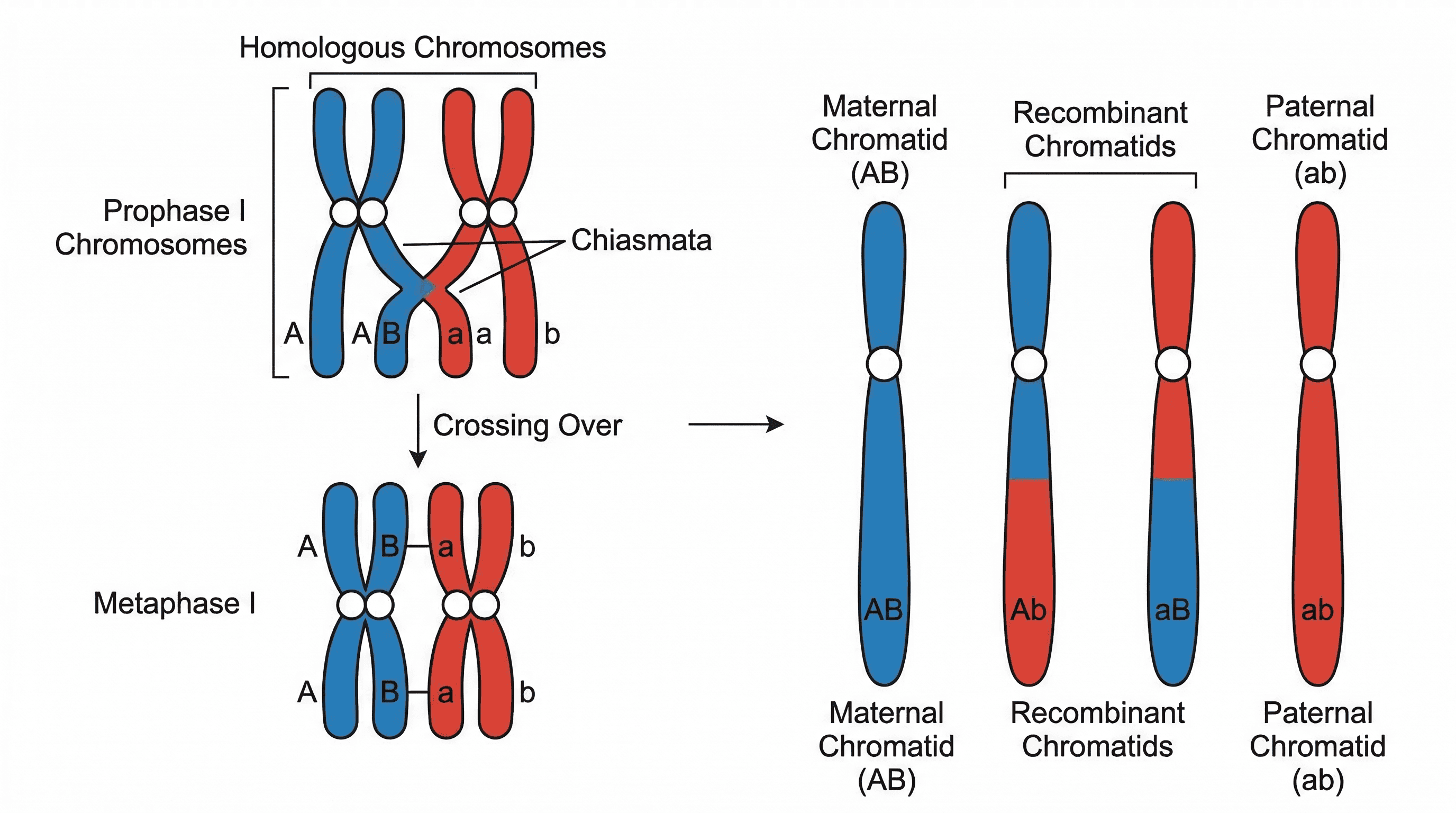
When explaining crossing over, specify 'non-sister chromatids' and 'homologous chromosomes' and link it to the production of 'genetic variation'.
independent assortment — The production of different combinations of alleles in daughter cells, as a result of the random alignment of bivalents on the equator of the spindle during metaphase I of meiosis.
The orientation of each homologous pair (bivalent) at the metaphase I plate is random and independent of other pairs. This leads to a vast number of possible combinations of chromosomes in the resulting gametes, contributing significantly to genetic variation, like randomly lining up different colored pairs of socks.
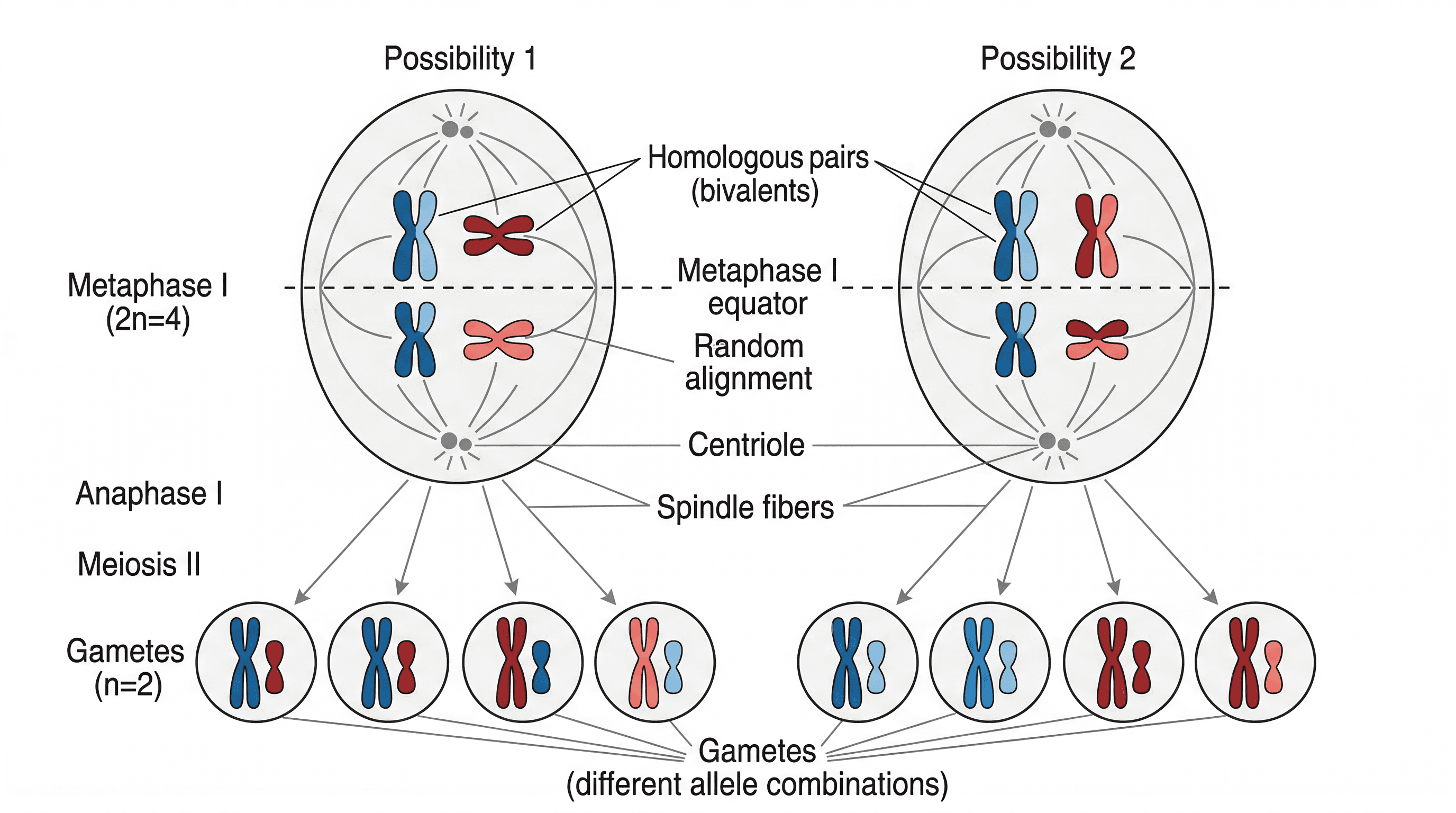
Students often confuse independent assortment with crossing over, but actually independent assortment refers to the random segregation of entire homologous chromosomes, while crossing over involves exchange of segments within chromosomes.
Link independent assortment directly to the 'random alignment of bivalents' in 'metaphase I' and its role in creating 'different combinations of alleles' in gametes.
Beyond meiosis, the random nature of fertilisation further enhances genetic variation. Any one of the genetically unique sperm cells can fertilise any one of the genetically unique egg cells. This random fusion of gametes ensures that each zygote formed is a unique combination of alleles from both parents, contributing to the diversity observed within a population.
locus (plural: loci) — The position of a gene on a chromosome.
Each gene occupies a specific locus on a particular chromosome. Homologous chromosomes have genes for the same traits at corresponding loci, though the alleles may differ, much like a specific address on a street.
allele — A variety of a gene.
Alleles are different forms of the same gene, arising from mutations, and they code for slightly different versions of a protein, leading to variations in a trait. For example, a gene for eye colour can have alleles for red or brown eyes, like different versions of a cake recipe.
genotype — The alleles possessed by an organism.
The genotype represents the genetic makeup of an individual for a particular trait, expressed using symbols (e.g., BB, Bb, bb). It determines the potential range of phenotypes, similar to the instruction code in a computer program.
phenotype — The observable features of an organism; it is affected by genes and also by environment.
The phenotype is the physical expression of an organism's genotype, influenced by both genetic factors and environmental conditions. Examples include coat colour, height, or blood group, much like the actual house built from a blueprint.
Students often confuse genotype with phenotype, but actually genotype is the genetic code, while phenotype is the observable characteristic.
When defining phenotype, always include that it is 'observable' and influenced by 'genes and environment' for a complete answer.
homozygous — Having two identical alleles of a gene.
An individual is homozygous for a gene if they have two copies of the same allele (e.g., BB or bb). This means they will express the trait associated with that allele, whether dominant or recessive, like having two scoops of the same ice cream flavor.
heterozygous — Having two different alleles of a gene.
An individual is heterozygous for a gene if they have two different alleles (e.g., Bb). In cases of complete dominance, the dominant allele's phenotype will be expressed, while the recessive allele is carried but not expressed, like having two scoops of different ice cream flavors.
dominant — A dominant allele has the same effect on phenotype, whether or not another allele is present.
A dominant allele expresses its trait even when only one copy is present in a heterozygous individual. It masks the effect of a recessive allele, like a loud voice in a conversation that is always heard.
Students often think that dominant alleles are always more common in a population, but dominance refers to the expression pattern of an allele, not its frequency.
recessive — A recessive allele only affects phenotype if no dominant allele is present.
A recessive allele only expresses its trait when two copies are present (homozygous recessive genotype). Its effect is masked by a dominant allele in a heterozygous individual, like a quiet voice that can only be heard if no loud voice is speaking.
multiple alleles — The existence of three or more alleles of a gene, as, for example, in the determination of A,B,O blood groups.
While an individual can only have two alleles for a given gene, a population can have multiple alleles for that gene. This increases the genetic diversity within the population, like having more than three different symbols that can appear in a slot machine.
When discussing multiple alleles, use the human ABO blood group system as a standard example, clearly showing the three alleles (IA, IB, IO).
codominant — Codominant alleles each affect phenotype when both of them are present.
In codominance, both alleles are fully expressed in the heterozygous individual, resulting in a phenotype that shows characteristics of both alleles, not an intermediate blend. The ABO blood group system (AB blood type) is a classic example, like two different colored paints both distinctly visible when mixed.
Students often confuse codominance with incomplete dominance; in codominance, both alleles are distinctly expressed, while in incomplete dominance, there is an intermediate phenotype.
When representing codominant alleles, use a capital letter for the gene and superscripts for the alleles (e.g., IA, IB) to clearly distinguish them from dominant/recessive alleles.
monohybrid inheritance — Inheritance of one gene.
Monohybrid crosses involve tracking the inheritance pattern of a single gene with two alleles. These crosses typically result in predictable phenotypic ratios (e.g., 3:1 in F2 generation for dominant/recessive traits), like focusing on just one specific feature of a car.
genetic diagram — A standard format in which the results of a genetic cross are predicted and explained.
Genetic diagrams, often incorporating Punnett squares, systematically illustrate the genotypes and phenotypes of parents, their gametes, and their potential offspring, along with expected ratios. They are essential tools for solving genetics problems, like a family tree that predicts traits.
Always include all headings (parental phenotypes, genotypes, gametes, offspring genotypes, phenotypes, and ratios) in a genetic diagram for full marks.
Punnett square — Part of a genetic diagram in which the genotypes of the offspring are worked out from the genotypes of the gametes.
The Punnett square is a grid used to combine the possible gametes from each parent to predict the genotypes and their frequencies in the offspring. It visually represents the random fusion of gametes, like a multiplication table for genetics.
Students often think a Punnett square is the entire genetic diagram, but actually it is only a component used to determine offspring genotypes.
F1 generation — The offspring resulting from the cross between individuals with a homozygous recessive and a homozygous dominant genotype.
The F1 generation (first filial generation) typically consists of all heterozygous individuals when the parental cross involves two pure-breeding (homozygous) parents with contrasting traits. These individuals are then often interbred to produce the F2 generation, like the puppies from a purebred black and white dog cross.
F2 generation — The offspring resulting from a cross between two F1 individuals.
The F2 generation (second filial generation) is produced by interbreeding individuals from the F1 generation. This generation typically exhibits the classic Mendelian ratios (e.g., 3:1 for monohybrid, 9:3:3:1 for dihybrid) due to the segregation and independent assortment of alleles, like the puppies produced when two F1 puppies breed.
test cross — A genetic cross in which an organism showing the dominant characteristic is crossed with a homozygous recessive organism; the phenotypes of the offspring can indicate whether the original organism is homozygous or heterozygous.
A test cross is used to determine the unknown genotype of an individual expressing a dominant phenotype. If any recessive offspring are produced, the unknown parent must be heterozygous; if all offspring show the dominant phenotype, the unknown parent is likely homozygous dominant, like a detective trying to figure out a secret ingredient.
sex chromosomes — The chromosomes that determine sex; in humans, these are the X and Y chromosomes.
Sex chromosomes carry genes that determine an individual's biological sex and also contain other genes unrelated to sex determination (sex-linked genes). In humans, XX results in female, and XY results in male, like the 'gender' setting on a device.
sex-linked gene — A gene found on a region of a sex chromosome that is not present on the other sex chromosome; in humans, most sex-linked genes are found on the X chromosome.
Sex-linked genes exhibit unique inheritance patterns because males only have one X chromosome, meaning they express any allele on their X chromosome, whether dominant or recessive. Females, with two X chromosomes, can be carriers for recessive sex-linked traits.
For sex-linked traits, always use X and Y notation with the allele as a superscript (e.g., X^H, X^h). Remember males only have one X chromosome.
carrier — An individual that possesses a particular allele as a single copy whose effect is masked by a dominant allele, so that the associated characteristic (such as a hereditary disease) is not displayed but may be passed to offspring.
Carriers are typically heterozygous for a recessive trait or disease. They do not express the phenotype themselves but can pass the recessive allele to their offspring, potentially leading to affected individuals in future generations.
dihybrid inheritance — The inheritance of two genes.
Dihybrid crosses involve tracking the inheritance patterns of two different genes simultaneously. When these genes are on different chromosomes, they assort independently, leading to characteristic phenotypic ratios like 9:3:3:1 in the F2 generation.
epistasis — The interaction of two genes at different loci; one gene may affect the expression of the other.
Epistasis occurs when the allele of one gene masks or modifies the phenotypic expression of alleles at a different gene locus. This interaction can lead to modified dihybrid ratios that deviate from the expected 9:3:3:1, as the genes do not act independently in determining the final phenotype.
autosomal linkage — The presence of two genes on the same autosome, (any chromosome other than a sex chromosome) so that they tend to be inherited together and do not assort independently.
When genes are located on the same chromosome, they are said to be linked. These genes tend to be inherited together because the chromosome is passed as a unit during meiosis. This means they do not assort independently, leading to different phenotypic ratios than expected for unlinked genes.
Students often assume independent assortment for all dihybrid crosses, but genes located on the same chromosome (linked genes) do not assort independently unless crossing over occurs.
While linked genes tend to be inherited together, crossing over can separate them. If crossing over occurs between two linked genes on homologous chromosomes, it can lead to the formation of recombinant gametes. The frequency of these recombinant gametes is proportional to the distance between the linked genes on the chromosome.
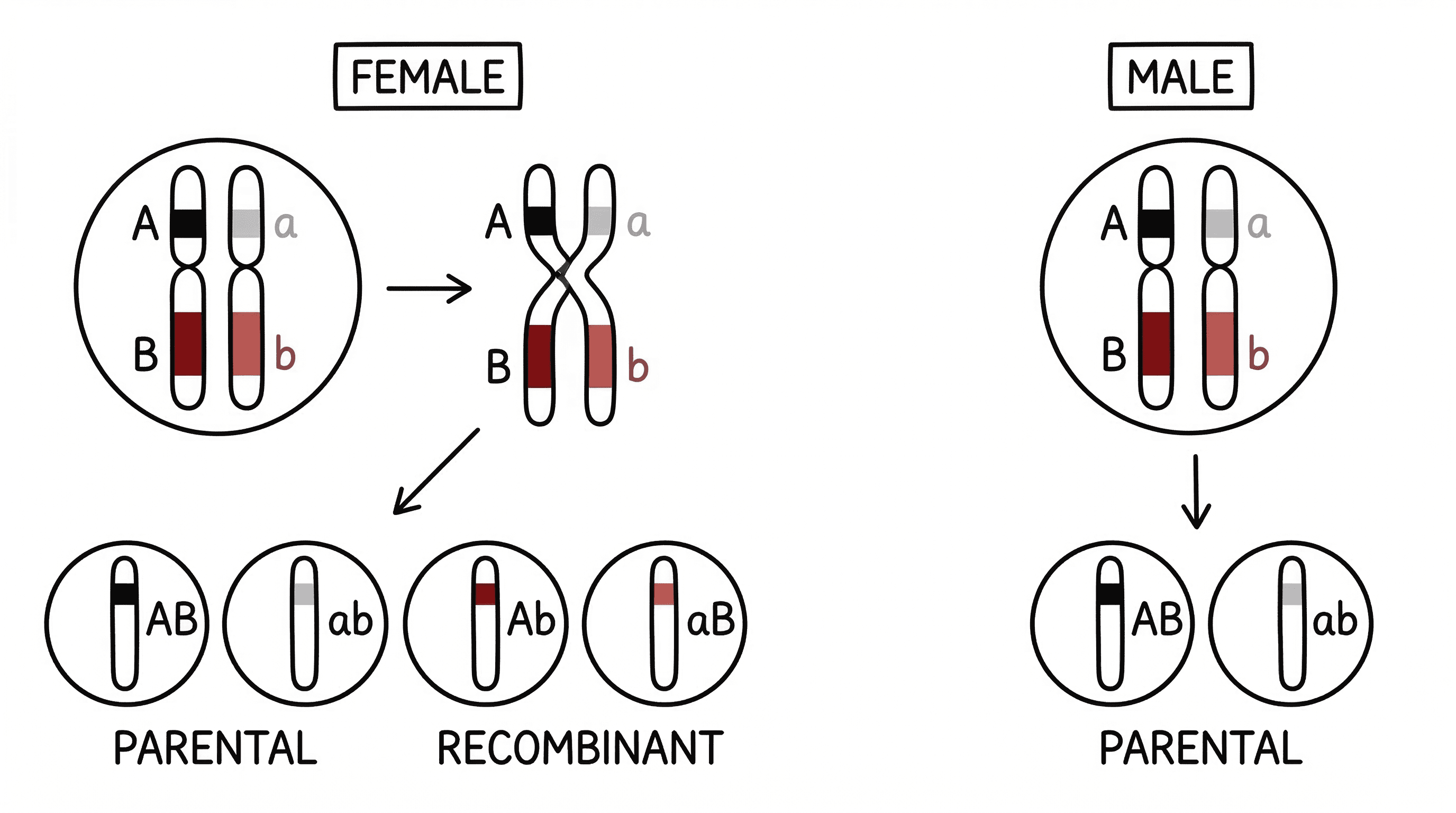
parental type — Offspring that show the same combinations of characteristics as their parents.
Parental types are offspring whose phenotypes match one of the parental phenotypes. In linkage studies, a higher proportion of parental types compared to recombinant types indicates that the genes are linked and crossing over is less frequent.
recombinant — Offspring that show different combinations of characteristics from their parents.
Recombinant offspring display new combinations of traits not seen in either parent. These arise from genetic recombination events, such as crossing over between linked genes or independent assortment of unlinked genes.
chi-squared (χ2) test — A statistical test that is used to determine whether differences between observed and expected results are significant.
The chi-squared test helps evaluate if observed phenotypic ratios in genetic crosses significantly deviate from expected Mendelian ratios, or if the differences are merely due to chance. It is a crucial tool for validating genetic hypotheses.
Chi-squared test
Used to determine if differences between observed and expected results are statistically significant. Compare calculated χ² value to critical value from a table based on degrees of freedom (number of classes - 1) and a chosen probability (e.g., 0.05).
When using the chi-squared test, always state your null hypothesis, show your calculation of the χ² value, determine degrees of freedom (n-1), and compare your value to the critical value to make a conclusion.
The relationship between genes, proteins, and phenotype is fundamental to understanding inheritance. Genes contain the instructions for making proteins, and these proteins then carry out various functions within the cell, ultimately determining an organism's observable characteristics. Mutations in genes can alter protein structure or function, leading to changes in phenotype, often resulting in genetic conditions.
Several human genetic conditions illustrate the gene-protein-phenotype link. The TYR gene codes for tyrosinase, an enzyme crucial for melanin production; mutations lead to albinism. The HBB gene codes for a subunit of haemoglobin; mutations cause sickle cell anaemia. The F8 gene codes for factor VIII, a clotting protein; mutations result in haemophilia. The HTT gene codes for huntingtin protein; mutations cause Huntington’s disease. In plants, the Le gene controls gibberellin production, influencing stem elongation.
Gene expression in prokaryotes is often controlled by operons, which are clusters of genes regulated by a single promoter. The lac operon in bacteria, for example, controls the production of enzymes needed for lactose metabolism. It is an inducible system, meaning the enzymes are only synthesised when lactose is present, ensuring efficient resource use.
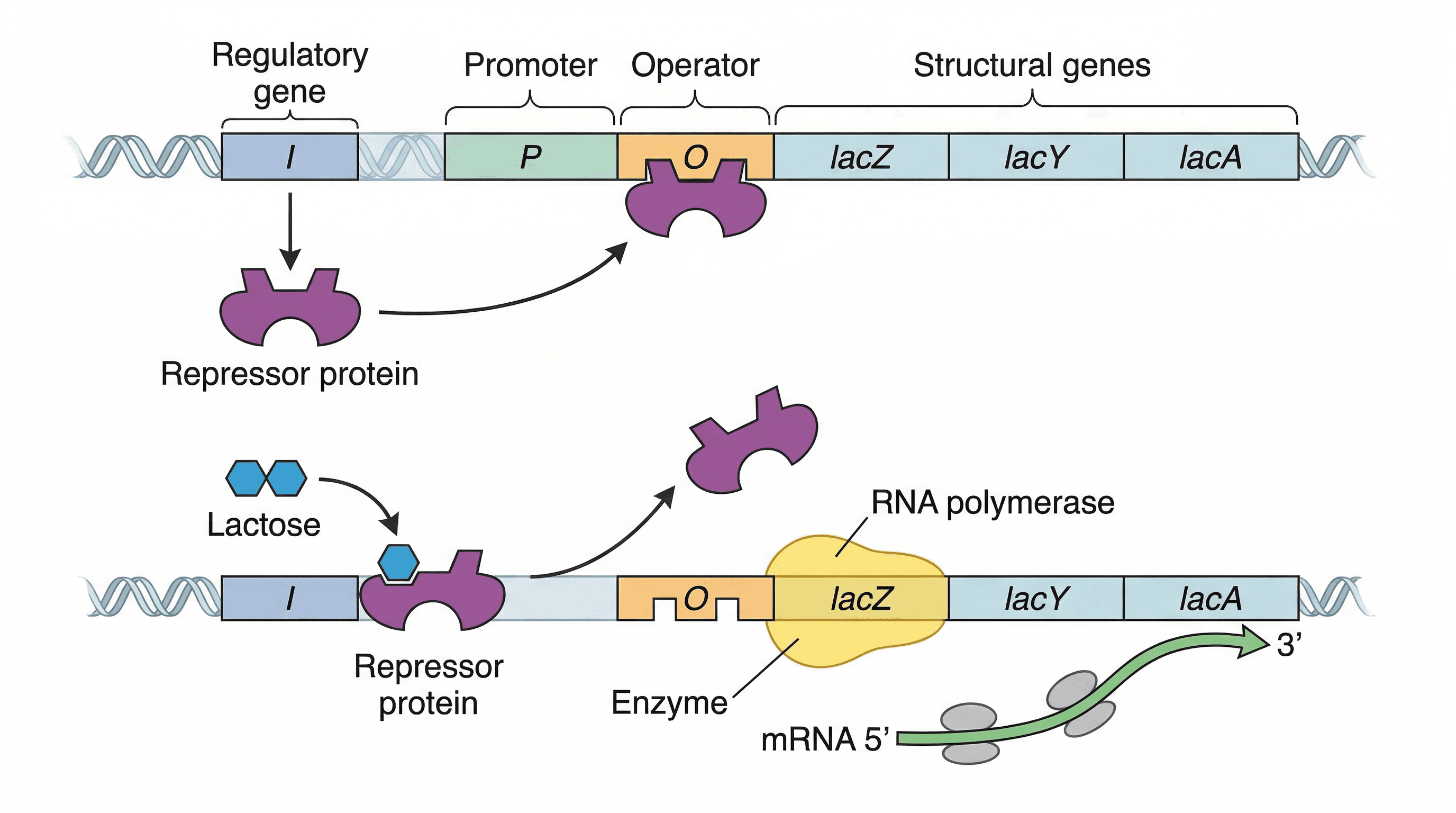
operon — A functional unit of transcription; a cluster of genes that are controlled by the same promoter.
Operons allow prokaryotes to efficiently regulate the expression of genes involved in a common metabolic pathway. All genes within an operon are transcribed together as a single mRNA molecule, ensuring coordinated protein production.
lac operon — An operon (see above) found in some bacteria that controls the production of β-galactosidase and two other structural proteins.
The lac operon is a classic example of an inducible operon. It contains structural genes for enzymes like β-galactosidase, which breaks down lactose, and is regulated by a repressor protein that binds to the operator in the absence of lactose.
β-galactosidase — An enzyme that catalyses the hydrolysis of lactose to glucose and galactose.
This enzyme is crucial for bacteria to utilise lactose as an energy source. Its production is regulated by the lac operon, ensuring it is only synthesised when lactose is available in the environment.
structural gene — A gene that codes for a protein that has a function within a cell.
Structural genes are the core components of operons, coding for the enzymes or proteins directly involved in a metabolic pathway or cellular process. Their expression is controlled by regulatory elements.
regulatory gene — A gene that codes for a protein that helps to control the expression of other genes.
Regulatory genes produce proteins, such as repressors or activators, that bind to specific DNA sequences to either inhibit or promote the transcription of structural genes. They are key to controlling gene expression.
inducible enzyme — An enzyme that is synthesised only when its substrate is present.
Inducible enzymes are part of metabolic pathways that are only required under specific environmental conditions. Their synthesis is 'induced' by the presence of their substrate, preventing wasteful production when not needed, as seen with β-galactosidase and lactose.
repressible enzyme — An enzyme that is normally produced, and whose synthesis is prevented by the presence of an effector.
Repressible enzymes are typically involved in anabolic pathways, where their product is continuously needed. Their synthesis is 'repressed' when the end-product accumulates, preventing overproduction and conserving energy.
Students often think operons are found in eukaryotes, but they are characteristic of prokaryotic gene regulation.
In eukaryotes, gene expression is controlled at multiple levels, with transcription factors playing a crucial role. These proteins bind to specific DNA sequences (promoters or enhancers) to either promote or inhibit the transcription of genes. This complex regulation allows for precise control over which genes are expressed, when, and in which cells.
transcription factor — A molecule that affects whether or not a gene is transcribed.
Transcription factors are proteins that regulate gene expression by binding to DNA and either facilitating or blocking the binding of RNA polymerase, thereby controlling the rate of transcription. They are essential for cell differentiation and development.
An example of eukaryotic gene control involves gibberellin hormones and DELLA proteins in plants. Gibberellins promote stem elongation by causing the degradation of DELLA proteins, which are repressors of growth. When gibberellin is present, DELLA proteins are removed, allowing genes for stem elongation to be expressed. The alleles Le and le control gibberellin production, thus influencing stem height.
When explaining genetic variation, always refer to the three key sources: crossing over, independent assortment, and random fertilisation.
Structure genetic diagrams precisely: 1. Parental Phenotype, 2. Parental Genotype, 3. Gametes, 4. Punnett Square, 5. Offspring Genotype(s), 6. Offspring Phenotype(s) & Ratio.
Recognise key phenotypic ratios: 3:1 (monohybrid), 9:3:3:1 (unlinked dihybrid), 1:2:1 (codominance). Deviations from 9:3:3:1 often indicate linkage or epistasis.
Definitions Bank
sexual reproduction
Reproduction involving the fusion of gametes (fertilisation) to produce a zygote.
gamete
A sex cell; during sexual reproduction, two gametes fuse together to form a zygote; gametes are usually haploid.
fertilisation
The fusing of the nuclei of two gametes, to form a zygote.
zygote
A cell formed by the fusion of the nuclei of two gametes; most zygotes are diploid.
diploid
Containing two complete sets of chromosomes; can be signified by the symbol 2n.
+41 more definitions
View all →Command Word Guide
| Describe | For meiosis, describe the key events in Meiosis I (homologous chromosome pairing, crossing over, reduction division) and Meiosis II (sister chromatid separation). For sexual reproduction, mention gamete fusion and zygote formation. |
| Explain | For genetic variation, explain *how* crossing over, independent assortment, and random fertilisation lead to new allele combinations. For gene-phenotype relationships, explain the role of the gene in coding for a protein and how this protein affects the observable characteristic. For gene control, explain the mechanism (e.g., lac operon components and their interactions, transcription factor binding). |
| Construct | For genetic diagrams, ensure all headings (parental phenotypes, genotypes, gametes, offspring genotypes, phenotypes, and ratios) are present and correct. Use appropriate notation for alleles (capital/lowercase, superscripts for codominance/sex-linkage). |
| Interpret | For genetic diagrams, correctly identify offspring genotypes and phenotypes and state their ratios. For chi-squared test results, interpret the calculated value against the critical value to determine statistical significance and draw a conclusion about the null hypothesis. |
+1 more
View all →Common Mistakes
Confusing homologous chromosomes with sister chromatids.
Homologous chromosomes are a pair (one from each parent) carrying the same genes, while sister chromatids are identical copies of a single chromosome joined at the centromere.
Thinking that dominant alleles are always more common in a population.
Dominance refers to the expression pattern of an allele (it masks a recessive allele), not its frequency in a population.
Mistaking codominance for incomplete dominance.
In codominance, both alleles are distinctly expressed (e.g., AB blood type), while in incomplete dominance, there is a blended, intermediate phenotype.
+23 more
View all →This chapter explores how variation within species drives evolution through natural and artificial selection. It details the mechanisms of genetic change, including genetic drift and the Hardy-Weinberg principle, and explains how these processes lead to the formation of new species.
genetic variation — Differences between the DNA base sequences of individuals within a species.
This variation is the raw material for natural selection and can arise from independent assortment, crossing over, random gamete fusion, random mating, and mutation. It leads to phenotypic variation, much like having many different cards and shuffling them in countless ways to create unique hands.
phenotypic variation — Differences between the observable characteristics of individuals within a species.
This variation is a result of the interaction between genetic and environmental factors and is what natural selection acts upon. For example, siblings share genes but might have different heights or weights due to their diet or exercise, showing phenotypic variation.
Students often confuse genetic variation with environmentally induced variation. Remember that only genetic variation is heritable and can be acted upon by natural selection; environmentally-induced variation is not passed on to offspring.
discontinuous variation — Differences between individuals of a species in which each one belongs to one of a small number of distinct categories, with no intermediates.
This type of variation is typically caused by different alleles at a single gene locus having large effects on the phenotype, with the environment having no effect. ABO blood groups are an example, much like choosing a shirt from a limited set of colours with no in-between shades.
continuous variation — Differences between individuals of a species in which each one can lie at any point in the range between the highest and lowest values.
This variation is affected by multiple genes (polygenes), each with small, often additive effects, and is also influenced by environmental factors. Height and mass are examples, similar to a dimmer switch for a light where you can set it to any brightness level.
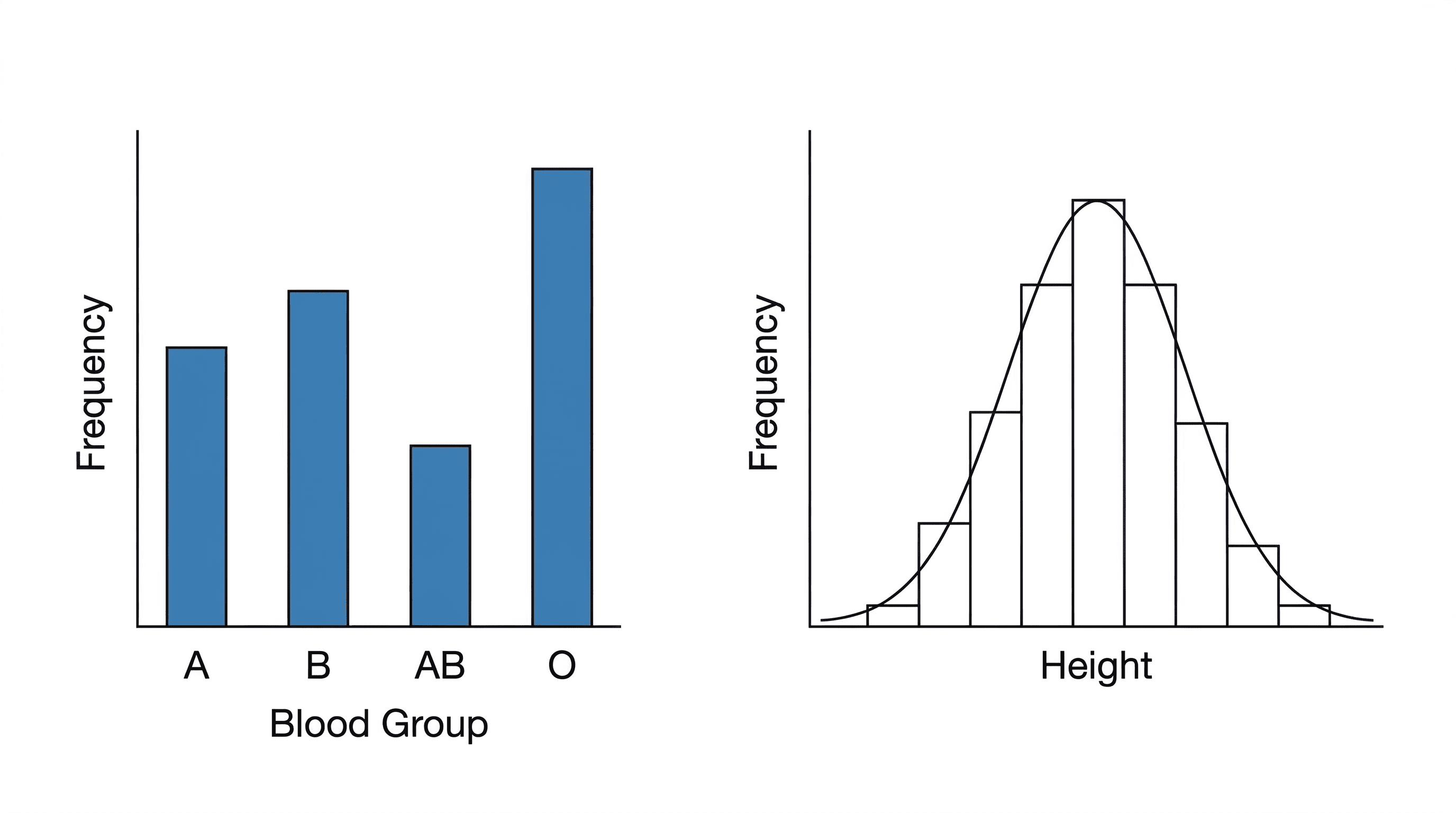
polygenes — A number of different genes at different loci that all contribute to a particular aspect of phenotype.
These genes typically have small, often additive effects, leading to continuous variation in traits like height or mass. The combined effect of many such genes creates a wide range of possible phenotypes, much like many different ingredients each adding a little bit to the overall flavour.
For continuous variation, remember to mention both polygenic inheritance (many genes, small additive effects) and environmental influence, and that it results in a range of phenotypes.
environmental factor — A feature of the environment of an organism that affects its survival.
These factors can be biotic (living organisms) or abiotic (non-living components) and exert selection pressures that influence which individuals survive and reproduce. They contribute to phenotypic variation but are not heritable, like the amount of sunlight affecting a plant's final size.
biotic factor — An environmental factor that is caused by living organisms (e.g. predation, competition).
These factors are interactions between organisms that can act as selection pressures, influencing survival and reproduction rates within a population. For example, predators selecting against certain prey phenotypes, similar to deer eating certain plants in a garden.
competition — The need for a resource by two organisms, when that resource is in short supply.
Competition can be intraspecific (within a species) or interspecific (between species) and acts as a selection pressure, favouring individuals better able to acquire the limited resource. This is like two children wanting the last slice of cake.
abiotic factor — An environmental factor that is caused by non-living components (e.g. soil pH, light intensity).
These physical and chemical conditions of an environment act as selection pressures, favouring individuals with adaptations that allow them to survive and thrive under those specific conditions. The temperature and rainfall in a desert are examples.
fitness — The ability of an organism to survive and reproduce.
Fitness is a measure of an organism's reproductive success, specifically its capacity to survive and transmit its alleles to its offspring. Higher fitness means a greater contribution to the gene pool of the next generation, much like having the energy to run a race again and produce more runners.
Students often equate fitness solely with physical strength or survival, but it specifically refers to reproductive success and the ability to pass on alleles.
selection pressure — An environmental factor that affects the chance of survival of an organism; organisms with one phenotype are more likely to survive and reproduce than those with a different phenotype.
Selection pressures drive natural selection by differentially favouring certain alleles or phenotypes, leading to changes in allele frequencies over generations. They can be biotic or abiotic, like a strong wind favouring trees with stronger roots.
natural selection — The process by which individuals with a particular set of alleles are more likely to survive and reproduce than those with other alleles; over time and many generations, the advantageous alleles become more frequent in the population.
This is a key mechanism of evolution, where environmental selection pressures act on phenotypic variation, leading to a gradual increase in the frequency of advantageous alleles and a decrease in disadvantageous ones. It acts like a filter, letting through only the best-adapted individuals.
Students sometimes think natural selection is a conscious process or that organisms 'try' to adapt. Remember that it's a passive process where the environment selects existing advantageous traits.
Ensure your explanation of natural selection includes variation, selection pressure, differential survival/reproduction, and changes in allele frequency over generations.
stabilising selection — Natural selection that tends to keep allele frequencies relatively constant over many generations.
This occurs when environmental conditions are stable, favouring individuals with intermediate phenotypes and selecting against extreme variations. It reduces the range of variation in a population, much like quality control rejecting screws that are too long or too short.
directional selection — Natural selection that causes a gradual change in allele frequency over many generations.
This occurs when there is a change in environmental conditions or a new allele arises, favouring individuals at one extreme of the phenotypic range. It shifts the mean phenotype of the population in a particular direction, like adjusting a machine to produce progressively longer screws.
disruptive selection — Natural selection that maintains relatively high frequencies of two different sets of alleles; individuals with intermediate features and allele sets are not selected for.
This occurs when conditions favour individuals at both extremes of the phenotypic range, while selecting against intermediate phenotypes. It can lead to polymorphism within a population and potentially speciation, similar to a fishing net with only very small or very large holes.
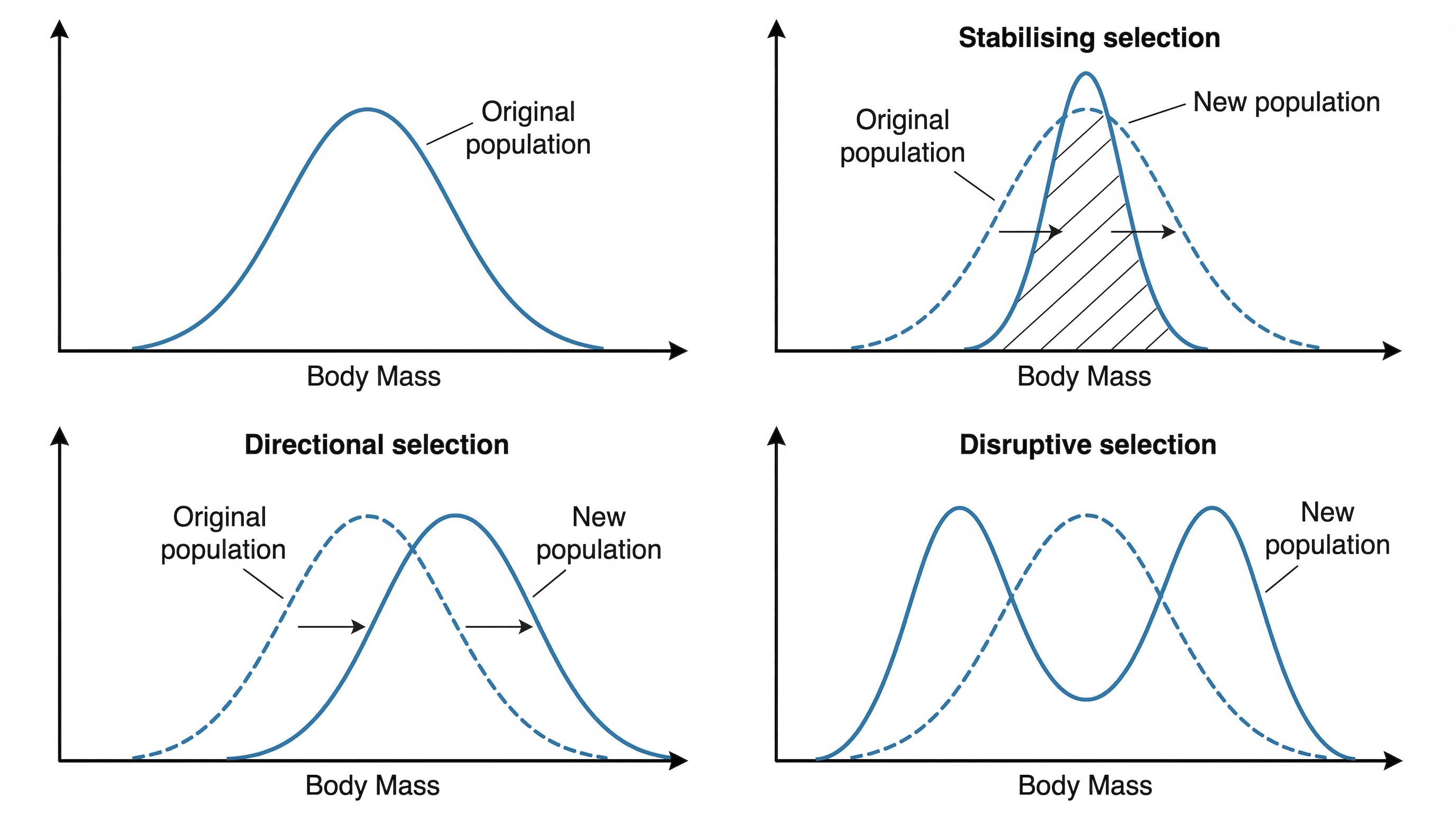
Directional selection drives changes in allele frequencies, leading to adaptations. A prominent example is the development of antibiotic resistance in bacteria. When antibiotics are used, they act as a selection pressure, killing susceptible bacteria. Any bacteria with pre-existing mutations that confer resistance survive and reproduce, passing on their advantageous alleles. Over generations, the frequency of resistant alleles increases, making the antibiotic less effective. Industrial melanism, exemplified by the peppered moth, is another case where environmental changes (soot pollution) favoured darker moths, shifting the population's phenotype.
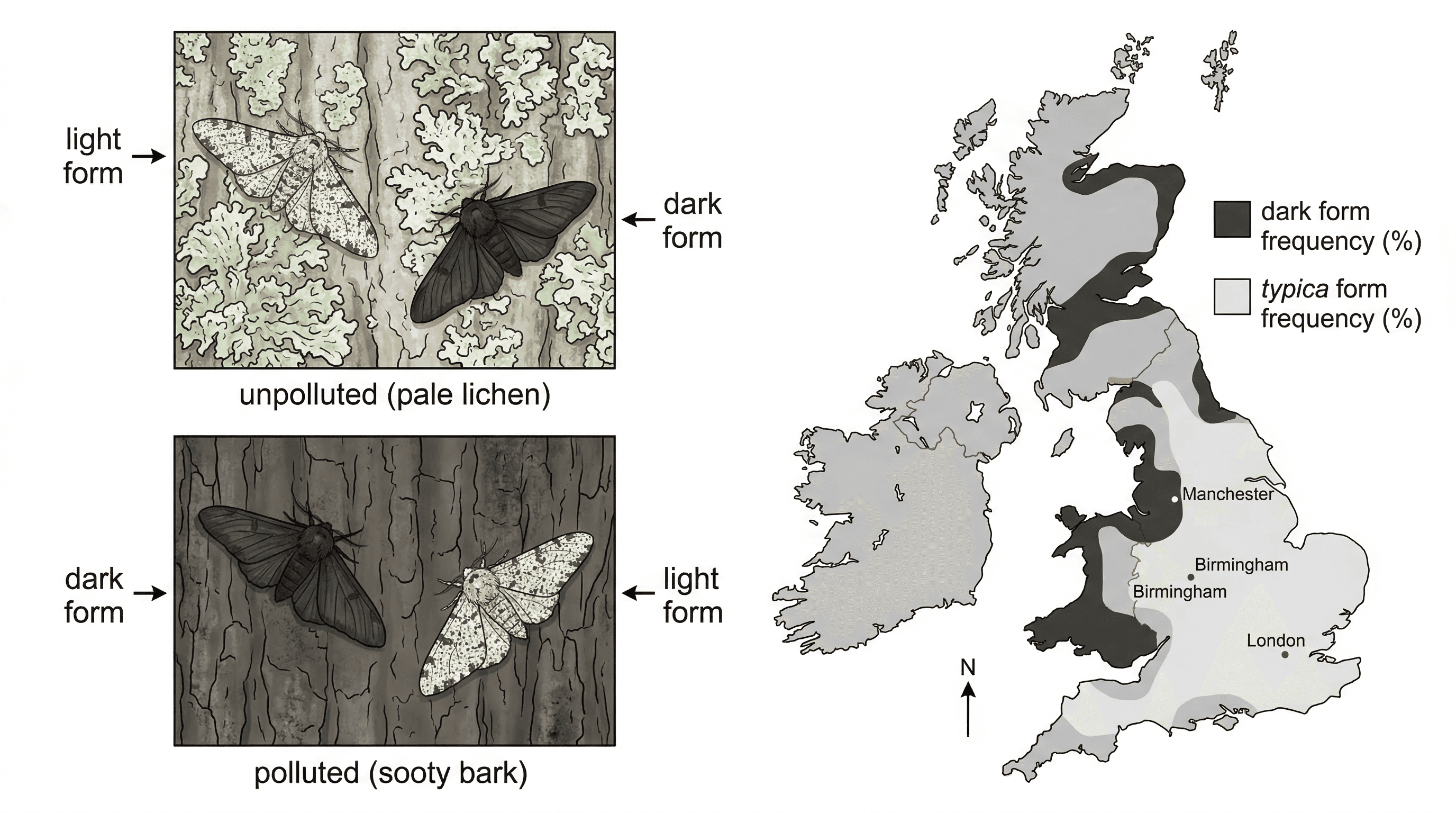
polymorphism — The continued existence of two or more different phenotypes in a species.
Polymorphism can be maintained by disruptive selection or other factors, ensuring genetic diversity within a population. It means that distinct forms of a trait coexist, like a species of flower that consistently produces both red and white blooms.
genetic drift — The gradual change in allele frequencies in a small population, where some alleles are lost or favoured just by chance and not by natural selection.
This random process has a more significant effect in small populations because chance events (e.g., which individuals happen to reproduce) can lead to disproportionate changes in allele frequencies, potentially leading to the loss of alleles. It's like randomly picking marbles from a small bag, where the proportions might change by chance.
Students may confuse genetic drift with natural selection. Remember that genetic drift is purely random, while natural selection is driven by environmental pressures.
gene pool — The complete range of DNA base sequences in all the organisms in a species or population.
It represents the total genetic diversity available within a population. Changes in allele frequencies within the gene pool are the basis of evolution, much like a library containing all the books and different editions available to a community.
founder effect — The reduction in a gene pool compared with the main populations of a species, resulting from only two or three individuals (with only a selection of the alleles in the gene pool) starting off a new population.
When a small group of individuals establishes a new population, the allele frequencies in this new population may differ significantly from the original population simply due to the limited genetic diversity of the founders. This is a form of genetic drift, similar to a new town's genetic makeup reflecting only its few founders.
evolutionary bottleneck — A period when the numbers of a species fall to a very low level, resulting in the loss of a large number of alleles and therefore a reduction in the gene pool of the species.
This event drastically reduces genetic diversity, making the surviving population more vulnerable to future environmental changes as it has fewer alleles to adapt with. It is a severe form of genetic drift, like pouring sand through a narrow bottleneck where only a fraction makes it through.
Emphasize that genetic drift is a 'chance' event and is most impactful in 'small populations', leading to random changes in allele frequencies. For the founder effect, highlight the 'small number of individuals' and the 'chance' selection of alleles they carry.
The Hardy-Weinberg principle provides a mathematical model to describe a population that is not evolving. It states that allele and genotype frequencies in a population will remain constant from generation to generation in the absence of other evolutionary influences. This principle serves as a null hypothesis against which to compare real populations to detect if evolution is occurring. The conditions for Hardy-Weinberg equilibrium include a large population size, random mating, no mutation, no gene flow (migration), and no natural selection.
Hardy–Weinberg Equation 1
Used to calculate allele frequencies in a population where only two alleles for a gene exist. 'p' is the frequency of the dominant allele, and 'q' is the frequency of the recessive allele.
Hardy–Weinberg Equation 2
Used to calculate genotype frequencies in a large, randomly mating population with no significant selective pressure, migration, or non-random mating. 'p^2' is the frequency of the homozygous dominant genotype, '2pq' is the frequency of the heterozygous genotype, and 'q^2' is the frequency of the homozygous recessive genotype.
For Hardy-Weinberg calculations, always start by identifying the frequency of the homozygous recessive phenotype, which is q². From q², you can calculate q, then p, and then all other frequencies.
artificial selection — The selection by humans of organisms with desired traits to survive and reproduce; also known as selective breeding.
Humans intentionally choose individuals with specific desirable characteristics to breed, aiming to enhance those traits in future generations. This process can lead to extreme phenotypes and reduced genetic diversity, much like a chef choosing specific ingredients to create a dish with a particular flavour.
Artificial selection, or selective breeding, involves humans acting as the selective agent to propagate desired traits. This has been used to improve milk yield in dairy cattle by breeding cows with high milk production. In crops, it's used to introduce disease resistance to varieties of wheat and rice. Inbreeding, breeding between close relatives, is often used to achieve homozygosity for desired traits, but it carries the risk of inbreeding depression. Outbreeding, breeding unrelated individuals, can lead to hybrid vigour, where offspring show increased ability to survive and grow well due to increased heterozygosity, as seen in maize hybridisation.
inbreeding — Breeding between organisms with similar genotypes, or that are closely related.
This practice increases the likelihood of offspring being homozygous for many genes, which can be used in selective breeding to fix desirable traits but also carries the risk of inbreeding depression. It's like a family only marrying within its own members, leading to a limited gene pool.
inbreeding depression — A loss of the ability to survive and grow well, due to breeding between close relatives; this increases the chance of harmful recessive alleles coming together in an individual and being expressed.
Repeated inbreeding increases homozygosity, which can expose deleterious recessive alleles that are normally masked in heterozygotes, leading to reduced vigour, fertility, and survival. This is similar to repeatedly copying a document from a copy, where errors accumulate.
Students might think inbreeding always produces weak offspring. Remember that it increases homozygosity which can fix desirable traits, though it also increases the risk of expressing harmful recessive alleles.
outbreeding — Breeding between individuals that are not closely related.
This practice increases heterozygosity in the offspring, which can lead to hybrid vigour and mask deleterious recessive alleles, improving overall fitness and yield. It's like bringing in new ingredients from different cuisines to create a more diverse and robust meal.
hybrid vigour — An increased ability to survive and grow well, as a result of outbreeding and therefore increased heterozygosity.
Also known as heterosis, it is the phenomenon where the offspring of genetically diverse parents show superior qualities (e.g., growth, yield, disease resistance) compared to either parent. This is often due to the masking of recessive deleterious alleles and the expression of advantageous dominant alleles, like mixing two different types of strong plants to get an even stronger one.
evolution — A process leading to the formation of new species from pre-existing species over time.
Evolution involves changes in the characteristics of species over time, primarily due to changes in allele frequencies within gene pools across many generations. Natural selection, genetic drift, and mutation are key mechanisms, much like a language slowly changing over centuries.
Students often think evolution is about individuals changing during their lifetime. Remember that it's about changes in populations over generations.
morphological — Relating to structural features.
Morphological features are observable physical characteristics of an organism, such as shape, size, and colour. They are often used in classification and to distinguish between species, like describing a car by its body shape and colour.
physiological — Relating to metabolic and other processes in a living organism.
Physiological features describe how an organism's body functions, including its metabolism, reproduction, and responses to the environment. These can also be used to differentiate species, similar to describing how a car's engine works.
reproductive isolation — The inability of two groups of organisms to breed with one another; two populations of the same species may be geographically separated, or two different species are unable to breed to produce fertile offspring.
This is a critical step in speciation, preventing gene flow between populations and allowing them to diverge genetically. It can arise from various pre-zygotic or post-zygotic barriers, like two groups of people who speak different languages and rarely intermarry.
genetically isolated — No longer able to breed together; there is no exchange of genes.
This is the ultimate outcome of reproductive isolation, where two populations or species cannot interbreed, meaning their gene pools are completely separate and they evolve independently. It's like two separate computer networks that cannot share files.
speciation — The production of new species.
Speciation is the evolutionary process by which new biological species arise. It occurs when populations become reproductively isolated, preventing gene flow and allowing them to diverge genetically over time.
geographical isolation — Separation by a geographical barrier, such as a stretch of water or a mountain range.
This physical barrier prevents gene flow between populations, leading to allopatric speciation. Over time, the separated populations experience different selection pressures and genetic drift, leading to genetic divergence.
allopatric speciation — The development of new species following geographical isolation.
In allopatric speciation, a physical barrier prevents gene flow between populations. Over time, these isolated populations accumulate genetic differences due to different selection pressures, mutations, and genetic drift, eventually leading to reproductive isolation and the formation of distinct species.
sympatric speciation — The development of new species without any geographical separation.
Sympatric speciation occurs when populations diverge into new species while inhabiting the same geographical area. This can happen due to ecological separation (e.g., different food sources) or behavioural separation (e.g., different mating rituals) that lead to reproductive isolation.
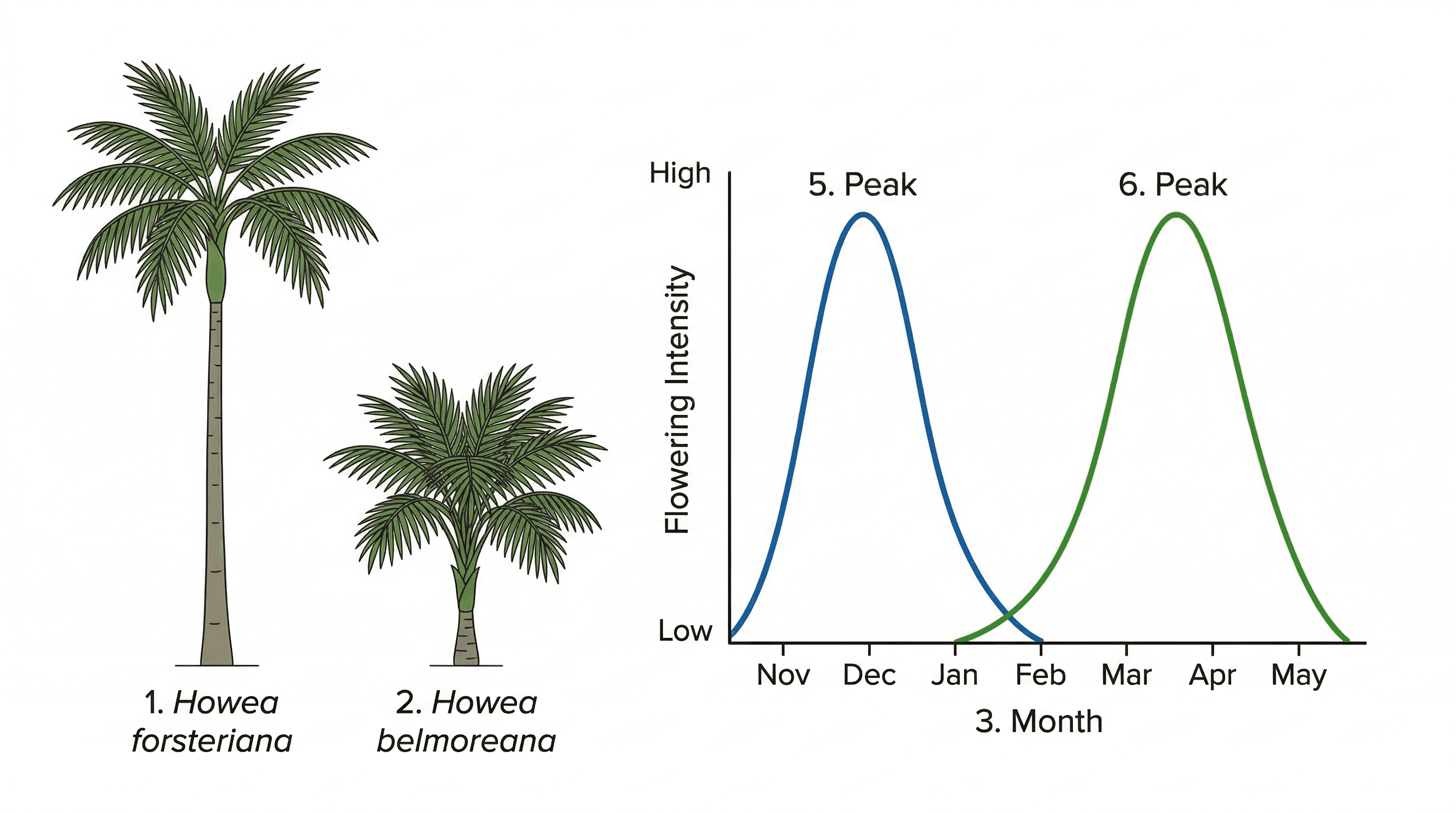
ecological separation — The separation of two populations because they live in different environments in the same area and so cannot breed together.
This is a mechanism for sympatric speciation, where populations within the same geographical region exploit different niches or habitats, reducing their chances of interbreeding and leading to genetic divergence.
behavioural separation — The separation of two populations because they have different behaviours which prevent them breeding together.
This is another mechanism for sympatric speciation, where differences in mating rituals, courtship displays, or activity times prevent interbreeding between populations, even if they are in the same area.
Students sometimes struggle to understand how sympatric speciation can occur without geographical isolation. Remember to consider ecological or behavioural separation as isolating mechanisms.
DNA sequencing provides a powerful tool for determining evolutionary relationships between species. By comparing the DNA base sequences of different organisms, scientists can identify similarities and differences. The more similar the DNA sequences, the more closely related the species are considered to be, indicating a more recent common ancestor. This molecular evidence complements morphological, physiological, and behavioural comparisons, offering a precise way to trace evolutionary lineages and construct phylogenetic trees.
In speciation questions, always state that 'reproductive isolation' is the key step that prevents gene flow between populations, leading to the evolution of separate species.
When comparing natural and artificial selection, clearly state the difference in the selection pressure (environment vs. humans) and the goal (survival vs. human benefit).
Definitions Bank
genetic variation
Differences between the DNA base sequences of individuals within a species.
phenotypic variation
Differences between the observable characteristics of individuals within a species.
discontinuous variation
Differences between individuals of a species in which each one belongs to one of a small number of distinct categories, with no intermediates.
continuous variation
Differences between individuals of a species in which each one can lie at any point in the range between the highest and lowest values.
polygenes
A number of different genes at different loci that all contribute to a particular aspect of phenotype.
+31 more definitions
View all →Command Word Guide
| Explain | When explaining natural selection, ensure you include variation, selection pressure, differential survival/reproduction, and changes in allele frequency over generations. For speciation, clearly state how reproductive isolation prevents gene flow. |
| Describe | When describing continuous variation, mention both polygenic inheritance and environmental influence, and that it results in a range of phenotypes. For discontinuous variation, emphasize 'distinct categories' and 'no intermediates' and link to single gene control. |
| Discuss | When discussing artificial selection, include both its benefits (e.g., fixing desired traits) and drawbacks (e.g., inbreeding depression, reduced genetic diversity). For evolutionary relationships, mention DNA sequencing alongside morphological, physiological, and behavioural features. |
| Calculate | For Hardy-Weinberg calculations, show all steps clearly, starting from q² to find q, then p, and then p² and 2pq. Ensure the final sum of genotype frequencies equals 1. |
+1 more
View all →Common Mistakes
Confusing genetic variation with environmentally induced variation.
Only genetic variation is heritable and can be acted upon by natural selection. Environmentally-induced variation is not passed on to offspring.
Thinking natural selection is a conscious process or that organisms 'try' to adapt.
Natural selection is a passive process where the environment selects pre-existing advantageous traits; organisms do not consciously adapt.
Confusing genetic drift with natural selection.
Genetic drift is a random process due to chance, whereas natural selection is non-random, driven by environmental pressures.
+3 more
View all →This chapter explores the concept of species and how organisms are classified into a hierarchical system of three domains and four eukaryotic kingdoms. It defines biodiversity at multiple levels, outlines methods for its assessment using sampling and statistical tools, and discusses the critical reasons for its maintenance, causes of extinction, and various conservation strategies.
biological species — A group of organisms with similar morphology and physiology, which can breed together to produce fertile offspring and are reproductively isolated from other species.
This concept defines species based on their ability to interbreed and produce fertile offspring, emphasizing reproductive isolation as a key criterion. For example, two different breeds of dog can mate and have fertile puppies, making them the same biological species. However, a dog and a cat cannot produce fertile offspring, so they are different biological species.
Students often think that if two organisms look similar, they must be the same species, but actually, the ability to produce fertile offspring is the defining characteristic of a biological species.
When asked to define 'species', ensure you include 'produce fertile offspring' and 'reproductively isolated' for full marks, especially in 'Define' or 'Explain' questions.
morphological species — A group of organisms that share many physical features that distinguish them from other species.
This concept is often used when the biological species concept cannot be applied, such as for fossil species or organisms that reproduce asexually. It relies on observable physical characteristics like outward appearance (morphology) and anatomy. For instance, different types of roses in a garden all look like roses, sharing similar flower and leaf structures, allowing classification based on morphology.
Be prepared to discuss the limitations of the morphological species concept, such as cryptic species (look alike but are reproductively isolated) or phenotypic variation within a single species.
ecological species — A population of individuals of the same species living in the same area at the same time.
This concept focuses on a population of organisms that share common features and occupy the same habitat at the same time, highlighting their ecological role and interactions within a specific environment. For example, a group of deer living in a particular forest at the same time, all sharing the same food sources and interacting with each other, would be considered an ecological species within that forest.
Students often think 'ecological species' refers to the species' role, but actually, it specifically refers to a population of individuals of the same species coexisting in the same place and time.
population — All of the organisms of the same species present in the same place and at the same time that can interbreed with one another.
A population is a fundamental unit in ecology and evolution, representing a group of individuals of a single species that share a common gene pool and interact within a defined geographical area and time frame. All the students in a specific biology class at a particular school during one academic year form a population of students, as they are all of the same 'species' (students), in the same place, at the same time, and can interact.
When defining 'population', ensure you specify 'same species', 'same place', and 'same time' to achieve full accuracy.
biological classification — The organisation of living and extinct organisms into systematic groups based on similarities and differences between species.
This process helps biologists manage the vast diversity of life, making it easier to study, understand, and remember the key features of different organisms by grouping them into categories. It's like organizing books in a library: you group similar books (e.g., fiction, non-fiction) into categories, then subcategories, making it easier to find and understand the collection.
When asked to 'Explain' the purpose of classification, mention both 'understanding relationships' and 'ease of study/communication'.
taxonomy — The study and practice of naming and classifying species and groups of species within the hierarchical classification scheme.
Taxonomy is the scientific discipline concerned with the identification, naming, and classification of organisms, involving the assignment of organisms to specific taxonomic units (taxa) within a hierarchical system. A librarian who creates the system for categorizing and naming all the books in a library is doing taxonomy for books.
Students often confuse 'taxonomy' with 'classification', but actually, taxonomy is the broader study and practice of classification, including the naming aspect.
Ensure you include both 'naming' and 'classifying' when defining taxonomy, as both are crucial aspects of the discipline.
hierarchical classification — The arrangement of organisms into groups of different rank, from species (lowest) to domain (highest), where similar groups are nested within larger, more inclusive groups.
This system organizes life into a series of nested categories, allowing for a systematic way to show evolutionary relationships and shared characteristics, moving from very specific groups to very broad ones. It's like a set of Russian nesting dolls: each smaller doll (species) fits inside a slightly larger one (genus), and so on, up to the largest doll (domain).
Be able to list the taxonomic ranks in order (Domain, Kingdom, Phylum, Class, Order, Family, Genus, Species) and explain why it is hierarchical (groups within groups).
taxonomic rank — One of the groups used in the hierarchical classification system for organisms, such as species, genus, family, order, class, phylum, kingdom and domain.
Each taxonomic rank represents a level in the hierarchy, with organisms at lower ranks sharing more specific characteristics and being more closely related than those at higher ranks. In a company, 'CEO', 'Manager', and 'Employee' are different ranks; similarly, 'Kingdom', 'Phylum', and 'Species' are different ranks in biological classification.
taxon (plural: taxa) — A taxonomic group of any rank, such as a particular species (e.g. Giraffa camelopardalis), a family (e.g. Elephantidae), a class (e.g. Mammalia) or a kingdom (e.g. Plantae).
A taxon is a named group of organisms at any level of the classification hierarchy, representing a concrete group of organisms that share common characteristics and are distinct from other groups. If 'class' is a taxonomic rank, then 'Mammalia' is a specific taxon within that rank, referring to all mammals.
domain — The highest taxonomic rank.
The domain is the broadest category in the hierarchical classification system, dividing all life into three major groups: Bacteria, Archaea, and Eukarya, based on fundamental differences in cell structure and molecular biology. Think of the 'domain' as the largest continent on Earth, encompassing many different countries (kingdoms) and cities (species).
Students often think 'kingdom' is the highest rank, but actually, 'domain' was introduced later to reflect deeper evolutionary divergences, especially among prokaryotes.
When discussing domains, ensure you mention the three specific domains (Bacteria, Archaea, Eukarya) and the key differences that led to their establishment, particularly the molecular evidence.
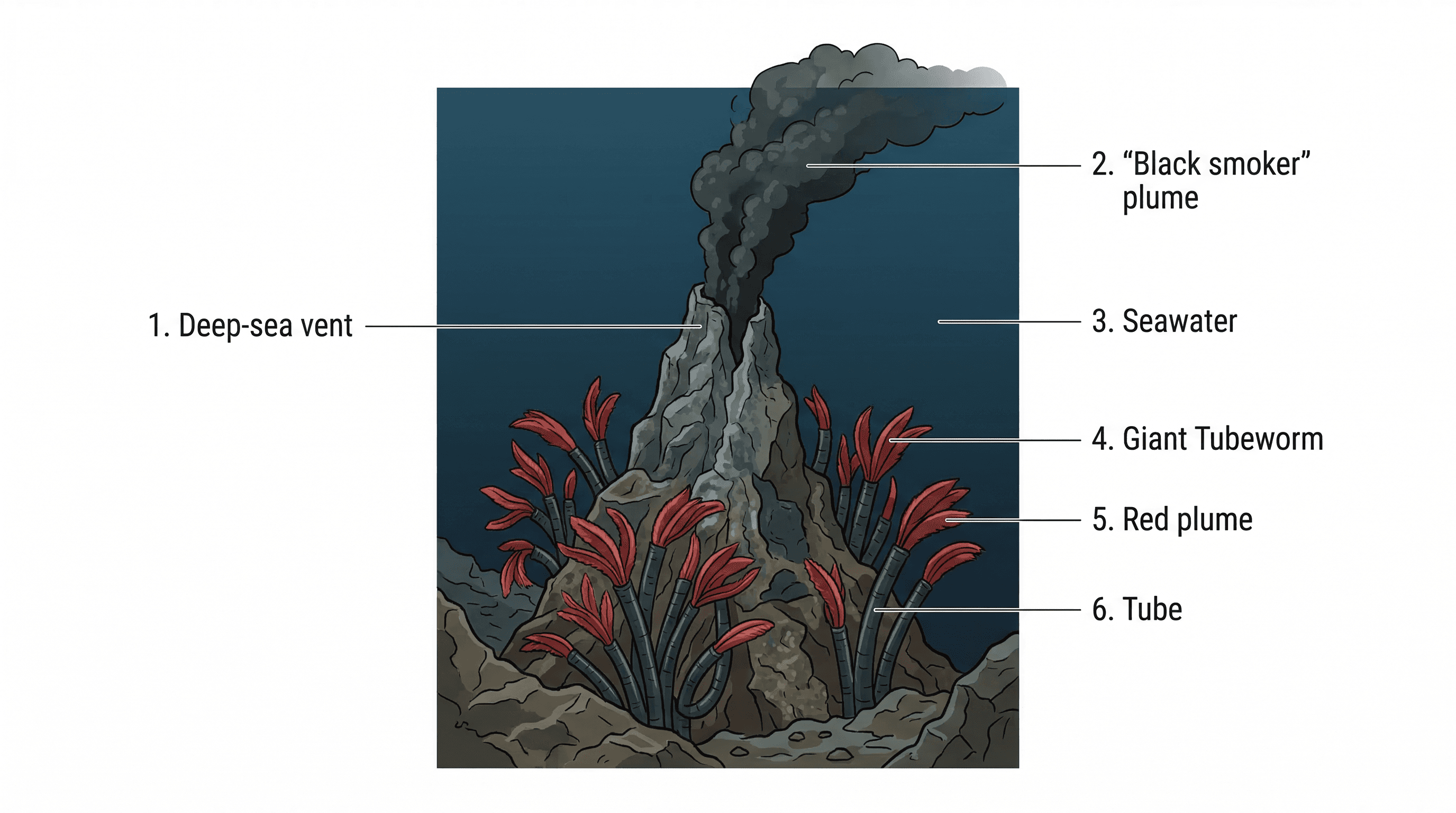

kingdom — The taxonomic rank below domain.
Kingdoms are broad groupings within domains, further categorizing organisms based on general characteristics such as cell type, mode of nutrition, and body organization. The Eukarya domain is divided into four kingdoms: Protoctista, Fungi, Plantae, and Animalia. If 'domain' is a continent, then 'kingdom' is a large country within that continent, containing many different regions (phyla) and cities (species).
Be able to list the four eukaryotic kingdoms and their defining characteristics, especially for 'Outline' or 'Describe' questions.
Bacteria — The domain that contains all prokaryotic organisms except those classified as Archaea.
Bacteria are single-celled prokaryotes characterized by the absence of a nucleus and membrane-bound organelles, having circular DNA without histone proteins, and cell walls containing peptidoglycans. Think of bacteria as the 'classic' single-celled organisms, like the common germs you might hear about, distinct from the more 'extreme' archaeans.
Archaea — The domain of prokaryotic organisms that resemble bacteria but share some features with eukaryotes.
Archaeans are prokaryotes that often inhabit extreme environments and possess unique membrane lipids, ribosomal RNA sequences more similar to eukaryotes, and cell walls without peptidoglycans, distinguishing them from Bacteria. Imagine Archaea as the 'ancient survivors' of the microbial world, thriving in conditions that most other life forms cannot tolerate.
Highlight the 'extreme environments' and the unique molecular features (rRNA, membrane lipids) when describing Archaea, as these are key to their classification.
Eukarya — The domain that contains all eukaryotic organisms: protoctists, fungi, plants and animals.
Eukarya is characterized by cells possessing a nucleus and membrane-bound organelles, with DNA arranged in linear chromosomes associated with histone proteins, and larger (80S) ribosomes in the cytosol. Think of Eukarya as all the 'complex' life forms you can see with the naked eye, plus many microscopic ones, all sharing the fundamental feature of having cells with internal compartments.
When describing Eukarya, emphasize the presence of a nucleus and membrane-bound organelles as the defining characteristics, and mention the four kingdoms within it.
Protoctista — Kingdom of eukaryotic organisms which are single-celled or made up of groups of similar cells.
This is a highly diverse kingdom of eukaryotes that do not fit into the Fungi, Plantae, or Animalia kingdoms. They can be animal-like (protozoa) or plant-like (algae), with varying cell structures and modes of nutrition. Think of the Protoctista as the 'miscellaneous' drawer of eukaryotes; if it has a nucleus but isn't clearly a fungus, plant, or animal, it probably belongs here.
protoctist — A member of the Protoctista kingdom.
Protoctists are a diverse group of eukaryotic organisms, including single-celled forms like amoebas and paramecia, and simple multicellular forms like seaweeds, characterized by their varied cellular organization and nutritional strategies. A protoctist is like a 'biological odd job' organism; it's eukaryotic but doesn't quite fit into the more defined categories of plants, animals, or fungi.
Provide examples of protoctists (e.g., Amoeba, Volvox, seaweed) to illustrate their diversity when asked to 'Describe' them.
Fungi — Kingdom of eukaryotic organisms which do not photosynthesise and have cell walls but without cellulose.
Fungi are heterotrophic eukaryotes that obtain nutrients by absorbing organic compounds from their environment, often through decomposition or parasitism. Their cell walls are typically made of chitin, not cellulose, and they reproduce via spores. Think of fungi as the 'recyclers' of the biological world; they break down dead matter and absorb nutrients, much like a compost heap.
Students often confuse fungi with plants due to their stationary nature and cell walls, but actually, fungi are heterotrophic and have chitin cell walls, unlike plants which are autotrophic with cellulose walls.
Emphasize heterotrophic nutrition, chitin cell walls, and reproduction by spores as key distinguishing features of fungi in 'Outline' or 'Compare' questions.
Plantae — Kingdom of eukaryotic organisms which are multicellular, have cell walls that contain cellulose and can photosynthesise.
Plants are multicellular, autotrophic eukaryotes that produce their own food through photosynthesis, possess specialized cells forming tissues and organs, and have rigid cell walls primarily composed of cellulose. Think of plants as the 'primary producers' of most ecosystems, converting sunlight into energy, much like a solar panel converts light into electricity.
When describing Plantae, ensure you include multicellularity, autotrophic nutrition (photosynthesis), and cellulose cell walls as defining characteristics.
Animalia — Kingdom of eukaryotic organisms which are multicellular and heterotrophic and have a nervous system.
Animals are multicellular, heterotrophic eukaryotes that obtain food by ingestion, lack cell walls and chloroplasts, and are characterized by specialized cells, tissues, and organs, including a unique nervous system for communication. Think of animals as the 'consumers' of the biological world, actively seeking and ingesting food, and often moving and reacting to their environment.
Key features for Animalia include multicellularity, heterotrophic nutrition, absence of cell walls, and the presence of a nervous system for communication.
Viruses are classified separately from the three domains and four eukaryotic kingdoms. They are not considered living organisms in the same way as cellular life forms, as they lack cellular structure and metabolic machinery, relying instead on host cells for replication.
biodiversity — The variety of ecosystems and species in an area and the genetic diversity within each species.
Biodiversity encompasses three levels: the diversity of ecosystems and habitats, the number and relative abundance of different species, and the genetic variation within individual species. High biodiversity generally leads to more stable and resilient ecosystems. Imagine a diverse library with many different genres of books (ecosystems), many different authors (species), and many different editions or translations of each book (genetic diversity).
Students often think biodiversity is just about the number of species, but actually, it also includes ecosystem diversity and genetic diversity within species.
When asked to 'Explain the importance of biodiversity', ensure you address all three levels (ecosystem, species, genetic) and link them to ecosystem stability, resilience, and potential for future resources.
endemic — Of species, a species that is only found in a certain area and nowhere else.
Endemic species are geographically restricted to a particular region, making them particularly vulnerable to habitat loss or environmental changes in that specific area, as they have no other populations elsewhere. Think of a local restaurant that only serves a dish unique to that town; that dish is 'endemic' to that town because you can't find it anywhere else.
ecosystem — A relatively self-contained, interacting community of organisms, and the environment in which they live and with which they interact.
An ecosystem includes both the living (biotic) components, such as plants, animals, and microorganisms, and the non-living (abiotic) components, such as soil, water, and air, all interacting as a functional unit. A fish tank is a small ecosystem: the fish, plants, and bacteria are the community, and the water, gravel, and filter are the environment they interact with.
When defining 'ecosystem', ensure you mention both 'community of organisms' and 'environment' and their 'interactions' for a complete definition.
community — All of the living organisms, of all species, that are found in a particular ecosystem at a particular time.
A community refers to the collection of different populations of species that live and interact within a specific area, forming a complex web of relationships such as predation, competition, and symbiosis. In a neighborhood, all the different types of people (families, individuals, different professions) living and interacting together form a community.
habitat — The place where an organism, a population or a community lives.
A habitat is the specific physical environment where a species or group of species naturally resides, providing the necessary resources and conditions for their survival and reproduction. Your home address is your habitat; it's the specific place where you live.
niche — The role of an organism in an ecosystem; it is how the organism ‘fits into’ the ecosystem.
An organism's niche describes its specific role, including how it obtains energy, its interactions with other species (e.g., predator, prey, competitor), and its relationship with the physical environment (e.g., temperature, light requirements). If a habitat is an organism's address, its niche is its job description, including what it eats, who eats it, and how it affects its surroundings.
When defining 'niche', ensure you include aspects beyond just location, such as 'how it obtains energy' and 'how it interacts with both its physical environment and with other species'.
species diversity — All the species in an ecosystem.
Species diversity considers both species richness (the number of different species) and species evenness (the relative abundance of each species). Higher diversity implies a greater number of species and a more balanced distribution of individuals among those species. Imagine two classrooms: one has 20 students, all from different countries (high species richness). Another has 20 students, with 19 from one country and 1 from another (low species evenness). Species diversity considers both.
Students often equate species diversity solely with species richness, but actually, it also includes species evenness, meaning how balanced the populations of those species are.
When comparing species diversity, remember to consider both species richness and species evenness; a high number of species with one dominant species results in lower diversity than a similar number of species with more even abundances.
genetic diversity — All the alleles of all the genes in the genome of a species.
Genetic diversity refers to the variation in genetic material (alleles) within a single species. This diversity is crucial for a species' ability to adapt to changing environmental conditions, resist diseases, and survive long-term. Think of a deck of cards for a species; genetic diversity is like having a full deck with many different suits and numbers (alleles), rather than a deck with only aces of spades.
When discussing genetic diversity, link it directly to the ability of a population to adapt to changes in biotic and abiotic factors, such as disease resistance or climate change.
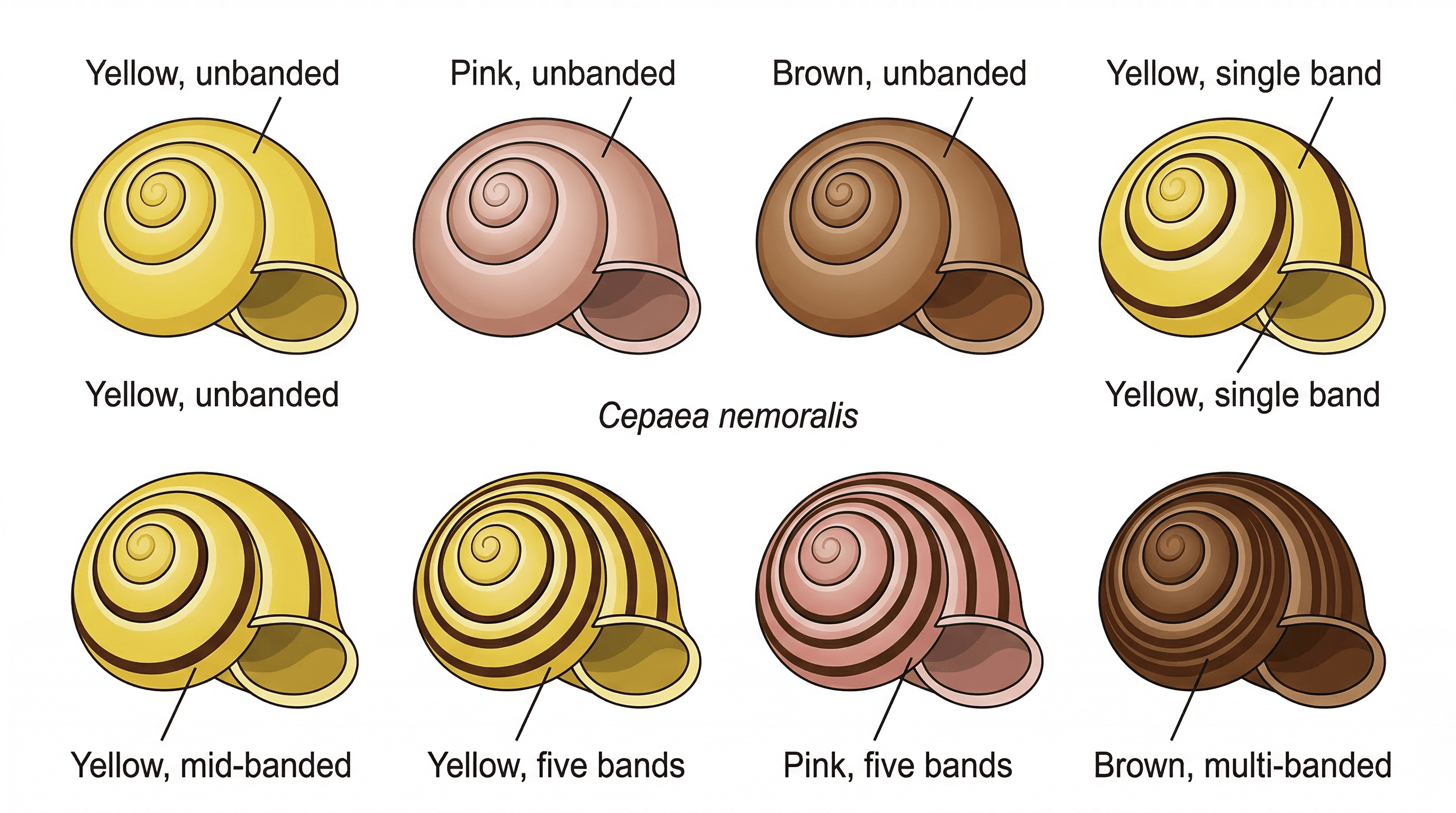
To assess species diversity, ecologists employ various sampling techniques to estimate the occurrence, abundance, and distribution of species within an ecosystem. This involves collecting organisms, making species lists, and using methods like random or systematic sampling to gather representative data.
random sampling — Method of investigating the abundance and/or distribution of populations which is determined by chance and shows no bias on the part of the person carrying out the sampling.
Random sampling ensures that every part of the study area has an equal chance of being sampled, thereby producing data that is representative of the entire area and minimizing observer bias. Picking names out of a hat to choose who gets a prize is random sampling; everyone has an equal chance, and the selection is unbiased.
Students often think 'random' means haphazard, but actually, it requires a systematic method (e.g., using random number generators for coordinates) to ensure true randomness and avoid bias.
When describing random sampling, always mention using a random number generator or similar method to ensure unbiased placement of quadrats.
quadrat — A square frame which is used to mark out an area for sampling populations of organisms.
Quadrats are used in ecological surveys to define a specific area for counting individuals, estimating percentage cover, or recording species frequency, particularly for sessile or slow-moving organisms. A quadrat is like a picture frame that you place on the ground to focus on and study a small, defined section of a larger landscape.
When describing quadrat use, mention its application for measuring species frequency, species density, or percentage cover, and the importance of appropriate size and random placement.
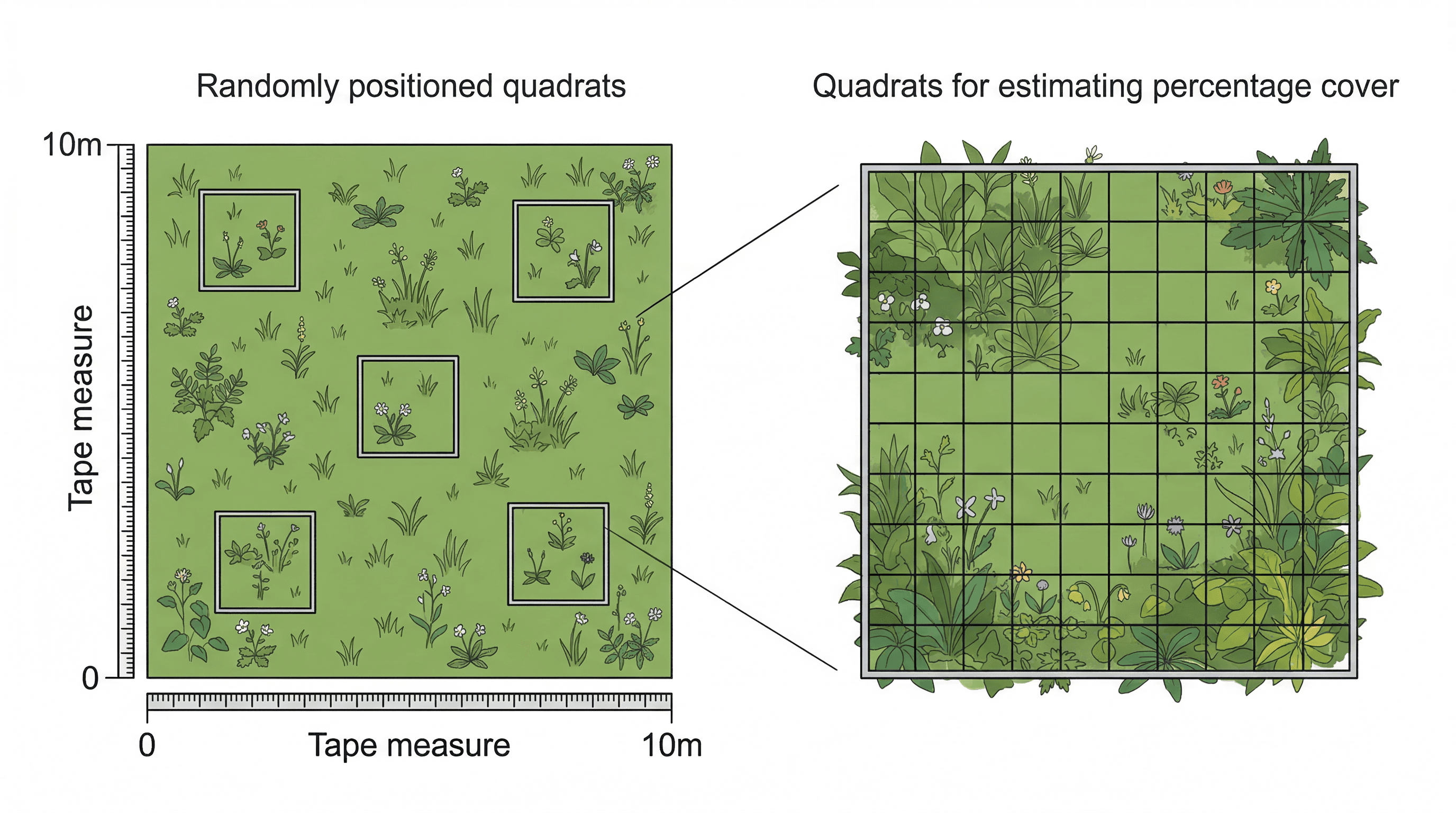
mark–release–recapture — A method of estimating the numbers of individuals in a population of mobile animals.
This technique involves capturing a sample of mobile animals, marking them, releasing them, and then recapturing a second sample to estimate the total population size. It assumes no immigration, emigration, births, or deaths between samples, and that marking does not affect survival or recapture probability.
Lincoln Index (Population Size Estimate)
Used for estimating population size of mobile animals using the mark–release–recapture technique. Assumes no immigration, emigration, births, or deaths between samples, and that marking does not affect survival or recapture probability.
Simpson’s index of diversity (D)
Used to calculate the biodiversity of a habitat. Values range from 0 (low diversity) to 1 (high diversity). A higher value indicates greater species diversity, considering both richness and evenness.
systematic sampling — A non-random method of investigating the abundance and/or distribution of populations in which the position of sampling points are determined by the person carrying out the sampling (e.g. at every 2 m along a transect).
Systematic sampling is used when there is a clear environmental gradient or pattern in the distribution of species, allowing researchers to investigate how species abundance or distribution changes along that gradient. If you want to see how house prices change as you move away from the city center, you might check prices every mile along a straight road; this is systematic, not random.
Students often think systematic sampling is always inferior to random sampling, but actually, it is more appropriate and effective when investigating environmental gradients or patterns.
Specify that systematic sampling is used when conditions are 'not uniform' or when investigating 'gradients' (e.g., along a transect) to justify its use over random sampling.
transect — A line marked by a tape measure along which samples are taken, either by noting the species at equal distances (line transect) or placing quadrats at regular intervals (belt transect).
Transects are used in systematic sampling to study how species distribution and abundance change along an environmental gradient, such as from the seashore to inland, or up a mountain slope. They provide a structured way to record data at regular intervals.
Kite diagrams are a visual tool used to display data on species abundance along a transect. They effectively illustrate how the distribution and abundance of different species vary with changes in environmental factors. The width of the 'kite' at each sampling point represents the abundance of a particular species.
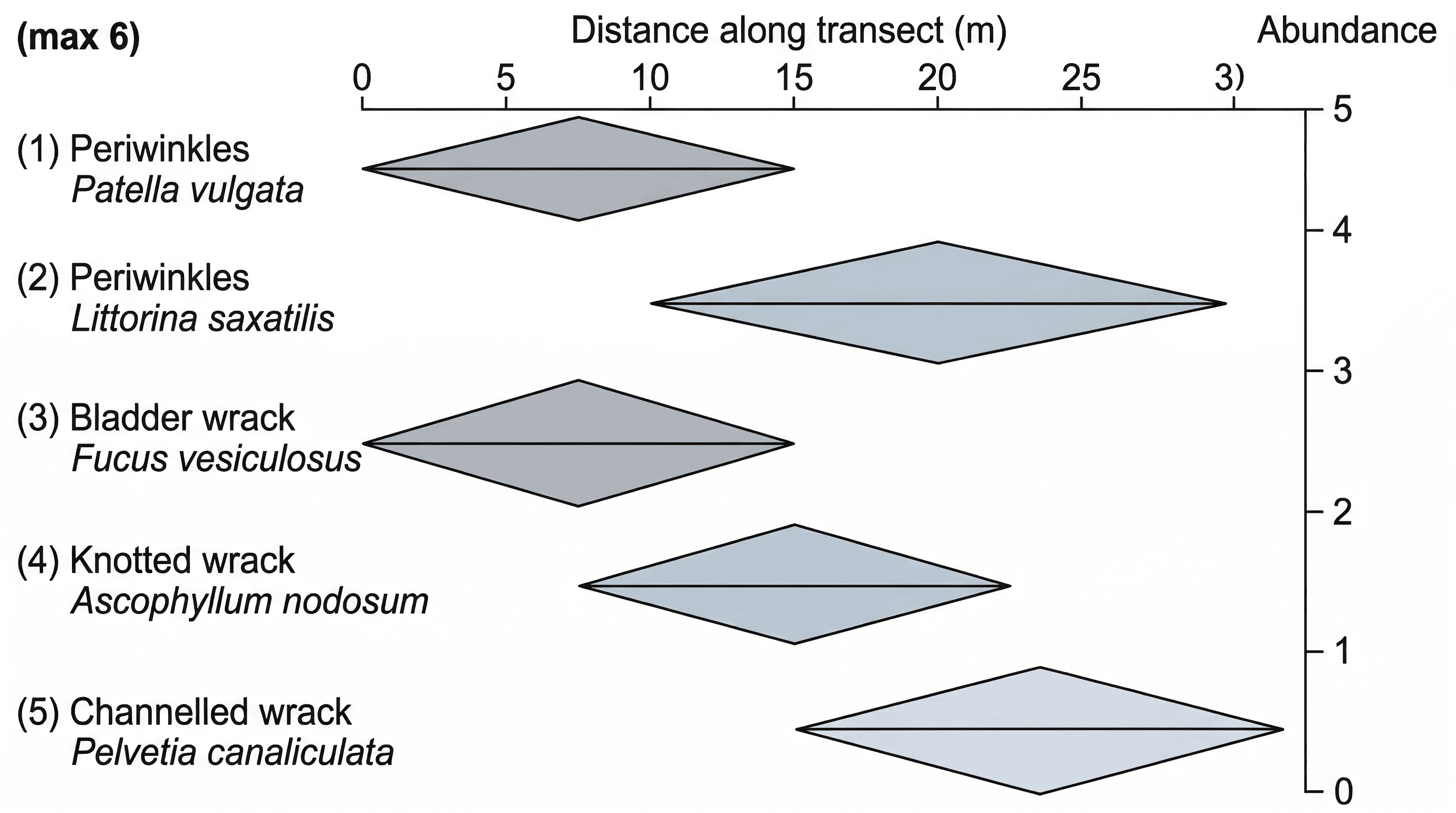
Correlation refers to a statistical relationship between two variables. It indicates whether two variables tend to change together, but it does not imply that one variable causes the other. Statistical methods like Spearman's rank correlation and Pearson's linear correlation are used to quantify the strength and direction of these relationships.
Students often assume correlation implies causation, but statistical tests like Pearson's and Spearman's only show a relationship, not cause and effect.
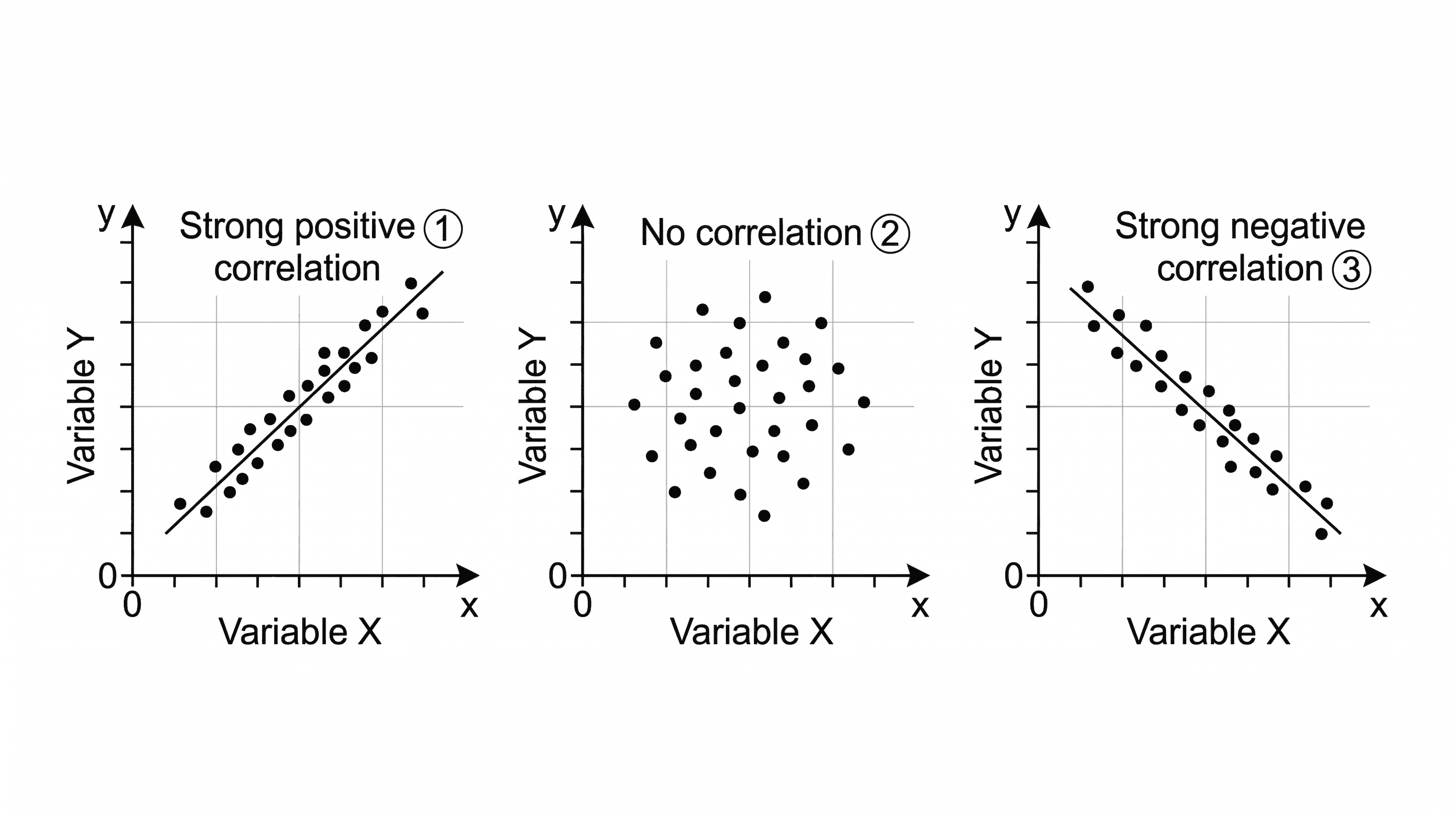
Spearman’s rank correlation — A statistical test to determine if there is a correlation between two variables when one or both of them are not normally distributed.
Spearman's rank correlation coefficient (r_s) is a non-parametric measure of the strength and direction of association between two ranked variables. It is particularly useful when data are ordinal or when the assumptions for Pearson's correlation are not met.
Spearman’s rank correlation coefficient (rs)
Used to determine if there is a correlation between two variables when one or both are not normally distributed, or when using ordinal data. Values range from -1 (perfect negative correlation) to +1 (perfect positive correlation).
Pearson’s linear correlation — A statistical test used to determine if there is a linear correlation between two variables that are normally distributed.
Pearson's linear correlation coefficient (r) measures the strength and direction of a linear relationship between two continuous variables. It requires that both variables are normally distributed and that the relationship is linear.
Pearson’s linear correlation coefficient (r)
Used to determine if there is a linear correlation between two continuous variables that are normally distributed. Values range from -1 (perfect negative linear correlation) to +1 (perfect positive linear correlation). Correlation does not imply causation.
Maintaining biodiversity is crucial for ecological stability, providing essential ecosystem services, and offering potential resources for human benefit. The loss of biodiversity, often due to human activities, leads to extinctions and reduces the resilience of ecosystems to environmental changes.
Species may become extinct due to various factors, including habitat loss and fragmentation, climate change, pollution, overexploitation, and the introduction of invasive alien species. These factors often interact, accelerating the rate of species decline.
alien species — A species that has moved into a new ecosystem where it was previously unknown; also known as invasive species.
Alien species, also known as invasive species, can pose a significant threat to native biodiversity by outcompeting native species for resources, preying on them, or introducing diseases. Controlling these species is often necessary to protect local ecosystems.
Conservation efforts aim to protect endangered species and their habitats through both in situ (in their natural habitat) and ex situ (outside their natural habitat) methods. In situ strategies include establishing national parks and protected areas, while ex situ methods involve zoos, botanic gardens, seed banks, and assisted reproduction techniques.
assisted reproduction — Any technique that is involved in treating infertility or protecting a female mammal of an endangered species from the health risks of pregnancy.
Assisted reproduction techniques are vital in the conservation of endangered mammal species, helping to increase population numbers and maintain genetic diversity. These methods include artificial insemination, in vitro fertilisation, and embryo transfer.
artificial insemination (AI) — Injection of semen collected from a male into the uterus.
Artificial insemination is a technique used in assisted reproduction where semen is collected from a male and directly injected into the uterus of a female. This method can overcome geographical barriers or behavioral incompatibilities between individuals, facilitating breeding in endangered species.
in vitro fertilisation (IVF) — The fertilisation of an egg that occurs outside the body of a female (e.g. in a Petri dish).
In vitro fertilisation involves the fertilization of an egg by sperm outside the body, typically in a laboratory setting. The resulting embryos can then be transferred into a surrogate mother or cryopreserved for future use, offering a way to breed endangered species with limited reproductive success.
embryo transfer — Embryos are removed from the uterus of a female mammal of an endangered species shortly after fertilisation and transferred to surrogate females to bring to full term.
Embryo transfer involves moving embryos from a donor female to a surrogate mother. This allows a single endangered female to produce more offspring than she could naturally, as multiple surrogates can carry pregnancies to term, maximizing reproductive output.
surrogacy — A female becomes pregnant with an embryo from another female and carries it to full term; embryos can be conceived naturally, by AI or by IVF.
Surrogacy in conservation involves a female carrying an embryo from another female, often of an endangered species, to full term. This can be achieved through natural conception, artificial insemination, or in vitro fertilisation, providing a means to increase the birth rate of vulnerable populations.
frozen zoo — A facility where genetic materials taken from animals are stored at very low temperatures (–196 °C); sperm, eggs, embryos and tissue samples are examples of these genetic materials.
Frozen zoos are critical ex situ conservation facilities that store genetic material such as sperm, eggs, embryos, and tissue samples from endangered animals at ultra-low temperatures. This preserves genetic diversity for future breeding programs or reintroduction efforts, acting as a genetic safeguard against extinction.
seed bank — Facility where seeds are dried and kept in cold storage to conserve plant biodiversity.
Seed banks are vital for plant conservation, storing seeds under controlled conditions (dried and cold) to maintain their viability for long periods. This preserves the genetic diversity of plant species, including endangered ones and those with agricultural importance, for future use or reintroduction.
Global conservation efforts are supported by international organisations like CITES and IUCN, which play crucial roles in regulating trade, assessing species' conservation status, and coordinating worldwide initiatives to protect biodiversity.
CITES — The Convention on International Trade in Endangered Species of Wild Fauna and Flora.
CITES is an international agreement that regulates the international trade of wild animals and plants to ensure that such trade does not threaten their survival. It lists species in appendices according to their level of threat, imposing varying degrees of control on their trade.
IUCN — The International Union for Conservation of Nature.
The IUCN is a global authority on the status of the natural world and the measures needed to safeguard it. It publishes the Red List of Threatened Species, which assesses the conservation status of species worldwide, guiding conservation action and policy.
For all calculations (Simpson's, Lincoln, Spearman's), write out the formula, show your full working, and give your answer to an appropriate number of significant figures.
When interpreting a calculated correlation coefficient, always state the strength (strong/weak), direction (positive/negative), and conclude by stating that correlation does not prove causation.
When asked for reasons to maintain biodiversity, structure your answer with ecological, economic, and ethical points.
Be prepared to draw and interpret kite diagrams. Remember the width of the kite at any point along the transect line represents the abundance of the species.
When describing sampling techniques, be specific. For random sampling, state you would use a grid and random number generator to place quadrats. For systematic, mention laying a transect line and sampling at regular intervals.
Definitions Bank
biological species
A group of organisms with similar morphology and physiology, which can breed together to produce fertile offspring and are reproductively isolated from other species.
morphological species
A group of organisms that share many physical features that distinguish them from other species.
ecological species
A population of individuals of the same species living in the same area at the same time.
population
All of the organisms of the same species present in the same place and at the same time that can interbreed with one another.
biological classification
The organisation of living and extinct organisms into systematic groups based on similarities and differences between species.
+38 more definitions
View all →Command Word Guide
| Define | Provide a precise, one-sentence biological definition. For 'species', include 'fertile offspring' and 'reproductively isolated'. For 'ecosystem', include 'community', 'environment', and 'interactions'. |
| Explain | Go beyond a definition; provide reasons, mechanisms, or implications. For 'classification', explain its purpose (understanding relationships, ease of study). For 'biodiversity', explain its importance at ecosystem, species, and genetic levels. |
| Outline | Give the main features or general principles. For 'domains' or 'kingdoms', list them and state their key distinguishing characteristics without excessive detail. |
| Discuss | Present a balanced argument or consider different aspects of a topic. For 'reasons for maintaining biodiversity', include ecological, economic, and ethical points. For 'species concepts', compare their strengths and limitations. |
+2 more
View all →Common Mistakes
Thinking organisms that look similar must be the same species.
The biological species concept defines species by their ability to interbreed and produce fertile offspring, and being reproductively isolated from other species, not just by appearance.
Confusing 'taxonomy' with 'classification'.
Taxonomy is the broader scientific study and practice of naming and classifying organisms, while classification is the act of organizing them into groups.
Believing Kingdom is the highest taxonomic rank.
Domain is the highest taxonomic rank, above Kingdom, reflecting deeper evolutionary divergences.
+4 more
View all →This chapter explores genetic technology, detailing techniques like genetic engineering, PCR, gel electrophoresis, and microarrays. It covers the tools used, such as restriction enzymes and vectors, and discusses applications in medicine and agriculture, alongside their significant social and ethical implications.
genetic engineering — Any procedure in which the genetic information in an organism is changed by altering the base sequence of a gene or by introducing a gene from another organism; the organism is then said to be a genetically modified organism (GMO).
Genetic engineering allows for targeted changes to an organism's DNA, often involving the transfer of a single gene between different species. Imagine editing a specific sentence in a book (the genome) to change a character's trait, rather than just picking the best existing books from a library.
Students often think genetic engineering is the same as selective breeding, but actually genetic engineering involves direct manipulation of DNA, often across species, while selective breeding relies on choosing individuals with desirable traits for reproduction.
genetically modified organism (GMO) — Any organism that has had its DNA changed in a way that does not occur naturally or by selective breeding.
GMOs are organisms whose genetic material has been altered using genetic engineering techniques. This alteration can involve adding, deleting, or changing genes to introduce new characteristics, such as disease resistance or enhanced growth. Think of a car that has been specifically engineered with a new engine or feature that wasn't part of its original design or a standard upgrade.
recombinant DNA (rDNA) — DNA made by artificially joining together pieces of DNA from two or more different species.
rDNA is a key product of genetic engineering, where a desired gene from one organism is combined with the DNA of a vector (like a plasmid) from another. This allows the gene to be introduced and expressed in a new host. Think of cutting a paragraph from one book and pasting it into another book, creating a new combined text.
Students often think rDNA is just any altered DNA, but actually it specifically refers to DNA constructed from two or more different sources, often different species.
transgenic organism — Any organism that contains DNA from another source, such as from another individual of the same species or from a different species.
A transgenic organism is a type of GMO that has incorporated foreign DNA into its genome. This foreign DNA can originate from the same species or a different one, leading to the expression of new traits. Like a computer program that has a new piece of code (DNA) added from a different program to give it a new function.
Use precise terminology. Refer to 'restriction endonucleases', 'vectors', 'recombinant DNA', and 'transgenic organisms' rather than vague terms like 'genetic scissors' or 'changed organism'.
Genetic engineering relies on a suite of molecular tools to manipulate DNA. These include enzymes for cutting and joining DNA, and vectors for delivering genes into host cells. Understanding these tools is fundamental to performing genetic modifications.
restriction endonuclease (restriction enzyme) — An enzyme, originally derived from bacteria, that cuts DNA molecules; each type of restriction enzyme cuts only at a particular sequence of bases.
Restriction enzymes are molecular 'scissors' that recognize specific palindromic sequences (restriction sites) on DNA and cleave the phosphodiester backbone. They are essential for cutting out desired genes and opening plasmid vectors, often creating 'sticky ends'. Like a very precise pair of scissors that only cuts paper at specific, pre-marked lines.
Students often think restriction enzymes cut DNA randomly, but actually each enzyme recognizes and cuts at a very specific base sequence.
Emphasize the 'specificity' of restriction enzymes – they cut at 'particular sequences of bases' or 'restriction sites' – as this is a key characteristic.
sticky ends — Short lengths of unpaired bases that form hydrogen bonds with complementary sequences of bases on other pieces of DNA cut with the same restriction enzyme.
Sticky ends are created by restriction enzymes that make staggered cuts in DNA, leaving overhangs of single-stranded DNA. These overhangs are crucial for genetic engineering as they allow DNA fragments from different sources, cut with the same enzyme, to anneal together. Like two pieces of a jigsaw puzzle that have complementary shapes, allowing them to fit together perfectly.
cDNA — Double-stranded complementary DNA formed from an mRNA template using reverse transcriptase and DNA polymerase.
cDNA is a DNA copy of an mRNA molecule, lacking introns because it's synthesized from processed mRNA. This makes it particularly useful in genetic engineering for expressing eukaryotic genes in prokaryotic hosts, which cannot splice introns. Imagine making a clean, edited transcript (cDNA) from a raw recording (mRNA) that already had all the irrelevant parts (introns) removed.
Highlight that cDNA is derived from mRNA and thus lacks introns, which is a key advantage when expressing eukaryotic genes in bacteria.
vector — A means of delivering genes into a cell used in gene technology; e.g. plasmids and viruses.
Vectors are crucial tools in genetic engineering, acting as carriers to transport desired genes into host cells. Plasmids and viruses are commonly used because they can naturally enter cells and integrate genetic material. Think of a delivery truck (the vector) carrying a package (the gene) to a specific house (the cell).
When asked for examples of vectors, always include both plasmids and viruses, as they are the most common and distinct types.
promoter — A length of DNA that includes the binding site for RNA polymerase where transcription of a gene or genes begins; in eukaryotes, promoters also have sites for binding of transcription factors.
Promoters are regulatory DNA sequences located upstream of a gene that control its expression. RNA polymerase binds to the promoter to initiate transcription, and in eukaryotes, transcription factors are also required to facilitate this binding and regulate gene activity. Like the 'on/off' switch and 'start' button for a machine (the gene), determining when and where it begins to operate.
Remember to mention the role of a promoter. A gene inserted into a vector will not be expressed unless it is placed after a suitable promoter sequence that allows RNA polymerase to bind and begin transcription.
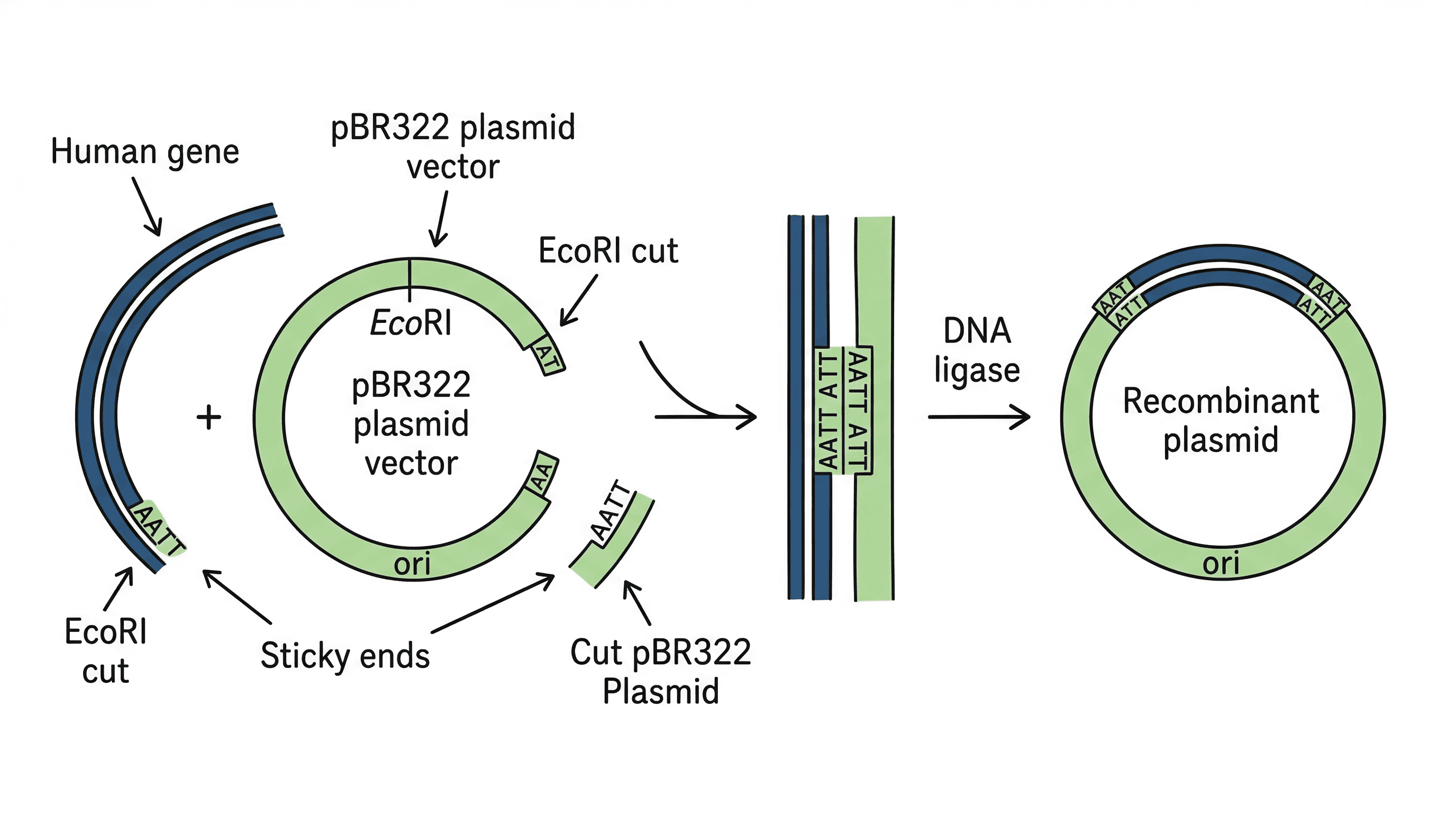
The process of creating recombinant DNA involves isolating a desired gene and inserting it into a plasmid vector. This is achieved by cutting both the gene and the plasmid with the same restriction enzyme, creating complementary sticky ends. These sticky ends then anneal, and DNA ligase forms phosphodiester bonds to seal the gene into the plasmid, forming recombinant DNA.
When describing how to create a recombinant plasmid, always state that the *same* restriction enzyme must be used on both the gene and the plasmid to create complementary sticky ends.
gene editing — A form of genetic engineering in which the genome of an organism can be changed by deleting, inserting or replacing a length of DNA using a method such as the Crispr/Cas9 system.
Gene editing, particularly with Crispr/Cas9, allows for precise, targeted modifications to an organism's DNA at specific locations. This offers greater control and accuracy compared to older genetic engineering methods that inserted DNA randomly. Instead of randomly inserting a new paragraph into a book, gene editing is like using a word processor to precisely cut, paste, or replace specific words or sentences.
When discussing gene editing, highlight its key advantage: the ability to make *precise* and *targeted* changes (deleting, inserting, or replacing) at *specific* locations in the genome.
Genetic technology often requires the amplification of specific DNA sequences or the separation of DNA fragments by size. Techniques like Polymerase Chain Reaction (PCR) and gel electrophoresis are fundamental for these purposes, enabling detailed analysis of genetic material.
polymerase chain reaction (PCR) — An automated process that amplifies selected regions of DNA using alternate stages of polynucleotide separation (denaturation of DNA) and DNA synthesis catalysed by DNA polymerase.
PCR is a powerful technique used to create millions of copies of a specific DNA segment from a very small initial sample. It involves repeated cycles of heating to separate DNA strands, cooling to allow primers to bind, and then DNA synthesis by a heat-stable DNA polymerase. Imagine a molecular photocopier that can make billions of copies of a specific page (DNA segment) from a single original page.
Students often think PCR copies the entire DNA sample, but actually it only amplifies *selected regions* of DNA, defined by the primers used.
For PCR questions, clearly state the purpose of each temperature stage: ~95°C to denature DNA (separate strands), ~55-65°C to anneal primers, and ~72°C for extension by heat-stable Taq polymerase.
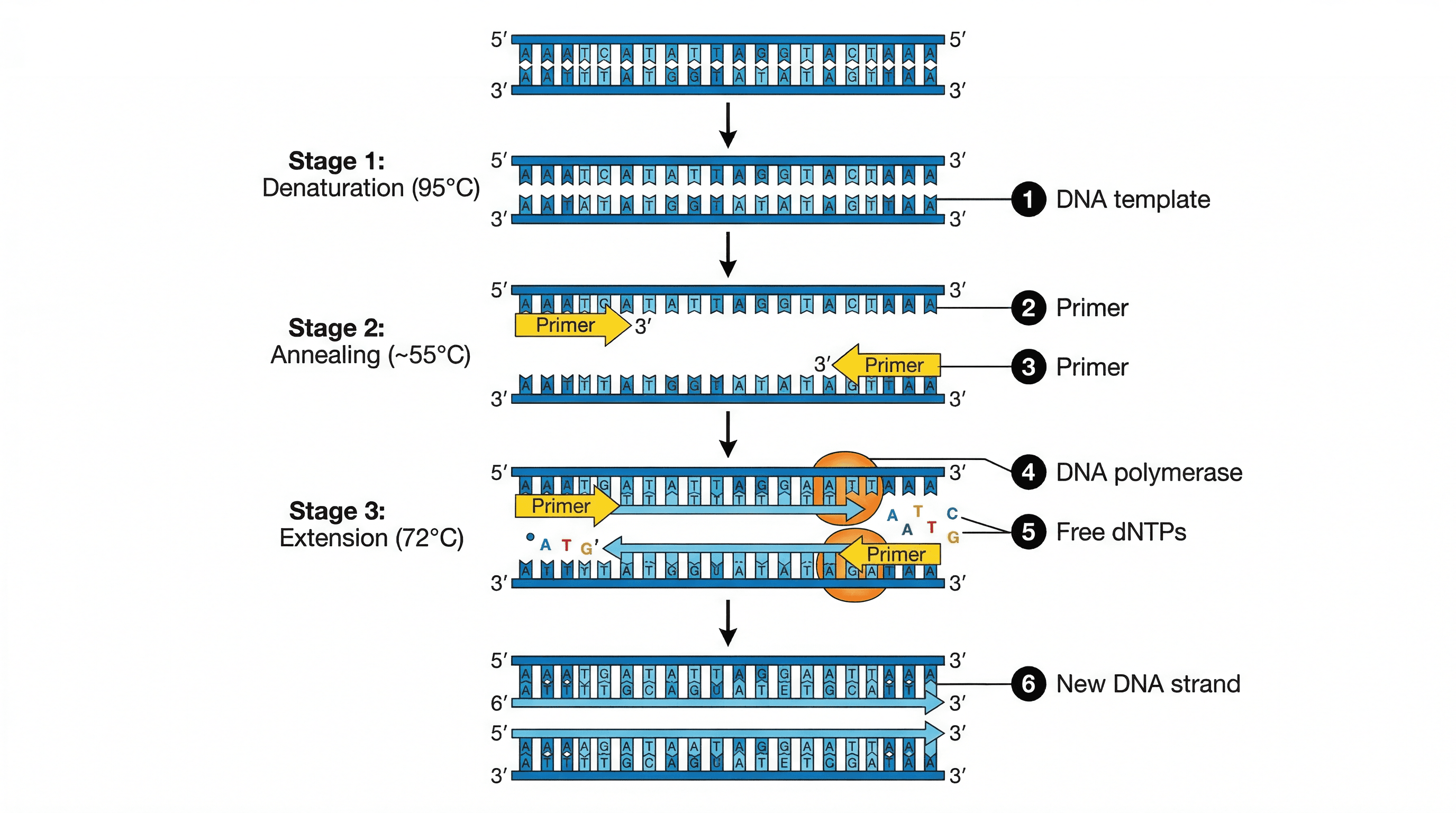
gel electrophoresis — The separation of charged molecules (e.g. DNA) by differential movement through a gel in an electric field; the degree of movement is dependent on the mass of the fragments of DNA.
Gel electrophoresis separates DNA fragments based on their size and charge. DNA, being negatively charged, moves towards the positive electrode. Smaller fragments move faster and further through the gel matrix than larger ones, allowing for size-based separation. Like a race where smaller, lighter runners (DNA fragments) can move faster through a crowded obstacle course (the gel) than larger, heavier runners.
Remember that DNA fragments are negatively charged and move towards the anode (+). The key factor for separation is size: smaller fragments travel further.
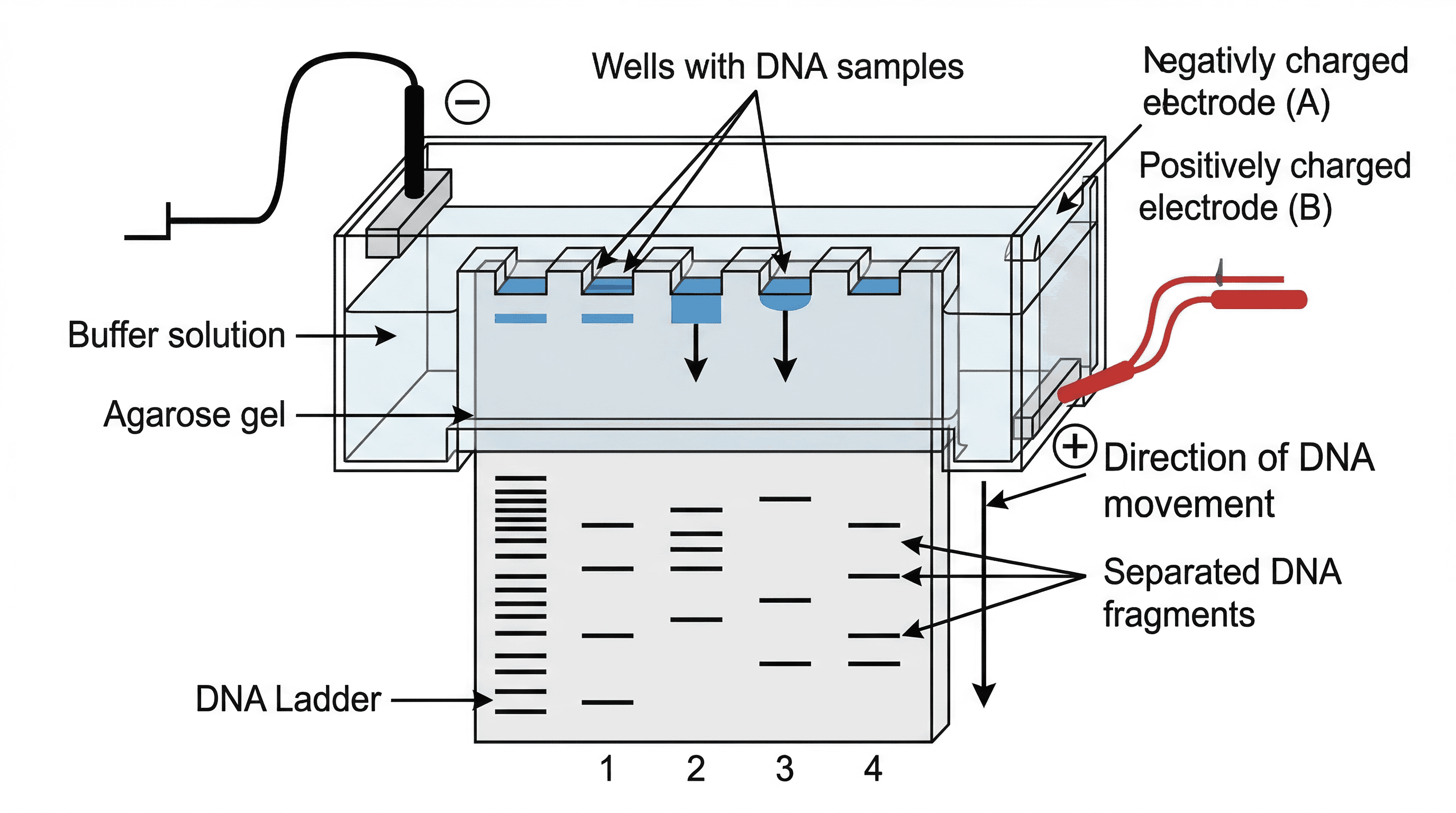
Modern genetic technologies generate vast amounts of data, necessitating advanced methods for analysis and storage. Microarrays allow for simultaneous analysis of gene expression, while bioinformatics provides the computational tools to manage and interpret complex biological datasets, including gene sequences and genomes.
microarray (also known as gene or DNA chips) — Slides that are printed with thousands of tiny spots in defined positions, with each spot containing a known DNA sequence; the DNA molecules attached to each slide act as probes to detect lengths of DNA or RNA with complementary sequences.
Microarrays are used to simultaneously analyze the expression of thousands of genes or to compare gene presence between different samples. Fluorescently labeled DNA or RNA samples are hybridized to the probes on the chip, and the resulting fluorescence indicates gene activity or presence. Imagine a library with thousands of tiny, labeled books (probes) on a single shelf. You can quickly see which of your own books (sample DNA/RNA) match and bind to the library's collection.
When describing microarrays, focus on their ability to analyze *many genes simultaneously* and their use of *hybridisation* with labeled samples to detect gene presence or expression.
DNA hybridisation — Binding together of two molecules of single-stranded DNA by complementary base pairing.
DNA hybridisation is the process where two complementary single-stranded DNA molecules (or DNA and RNA) anneal to form a stable double-stranded molecule. This principle is fundamental to techniques like microarrays and gene probes, allowing specific sequences to be identified. Like two halves of a zipper (single DNA strands) coming together and interlocking (base pairing) to form a complete zipper (double-stranded DNA).
gene probe — A length of DNA that has a complementary base sequence to another piece of DNA that you are trying to detect.
Gene probes are short, single-stranded DNA sequences, often labelled with a fluorescent or radioactive tag, used to identify specific DNA fragments. They bind to complementary sequences through DNA hybridisation, making the target DNA detectable. Like a specific key (probe) that only fits and lights up when it finds its matching lock (target DNA sequence).
Students often think a gene probe is the same as a primer, but actually a probe is used for detection and identification, while a primer is used to initiate DNA synthesis in PCR.
bioinformatics — The collection, processing and analysis of biological information and data using computer software.
Bioinformatics integrates biology, computer science, and statistics to manage and interpret large biological datasets, such as gene sequences, protein structures, and gene expression profiles. It is essential for making sense of the vast amount of data generated by modern genetic technologies. Like a digital librarian and data scientist for biological information, organizing, searching, and finding patterns in vast amounts of genetic data.
genome — The complete set of genes or genetic material present in a cell or an organism; the genome of a eukaryote includes the DNA in the nucleus and in the mitochondria; the genomes of plants include chloroplast DNA.
The genome encompasses all the hereditary information of an organism, encoded in its DNA (or RNA for some viruses). It includes both coding and non-coding regions, and in eukaryotes, it extends beyond nuclear DNA to include mitochondrial and chloroplast DNA. The complete instruction manual for building and operating an organism.
Genetic technology has profound applications in medicine, from producing therapeutic proteins to diagnosing and treating genetic disorders. Recombinant proteins, genetic screening, and gene therapy represent key areas where these advancements are transforming healthcare.
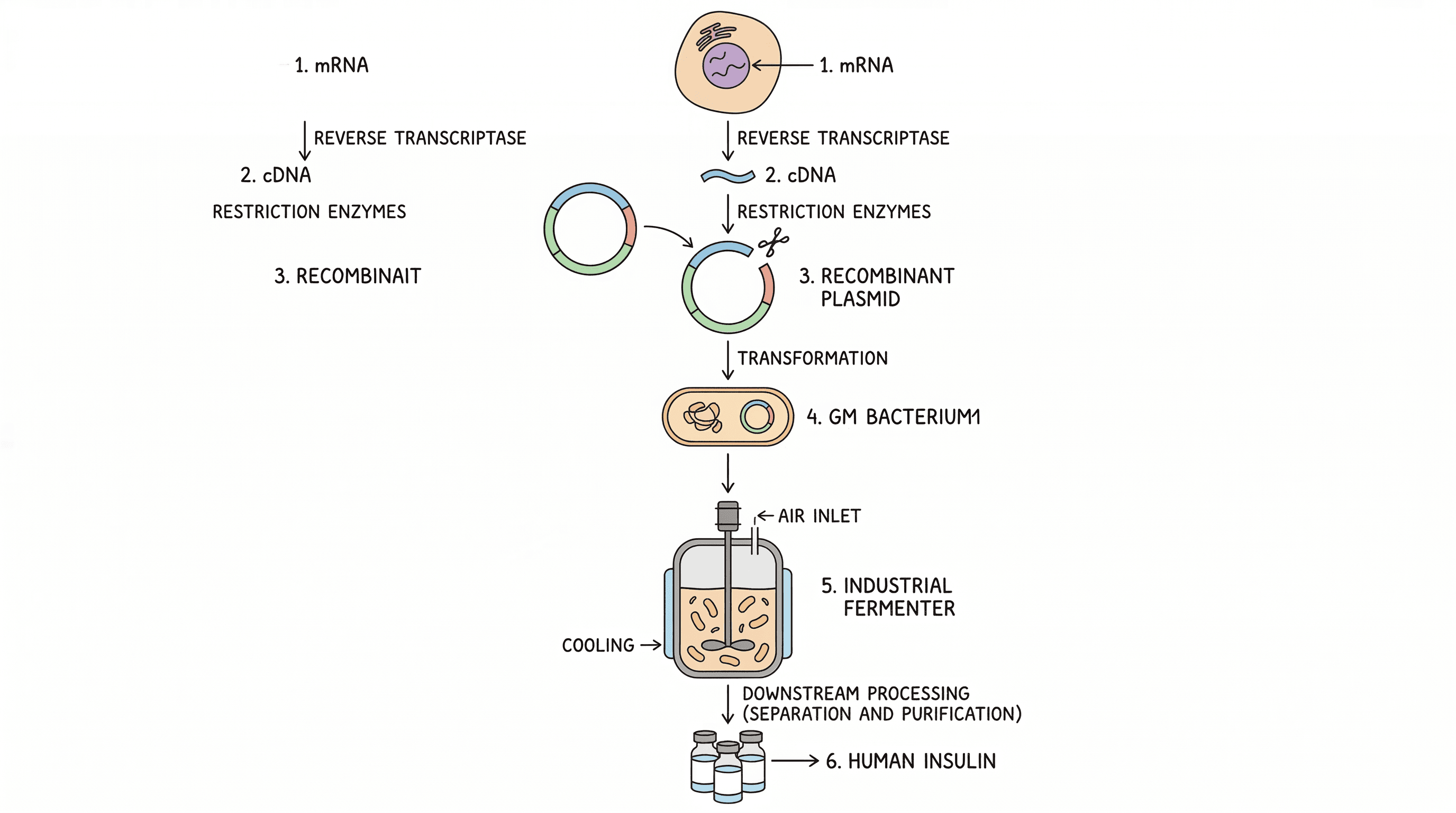
genetic screening — Testing an embryo, fetus or adult to find out whether a particular allele is present.
Genetic screening involves analyzing an individual's DNA to identify alleles associated with genetic diseases, even in the absence of symptoms. This can inform reproductive decisions, allow for early intervention, or guide preventative treatments. Like a detective searching for a specific clue (allele) in a person's genetic blueprint (DNA) to predict a future event (disease).
cystic fibrosis (CF) — A genetic disease caused by recessive alleles of the CFTR (cystic fibrosis transmembrane regulator) gene.
Cystic fibrosis is a common genetic disorder affecting mucus and sweat glands, leading to thick, sticky mucus that obstructs airways and ducts. It is caused by mutations in the CFTR gene, which codes for a chloride ion channel protein, impairing water movement across cell membranes. Imagine a faulty pump (CFTR protein) that can't move water properly, causing a thick, sticky sludge (mucus) to build up instead of a free-flowing liquid.
gene therapy — Treatment of a genetic disorder by inserting genetically corrected cells into the body or introducing functioning genes directly into affected cells.
Gene therapy aims to correct genetic defects by introducing functional genes into a patient's cells to replace or supplement faulty ones. While promising, it faces challenges in effectively delivering genes to target cells and ensuring their stable and safe expression. Like replacing a broken part (faulty gene) in a machine (cell) with a working part (functional gene) to restore its proper function.
Students often think gene therapy is a guaranteed cure for all genetic diseases, but actually it has proven very difficult, with limited successful treatments and ongoing challenges in delivery and safety.
When discussing gene therapy, acknowledge both its potential (curing genetic diseases) and the significant challenges (vector delivery, immune response, temporary effects, ethical concerns).
Genetic technology has revolutionized agriculture through the development of genetically modified organisms (GMOs). These modifications aim to improve crop yields, enhance nutritional value, and provide resistance to pests and herbicides, addressing global food security challenges.
Bt toxin — Insecticidal toxin produced by the bacterium Bacillus thuringiensis; the gene for Bt toxin is transferred to crop plants to make them resistant to insect pests.
Bt toxin is a natural insecticide that is harmless to humans and other animals but lethal to specific insect pests. By incorporating the Bt toxin gene into crop plants, these plants can produce their own insecticide, reducing the need for external pesticide sprays. Like giving a plant its own built-in defense system, so it can produce a natural repellent against specific attackers.
bacteriophage (phage) — A type of virus that infects bacteria; phages have double-stranded DNA as their genetic material.
Bacteriophages are viruses that specifically target and replicate within bacteria. Bacteria evolved restriction enzymes as a defense mechanism against these phages, cutting their DNA into harmless fragments. Like a specialized predator (phage) that only hunts a particular type of prey (bacteria).
The widespread application of genetic technology in medicine and food production raises significant social and ethical questions. These include concerns about safety, accessibility, potential misuse, and the long-term impact on human health and ecosystems, requiring careful consideration and regulation.
For questions on the social or ethical implications of genetic technology (e.g., GMOs, gene therapy), provide a balanced argument, discussing both potential benefits and potential risks or concerns.
Definitions Bank
genetic engineering
Any procedure in which the genetic information in an organism is changed by altering the base sequence of a gene or by introducing a gene from another organism; the organism is then said to be a genetically modified organism (GMO).
recombinant DNA (rDNA)
DNA made by artificially joining together pieces of DNA from two or more different species.
transgenic organism
Any organism that contains DNA from another source, such as from another individual of the same species or from a different species.
genetically modified organism (GMO)
Any organism that has had its DNA changed in a way that does not occur naturally or by selective breeding.
vector
A means of delivering genes into a cell used in gene technology; e.g. plasmids and viruses.
+17 more definitions
View all →Command Word Guide
| Explain | When asked to 'explain' genetic engineering, ensure you mention both altering existing base sequences and introducing genes from other organisms, and the resulting GMO. |
| Describe | When describing how to create a recombinant plasmid, always state that the *same* restriction enzyme must be used on both the gene and the plasmid to create complementary sticky ends. |
| Explain | When explaining PCR, ensure you describe the three distinct temperature-dependent stages: denaturation, annealing (with primers), and extension (with DNA polymerase). |
| Describe | When describing microarrays, focus on their ability to analyze *many genes simultaneously* and their use of *hybridisation* with labeled samples to detect gene presence or expression. |
+1 more
View all →Common Mistakes
Confusing genetic engineering with selective breeding.
Genetic engineering is the direct manipulation of DNA, often between species, while selective breeding relies on choosing individuals with desirable traits for reproduction.
Thinking restriction enzymes cut DNA randomly.
Restriction enzymes are highly specific and only cut at particular base sequences called recognition sites.
Assuming PCR copies the entire DNA sample.
PCR only amplifies the specific target region located between the two primers.
+3 more
View all →Generated by Nexelia Academy · nexeliaacademy.com